Carlos Uribe
Artificial intelligence for simplified patient-centered dosimetry in radiopharmaceutical therapies
Oct 14, 2025Abstract:KEY WORDS: Artificial Intelligence (AI), Theranostics, Dosimetry, Radiopharmaceutical Therapy (RPT), Patient-friendly dosimetry KEY POINTS - The rapid evolution of radiopharmaceutical therapy (RPT) highlights the growing need for personalized and patient-centered dosimetry. - Artificial Intelligence (AI) offers solutions to the key limitations in current dosimetry calculations. - The main advances on AI for simplified dosimetry toward patient-friendly RPT are reviewed. - Future directions on the role of AI in RPT dosimetry are discussed.
How To Segment in 3D Using 2D Models: Automated 3D Segmentation of Prostate Cancer Metastatic Lesions on PET Volumes Using Multi-Angle Maximum Intensity Projections and Diffusion Models
Jul 26, 2024Abstract:Prostate specific membrane antigen (PSMA) positron emission tomography/computed tomography (PET/CT) imaging provides a tremendously exciting frontier in visualization of prostate cancer (PCa) metastatic lesions. However, accurate segmentation of metastatic lesions is challenging due to low signal-to-noise ratios and variable sizes, shapes, and locations of the lesions. This study proposes a novel approach for automated segmentation of metastatic lesions in PSMA PET/CT 3D volumetric images using 2D denoising diffusion probabilistic models (DDPMs). Instead of 2D trans-axial slices or 3D volumes, the proposed approach segments the lesions on generated multi-angle maximum intensity projections (MA-MIPs) of the PSMA PET images, then obtains the final 3D segmentation masks from 3D ordered subset expectation maximization (OSEM) reconstruction of 2D MA-MIPs segmentations. Our proposed method achieved superior performance compared to state-of-the-art 3D segmentation approaches in terms of accuracy and robustness in detecting and segmenting small metastatic PCa lesions. The proposed method has significant potential as a tool for quantitative analysis of metastatic burden in PCa patients.
Thyroidiomics: An Automated Pipeline for Segmentation and Classification of Thyroid Pathologies from Scintigraphy Images
Jul 14, 2024



Abstract:The objective of this study was to develop an automated pipeline that enhances thyroid disease classification using thyroid scintigraphy images, aiming to decrease assessment time and increase diagnostic accuracy. Anterior thyroid scintigraphy images from 2,643 patients were collected and categorized into diffuse goiter (DG), multinodal goiter (MNG), and thyroiditis (TH) based on clinical reports, and then segmented by an expert. A ResUNet model was trained to perform auto-segmentation. Radiomic features were extracted from both physician (scenario 1) and ResUNet segmentations (scenario 2), followed by omitting highly correlated features using Spearman's correlation, and feature selection using Recursive Feature Elimination (RFE) with XGBoost as the core. All models were trained under leave-one-center-out cross-validation (LOCOCV) scheme, where nine instances of algorithms were iteratively trained and validated on data from eight centers and tested on the ninth for both scenarios separately. Segmentation performance was assessed using the Dice similarity coefficient (DSC), while classification performance was assessed using metrics, such as precision, recall, F1-score, accuracy, area under the Receiver Operating Characteristic (ROC AUC), and area under the precision-recall curve (PRC AUC). ResUNet achieved DSC values of 0.84$\pm$0.03, 0.71$\pm$0.06, and 0.86$\pm$0.02 for MNG, TH, and DG, respectively. Classification in scenario 1 achieved an accuracy of 0.76$\pm$0.04 and a ROC AUC of 0.92$\pm$0.02 while in scenario 2, classification yielded an accuracy of 0.74$\pm$0.05 and a ROC AUC of 0.90$\pm$0.02. The automated pipeline demonstrated comparable performance to physician segmentations on several classification metrics across different classes, effectively reducing assessment time while maintaining high diagnostic accuracy. Code available at: https://github.com/ahxmeds/thyroidiomics.git.
Observer study-based evaluation of TGAN architecture used to generate oncological PET images
Nov 28, 2023Abstract:The application of computer-vision algorithms in medical imaging has increased rapidly in recent years. However, algorithm training is challenging due to limited sample sizes, lack of labeled samples, as well as privacy concerns regarding data sharing. To address these issues, we previously developed (Bergen et al. 2022) a synthetic PET dataset for Head and Neck (H and N) cancer using the temporal generative adversarial network (TGAN) architecture and evaluated its performance segmenting lesions and identifying radiomics features in synthesized images. In this work, a two-alternative forced-choice (2AFC) observer study was performed to quantitatively evaluate the ability of human observers to distinguish between real and synthesized oncological PET images. In the study eight trained readers, including two board-certified nuclear medicine physicians, read 170 real/synthetic image pairs presented as 2D-transaxial using a dedicated web app. For each image pair, the observer was asked to identify the real image and input their confidence level with a 5-point Likert scale. P-values were computed using the binomial test and Wilcoxon signed-rank test. A heat map was used to compare the response accuracy distribution for the signed-rank test. Response accuracy for all observers ranged from 36.2% [27.9-44.4] to 63.1% [54.8-71.3]. Six out of eight observers did not identify the real image with statistical significance, indicating that the synthetic dataset was reasonably representative of oncological PET images. Overall, this study adds validity to the realism of our simulated H&N cancer dataset, which may be implemented in the future to train AI algorithms while favoring patient confidentiality and privacy protection.
FlowNet-PET: Unsupervised Learning to Perform Respiratory Motion Correction in PET Imaging
May 27, 2022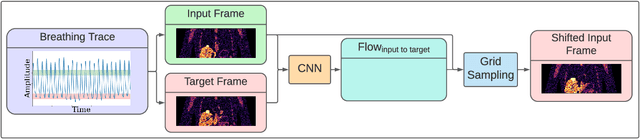
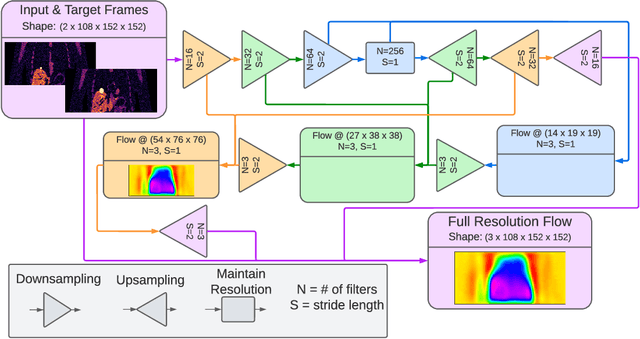
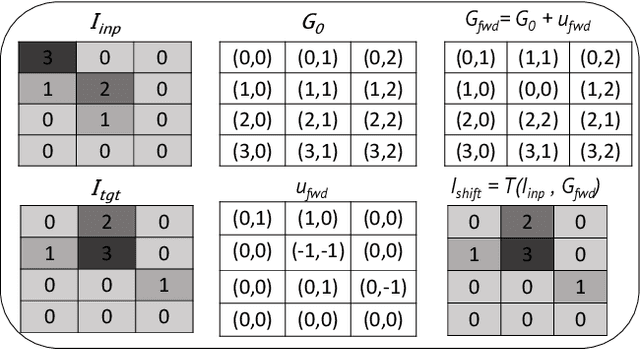
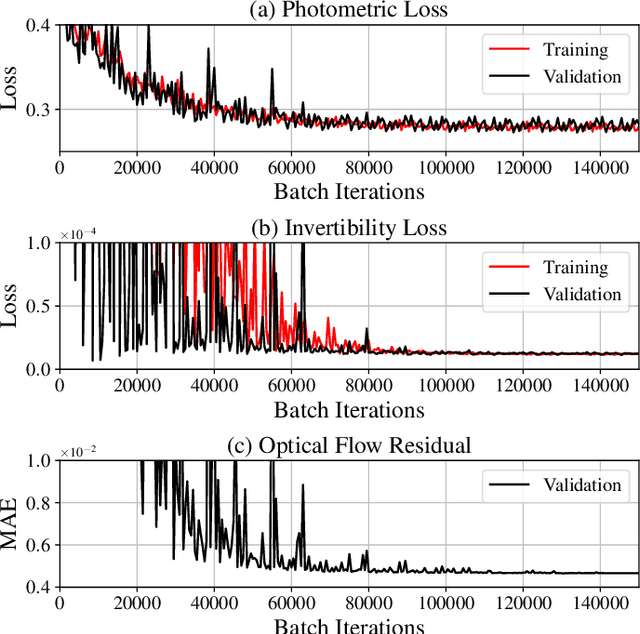
Abstract:To correct for breathing motion in PET imaging, an interpretable and unsupervised deep learning technique, FlowNet-PET, was constructed. The network was trained to predict the optical flow between two PET frames from different breathing amplitude ranges. As a result, the trained model groups different retrospectively-gated PET images together into a motion-corrected single bin, providing a final image with similar counting statistics as a non-gated image, but without the blurring effects that were initially observed. As a proof-of-concept, FlowNet-PET was applied to anthropomorphic digital phantom data, which provided the possibility to design robust metrics to quantify the corrections. When comparing the predicted optical flows to the ground truths, the median absolute error was found to be smaller than the pixel and slice widths, even for the phantom with a diaphragm movement of 21 mm. The improvements were illustrated by comparing against images without motion and computing the intersection over union (IoU) of the tumors as well as the enclosed activity and coefficient of variation (CoV) within the no-motion tumor volume before and after the corrections were applied. The average relative improvements provided by the network were 54%, 90%, and 76% for the IoU, total activity, and CoV, respectively. The results were then compared against the conventional retrospective phase binning approach. FlowNet-PET achieved similar results as retrospective binning, but only required one sixth of the scan duration. The code and data used for training and analysis has been made publicly available (https://github.com/teaghan/FlowNet_PET).
Tensor Radiomics: Paradigm for Systematic Incorporation of Multi-Flavoured Radiomics Features
Mar 12, 2022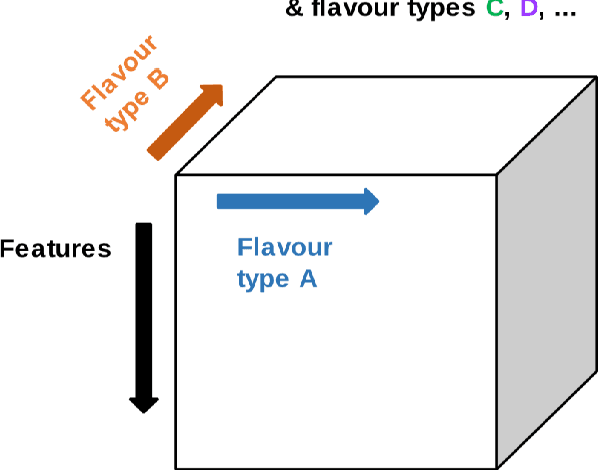
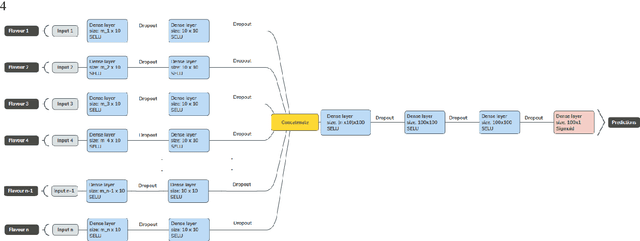
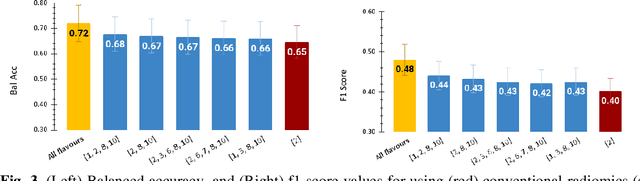
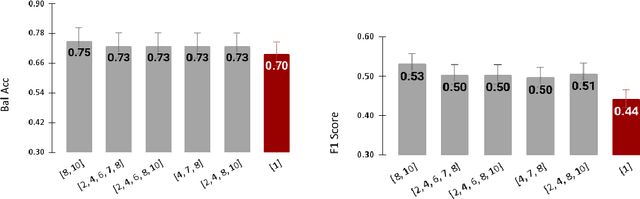
Abstract:Radiomics features extract quantitative information from medical images, towards the derivation of biomarkers for clinical tasks, such as diagnosis, prognosis, or treatment response assessment. Different image discretization parameters (e.g. bin number or size), convolutional filters, segmentation perturbation, or multi-modality fusion levels can be used to generate radiomics features and ultimately signatures. Commonly, only one set of parameters is used; resulting in only one value or flavour for a given RF. We propose tensor radiomics (TR) where tensors of features calculated with multiple combinations of parameters (i.e. flavours) are utilized to optimize the construction of radiomics signatures. We present examples of TR as applied to PET/CT, MRI, and CT imaging invoking machine learning or deep learning solutions, and reproducibility analyses: (1) TR via varying bin sizes on CT images of lung cancer and PET-CT images of head & neck cancer (HNC) for overall survival prediction. A hybrid deep neural network, referred to as TR-Net, along with two ML-based flavour fusion methods showed improved accuracy compared to regular rediomics features. (2) TR built from different segmentation perturbations and different bin sizes for classification of late-stage lung cancer response to first-line immunotherapy using CT images. TR improved predicted patient responses. (3) TR via multi-flavour generated radiomics features in MR imaging showed improved reproducibility when compared to many single-flavour features. (4) TR via multiple PET/CT fusions in HNC. Flavours were built from different fusions using methods, such as Laplacian pyramids and wavelet transforms. TR improved overall survival prediction. Our results suggest that the proposed TR paradigm has the potential to improve performance capabilities in different medical imaging tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge