Bruno Belucci
VertCoHiRF: Decentralized Vertical Clustering Beyond k-means
Feb 07, 2026Abstract:Vertical Federated Learning (VFL) enables collaborative analysis across parties holding complementary feature views of the same samples, yet existing approaches are largely restricted to distributed variants of $k$-means, requiring centralized coordination or the exchange of feature-dependent numerical statistics, and exhibiting limited robustness under heterogeneous views or adversarial behavior. We introduce VertCoHiRF, a fully decentralized framework for vertical federated clustering based on structural consensus across heterogeneous views, allowing each agent to apply a base clustering method adapted to its local feature space in a peer-to-peer manner. Rather than exchanging feature-dependent statistics or relying on noise injection for privacy, agents cluster their local views independently and reconcile their proposals through identifier-level consensus. Consensus is achieved via decentralized ordinal ranking to select representative medoids, progressively inducing a shared hierarchical clustering across agents. Communication is limited to sample identifiers, cluster labels, and ordinal rankings, providing privacy by design while supporting overlapping feature partitions and heterogeneous local clustering methods, and yielding an interpretable shared Cluster Fusion Hierarchy (CFH) that captures cross-view agreement at multiple resolutions.We analyze communication complexity and robustness, and experiments demonstrate competitive clustering performance in vertical federated settings.
kooplearn: A Scikit-Learn Compatible Library of Algorithms for Evolution Operator Learning
Dec 24, 2025



Abstract:kooplearn is a machine-learning library that implements linear, kernel, and deep-learning estimators of dynamical operators and their spectral decompositions. kooplearn can model both discrete-time evolution operators (Koopman/Transfer) and continuous-time infinitesimal generators. By learning these operators, users can analyze dynamical systems via spectral methods, derive data-driven reduced-order models, and forecast future states and observables. kooplearn's interface is compliant with the scikit-learn API, facilitating its integration into existing machine learning and data science workflows. Additionally, kooplearn includes curated benchmark datasets to support experimentation, reproducibility, and the fair comparison of learning algorithms. The software is available at https://github.com/Machine-Learning-Dynamical-Systems/kooplearn.
Automatic size and pose homogenization with spatial transformer network to improve and accelerate pediatric segmentation
Jul 06, 2021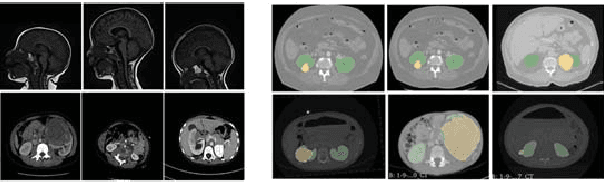

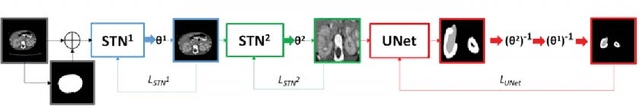
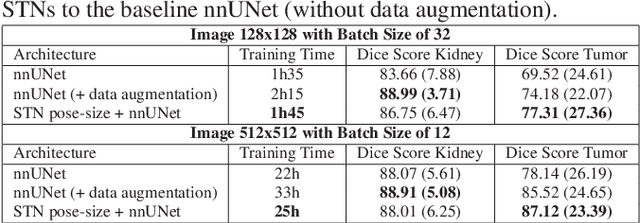
Abstract:Due to a high heterogeneity in pose and size and to a limited number of available data, segmentation of pediatric images is challenging for deep learning methods. In this work, we propose a new CNN architecture that is pose and scale invariant thanks to the use of Spatial Transformer Network (STN). Our architecture is composed of three sequential modules that are estimated together during training: (i) a regression module to estimate a similarity matrix to normalize the input image to a reference one; (ii) a differentiable module to find the region of interest to segment; (iii) a segmentation module, based on the popular UNet architecture, to delineate the object. Unlike the original UNet, which strives to learn a complex mapping, including pose and scale variations, from a finite training dataset, our segmentation module learns a simpler mapping focusing on images with normalized pose and size. Furthermore, the use of an automatic bounding box detection through STN allows saving time and especially memory, while keeping similar performance. We test the proposed method in kidney and renal tumor segmentation on abdominal pediatric CT scanners. Results indicate that the estimated STN homogenization of size and pose accelerates the segmentation (25h), compared to standard data-augmentation (33h), while obtaining a similar quality for the kidney (88.01\% of Dice score) and improving the renal tumor delineation (from 85.52\% to 87.12\%).
* ISBI 2021
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge