Banafshe Felfeliyan
FlexICL: A Flexible Visual In-context Learning Framework for Elbow and Wrist Ultrasound Segmentation
Oct 30, 2025Abstract:Elbow and wrist fractures are the most common fractures in pediatric populations. Automatic segmentation of musculoskeletal structures in ultrasound (US) can improve diagnostic accuracy and treatment planning. Fractures appear as cortical defects but require expert interpretation. Deep learning (DL) can provide real-time feedback and highlight key structures, helping lightly trained users perform exams more confidently. However, pixel-wise expert annotations for training remain time-consuming and costly. To address this challenge, we propose FlexICL, a novel and flexible in-context learning (ICL) framework for segmenting bony regions in US images. We apply it to an intra-video segmentation setting, where experts annotate only a small subset of frames, and the model segments unseen frames. We systematically investigate various image concatenation techniques and training strategies for visual ICL and introduce novel concatenation methods that significantly enhance model performance with limited labeled data. By integrating multiple augmentation strategies, FlexICL achieves robust segmentation performance across four wrist and elbow US datasets while requiring only 5% of the training images. It outperforms state-of-the-art visual ICL models like Painter, MAE-VQGAN, and conventional segmentation models like U-Net and TransUNet by 1-27% Dice coefficient on 1,252 US sweeps. These initial results highlight the potential of FlexICL as an efficient and scalable solution for US image segmentation well suited for medical imaging use cases where labeled data is scarce.
Sam2Rad: A Segmentation Model for Medical Images with Learnable Prompts
Sep 10, 2024



Abstract:Foundation models like the segment anything model require high-quality manual prompts for medical image segmentation, which is time-consuming and requires expertise. SAM and its variants often fail to segment structures in ultrasound (US) images due to domain shift. We propose Sam2Rad, a prompt learning approach to adapt SAM and its variants for US bone segmentation without human prompts. It introduces a prompt predictor network (PPN) with a cross-attention module to predict prompt embeddings from image encoder features. PPN outputs bounding box and mask prompts, and 256-dimensional embeddings for regions of interest. The framework allows optional manual prompting and can be trained end-to-end using parameter-efficient fine-tuning (PEFT). Sam2Rad was tested on 3 musculoskeletal US datasets: wrist (3822 images), rotator cuff (1605 images), and hip (4849 images). It improved performance across all datasets without manual prompts, increasing Dice scores by 2-7% for hip/wrist and up to 33% for shoulder data. Sam2Rad can be trained with as few as 10 labeled images and is compatible with any SAM architecture for automatic segmentation.
A Simple Framework Uniting Visual In-context Learning with Masked Image Modeling to Improve Ultrasound Segmentation
Mar 08, 2024



Abstract:Conventional deep learning models deal with images one-by-one, requiring costly and time-consuming expert labeling in the field of medical imaging, and domain-specific restriction limits model generalizability. Visual in-context learning (ICL) is a new and exciting area of research in computer vision. Unlike conventional deep learning, ICL emphasizes the model's ability to adapt to new tasks based on given examples quickly. Inspired by MAE-VQGAN, we proposed a new simple visual ICL method called SimICL, combining visual ICL pairing images with masked image modeling (MIM) designed for self-supervised learning. We validated our method on bony structures segmentation in a wrist ultrasound (US) dataset with limited annotations, where the clinical objective was to segment bony structures to help with further fracture detection. We used a test set containing 3822 images from 18 patients for bony region segmentation. SimICL achieved an remarkably high Dice coeffient (DC) of 0.96 and Jaccard Index (IoU) of 0.92, surpassing state-of-the-art segmentation and visual ICL models (a maximum DC 0.86 and IoU 0.76), with SimICL DC and IoU increasing up to 0.10 and 0.16. This remarkably high agreement with limited manual annotations indicates SimICL could be used for training AI models even on small US datasets. This could dramatically decrease the human expert time required for image labeling compared to conventional approaches, and enhance the real-world use of AI assistance in US image analysis.
Application Of Vision-Language Models For Assessing Osteoarthritis Disease Severity
Jan 12, 2024



Abstract:Osteoarthritis (OA) poses a global health challenge, demanding precise diagnostic methods. Current radiographic assessments are time consuming and prone to variability, prompting the need for automated solutions. The existing deep learning models for OA assessment are unimodal single task systems and they don't incorporate relevant text information such as patient demographics, disease history, or physician reports. This study investigates employing Vision Language Processing (VLP) models to predict OA severity using Xray images and corresponding reports. Our method leverages Xray images of the knee and diverse report templates generated from tabular OA scoring values to train a CLIP (Contrastive Language Image PreTraining) style VLP model. Furthermore, we incorporate additional contrasting captions to enforce the model to discriminate between positive and negative reports. Results demonstrate the efficacy of these models in learning text image representations and their contextual relationships, showcase potential advancement in OA assessment, and establish a foundation for specialized vision language models in medical contexts.
Self-supervised TransUNet for Ultrasound regional segmentation of the distal radius in children
Sep 18, 2023



Abstract:Supervised deep learning offers great promise to automate analysis of medical images from segmentation to diagnosis. However, their performance highly relies on the quality and quantity of the data annotation. Meanwhile, curating large annotated datasets for medical images requires a high level of expertise, which is time-consuming and expensive. Recently, to quench the thirst for large data sets with high-quality annotation, self-supervised learning (SSL) methods using unlabeled domain-specific data, have attracted attention. Therefore, designing an SSL method that relies on minimal quantities of labeled data has far-reaching significance in medical images. This paper investigates the feasibility of deploying the Masked Autoencoder for SSL (SSL-MAE) of TransUNet, for segmenting bony regions from children's wrist ultrasound scans. We found that changing the embedding and loss function in SSL-MAE can produce better downstream results compared to the original SSL-MAE. In addition, we determined that only pretraining TransUNet embedding and encoder with SSL-MAE does not work as well as TransUNet without SSL-MAE pretraining on downstream segmentation tasks.
Weakly Supervised Medical Image Segmentation With Soft Labels and Noise Robust Loss
Sep 16, 2022
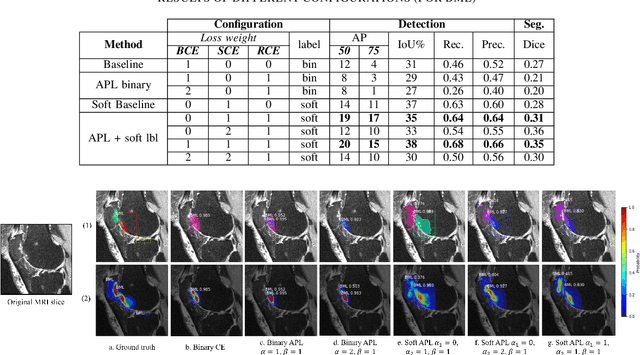
Abstract:Recent advances in deep learning algorithms have led to significant benefits for solving many medical image analysis problems. Training deep learning models commonly requires large datasets with expert-labeled annotations. However, acquiring expert-labeled annotation is not only expensive but also is subjective, error-prone, and inter-/intra- observer variability introduces noise to labels. This is particularly a problem when using deep learning models for segmenting medical images due to the ambiguous anatomical boundaries. Image-based medical diagnosis tools using deep learning models trained with incorrect segmentation labels can lead to false diagnoses and treatment suggestions. Multi-rater annotations might be better suited to train deep learning models with small training sets compared to single-rater annotations. The aim of this paper was to develop and evaluate a method to generate probabilistic labels based on multi-rater annotations and anatomical knowledge of the lesion features in MRI and a method to train segmentation models using probabilistic labels using normalized active-passive loss as a "noise-tolerant loss" function. The model was evaluated by comparing it to binary ground truth for 17 knees MRI scans for clinical segmentation and detection of bone marrow lesions (BML). The proposed method successfully improved precision 14, recall 22, and Dice score 8 percent compared to a binary cross-entropy loss function. Overall, the results of this work suggest that the proposed normalized active-passive loss using soft labels successfully mitigated the effects of noisy labels.
Self-Supervised-RCNN for Medical Image Segmentation with Limited Data Annotation
Jul 17, 2022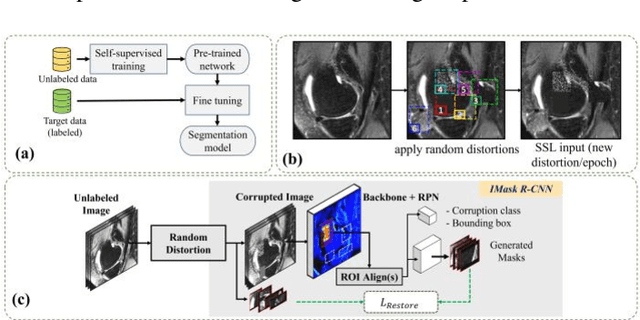
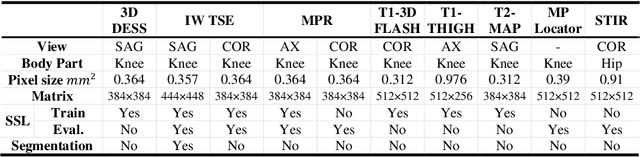
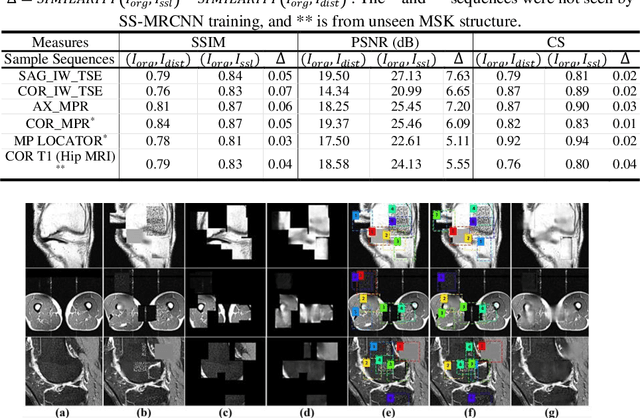
Abstract:Many successful methods developed for medical image analysis that are based on machine learning use supervised learning approaches, which often require large datasets annotated by experts to achieve high accuracy. However, medical data annotation is time-consuming and expensive, especially for segmentation tasks. To solve the problem of learning with limited labeled medical image data, an alternative deep learning training strategy based on self-supervised pretraining on unlabeled MRI scans is proposed in this work. Our pretraining approach first, randomly applies different distortions to random areas of unlabeled images and then predicts the type of distortions and loss of information. To this aim, an improved version of Mask-RCNN architecture has been adapted to localize the distortion location and recover the original image pixels. The effectiveness of the proposed method for segmentation tasks in different pre-training and fine-tuning scenarios is evaluated based on the Osteoarthritis Initiative dataset. Using this self-supervised pretraining method improved the Dice score by 20% compared to training from scratch. The proposed self-supervised learning is simple, effective, and suitable for different ranges of medical image analysis tasks including anomaly detection, segmentation, and classification.
Improved-Mask R-CNN: Towards an Accurate Generic MSK MRI instance segmentation platform
Jul 27, 2021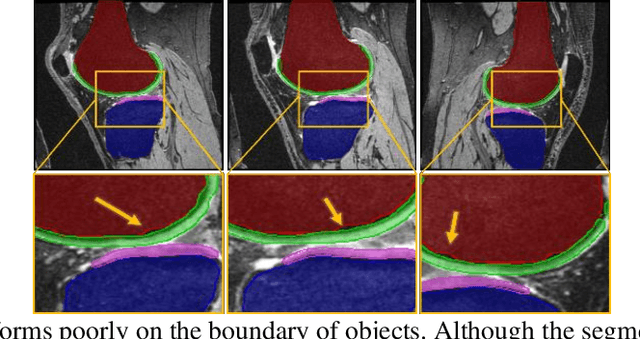
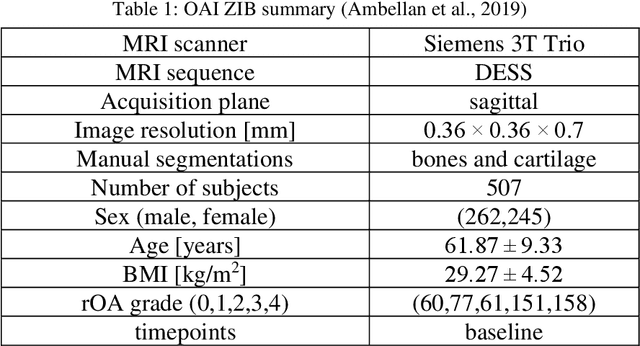
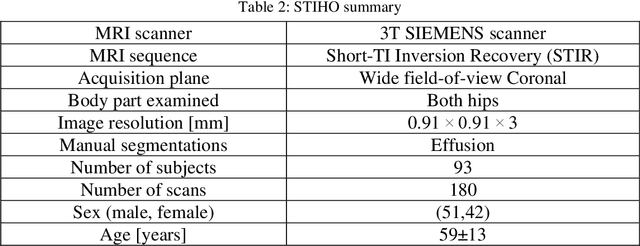
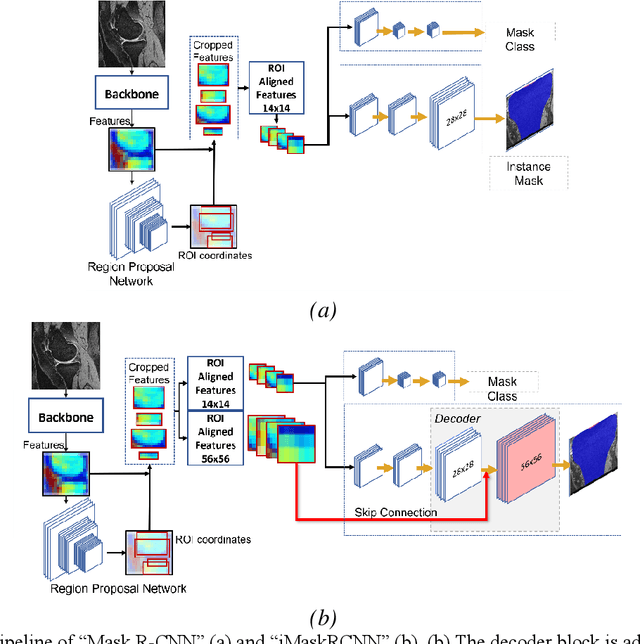
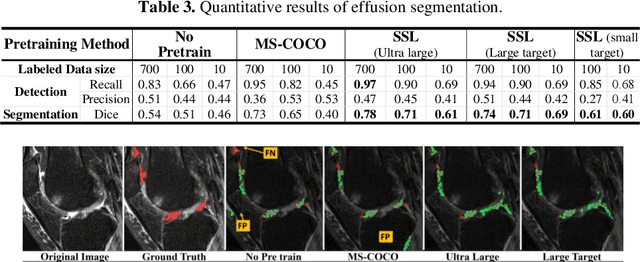
Abstract:Objective assessment of Magnetic Resonance Imaging (MRI) scans of osteoarthritis (OA) can address the limitation of the current OA assessment. Segmentation of bone, cartilage, and joint fluid is necessary for the OA objective assessment. Most of the proposed segmentation methods are not performing instance segmentation and suffer from class imbalance problems. This study deployed Mask R-CNN instance segmentation and improved it (improved-Mask R-CNN (iMaskRCNN)) to obtain a more accurate generalized segmentation for OA-associated tissues. Training and validation of the method were performed using 500 MRI knees from the Osteoarthritis Initiative (OAI) dataset and 97 MRI scans of patients with symptomatic hip OA. Three modifications to Mask R-CNN yielded the iMaskRCNN: adding a 2nd ROIAligned block, adding an extra decoder layer to the mask-header, and connecting them by a skip connection. The results were assessed using Hausdorff distance, dice score, and coefficients of variation (CoV). The iMaskRCNN led to improved bone and cartilage segmentation compared to Mask RCNN as indicated with the increase in dice score from 95% to 98% for the femur, 95% to 97% for tibia, 71% to 80% for femoral cartilage, and 81% to 82% for tibial cartilage. For the effusion detection, dice improved with iMaskRCNN 72% versus MaskRCNN 71%. The CoV values for effusion detection between Reader1 and Mask R-CNN (0.33), Reader1 and iMaskRCNN (0.34), Reader2 and Mask R-CNN (0.22), Reader2 and iMaskRCNN (0.29) are close to CoV between two readers (0.21), indicating a high agreement between the human readers and both Mask R-CNN and iMaskRCNN. Mask R-CNN and iMaskRCNN can reliably and simultaneously extract different scale articular tissues involved in OA, forming the foundation for automated assessment of OA. The iMaskRCNN results show that the modification improved the network performance around the edges.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge