B. Uma Shankar
Managing Diabetic Retinopathy with Deep Learning: A Data Centric Overview
Apr 02, 2026Abstract:Diabetic Retinopathy (DR) is a serious microvascular complication of diabetes, and one of the leading causes of vision loss worldwide. Although automated detection and grading, with Deep Learning (DL), can reduce the burden on ophthalmologists, it is constrained by the limited availability of high-quality datasets. Existing repositories often remain geographically narrow, contain limited samples, and exhibit inconsistent annotations or variable image quality; thereby, restricting their clinical reliability. This paper presents a comprehensive review and comparative analysis of fundus image datasets used in the management of DR. The study evaluates their usability across key tasks, including binary classification, severity grading, lesion localization, and multi-disease screening. It also categorizes the datasets by size, accessibility, and annotation type (such as image-level, lesion-level, and multi-disease). Finally, a recently published dataset is presented as a case study to illustrate broader challenges in dataset curation and usage. The review consolidates current knowledge while highlighting persistent gaps such as the lack of standardized lesion-level annotations and longitudinal data. It also outlines recommendations for future dataset development to support clinically reliable and explainable solutions in DR screening.
Weakly Supervised Patch Annotation for Improved Screening of Diabetic Retinopathy
Mar 04, 2026Abstract:Diabetic Retinopathy (DR) requires timely screening to prevent irreversible vision loss. However, its early detection remains a significant challenge since often the subtle pathological manifestations (lesions) get overlooked due to insufficient annotation. Existing literature primarily focuses on image-level supervision, weakly-supervised localization, and clustering-based representation learning, which fail to systematically annotate unlabeled lesion region(s) for refining the dataset. Expert-driven lesion annotation is labor-intensive and often incomplete, limiting the performance of deep learning models. We introduce Similarity-based Annotation via Feature-space Ensemble (SAFE), a two-stage framework that unifies weak supervision, contrastive learning, and patch-wise embedding inference, to systematically expand sparse annotations in the pathology. SAFE preserves fine-grained details of the lesion(s) under partial clinical supervision. In the first stage, a dual-arm Patch Embedding Network learns semantically structured, class-discriminative embeddings from expert annotated patches. Next, an ensemble of independent embedding spaces extrapolates labels to the unannotated regions based on spatial and semantic proximity. An abstention mechanism ensures trade-off between highly reliable annotation and noisy coverage. Experimental results demonstrate reliable separation of healthy and diseased patches, achieving upto 0.9886 accuracy. The annotation generated from SAFE substantially improves downstream tasks such as DR classification, demonstrating a substantial increase in F1-score of the diseased class and a performance gain as high as 0.545 in Area Under the Precision-Recall Curve (AUPRC). Qualitative analysis, with explainability, confirms that SAFE focuses on clinically relevant lesion patterns; and is further validated by ophthalmologists.
Collective Intelligent Strategy for Improved Segmentation of COVID-19 from CT
Dec 23, 2022Abstract:The devastation caused by the coronavirus pandemic makes it imperative to design automated techniques for a fast and accurate detection. We propose a novel non-invasive tool, using deep learning and imaging, for delineating COVID-19 infection in lungs. The Ensembling Attention-based Multi-scaled Convolution network (EAMC), employing Leave-One-Patient-Out (LOPO) training, exhibits high sensitivity and precision in outlining infected regions along with assessment of severity. The Attention module combines contextual with local information, at multiple scales, for accurate segmentation. Ensemble learning integrates heterogeneity of decision through different base classifiers. The superiority of EAMC, even with severe class imbalance, is established through comparison with existing state-of-the-art learning models over four publicly-available COVID-19 datasets. The results are suggestive of the relevance of deep learning in providing assistive intelligence to medical practitioners, when they are overburdened with patients as in pandemics. Its clinical significance lies in its unprecedented scope in providing low-cost decision-making for patients lacking specialized healthcare at remote locations.
Multi-Label Classification on Remote-Sensing Images
Jan 06, 2022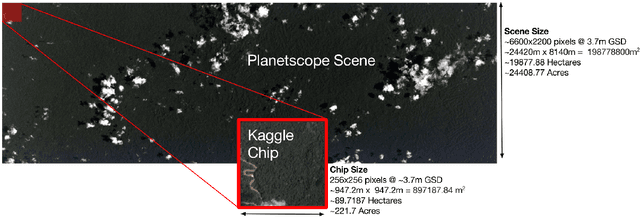
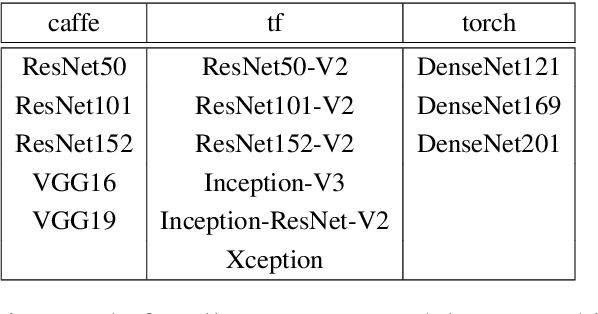

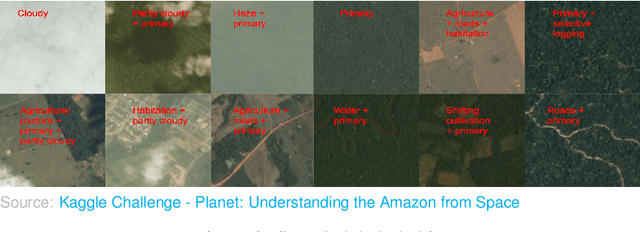
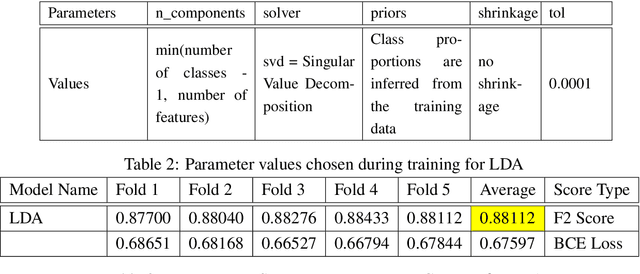
Abstract:Acquiring information on large areas on the earth's surface through satellite cameras allows us to see much more than we can see while standing on the ground. This assists us in detecting and monitoring the physical characteristics of an area like land-use patterns, atmospheric conditions, forest cover, and many unlisted aspects. The obtained images not only keep track of continuous natural phenomena but are also crucial in tackling the global challenge of severe deforestation. Among which Amazon basin accounts for the largest share every year. Proper data analysis would help limit detrimental effects on the ecosystem and biodiversity with a sustainable healthy atmosphere. This report aims to label the satellite image chips of the Amazon rainforest with atmospheric and various classes of land cover or land use through different machine learning and superior deep learning models. Evaluation is done based on the F2 metric, while for loss function, we have both sigmoid cross-entropy as well as softmax cross-entropy. Images are fed indirectly to the machine learning classifiers after only features are extracted using pre-trained ImageNet architectures. Whereas for deep learning models, ensembles of fine-tuned ImageNet pre-trained models are used via transfer learning. Our best score was achieved so far with the F2 metric is 0.927.
A CADe System for Gliomas in Brain MRI using Convolutional Neural Networks
Jun 20, 2018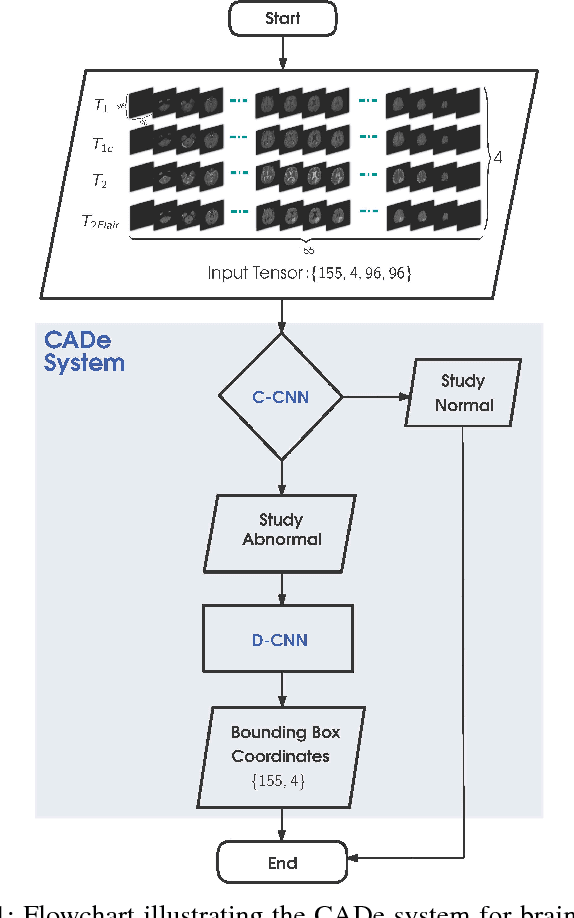
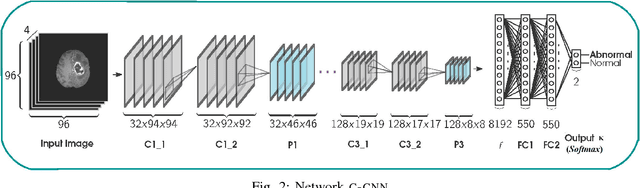
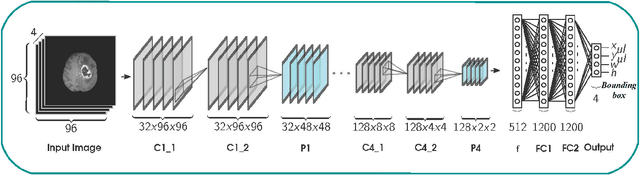
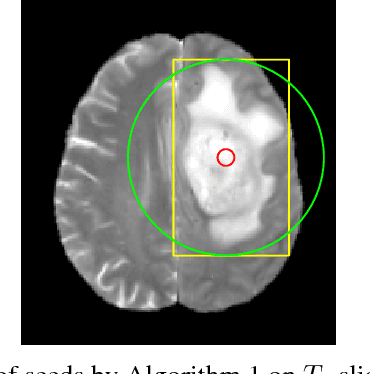
Abstract:Inspired by the success of Convolutional Neural Networks (CNN), we develop a novel Computer Aided Detection (CADe) system using CNN for Glioblastoma Multiforme (GBM) detection and segmentation from multi channel MRI data. A two-stage approach first identifies the presence of GBM. This is followed by a GBM localization in each "abnormal" MR slice. As part of the CADe system, two CNN architectures viz. Classification CNN (C-CNN) and Detection CNN (D-CNN) are employed. The CADe system considers MRI data consisting of four sequences ($T_1$, $T_{1c},$ $T_2$, and $T_{2FLAIR}$) as input, and automatically generates the bounding boxes encompassing the tumor regions in each slice which is deemed abnormal. Experimental results demonstrate that the proposed CADe system, when used as a preliminary step before segmentation, can allow improved delineation of tumor region while reducing false positives arising in normal areas of the brain. The GrowCut method, employed for tumor segmentation, typically requires a foreground and background seed region for initialization. Here the algorithm is initialized with seeds automatically generated from the output of the proposed CADe system, thereby resulting in improved performance as compared to that using random seeds.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge