Albert C. S. Chung
A Fine-Grain Error Map Prediction and Segmentation Quality Assessment Framework for Whole-Heart Segmentation
Jul 29, 2019
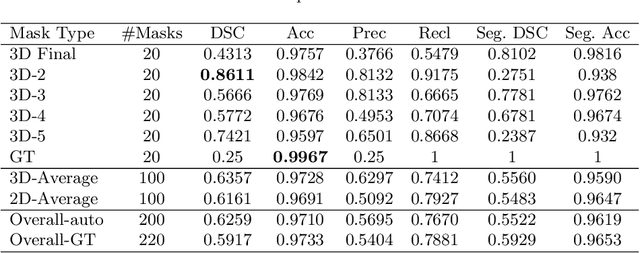
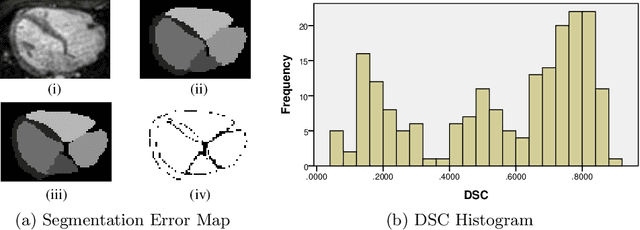
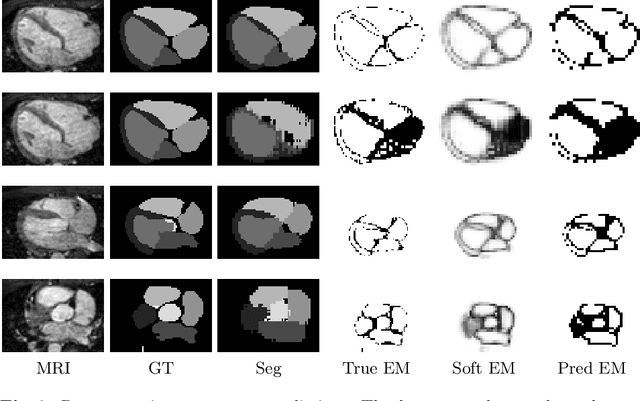
Abstract:When introducing advanced image computing algorithms, e.g., whole-heart segmentation, into clinical practice, a common suspicion is how reliable the automatically computed results are. In fact, it is important to find out the failure cases and identify the misclassified pixels so that they can be excluded or corrected for the subsequent analysis or diagnosis. However, it is not a trivial problem to predict the errors in a segmentation mask when ground truth (usually annotated by experts) is absent. In this work, we attempt to address the pixel-wise error map prediction problem and the per-case mask quality assessment problem using a unified deep learning (DL) framework. Specifically, we first formalize an error map prediction problem, then we convert it to a segmentation problem and build a DL network to tackle it. We also derive a quality indicator (QI) from a predicted error map to measure the overall quality of a segmentation mask. To evaluate the proposed framework, we perform extensive experiments on a public whole-heart segmentation dataset, i.e., MICCAI 2017 MMWHS. By 5-fold cross validation, we obtain an overall Dice score of 0.626 for the error map prediction task, and observe a high Pearson correlation coefficient (PCC) of 0.972 between QI and the actual segmentation accuracy (Acc), as well as a low mean absolute error (MAE) of 0.0048 between them, which evidences the efficacy of our method in both error map prediction and quality assessment.
A Unified Mammogram Analysis Method via Hybrid Deep Supervision
Aug 31, 2018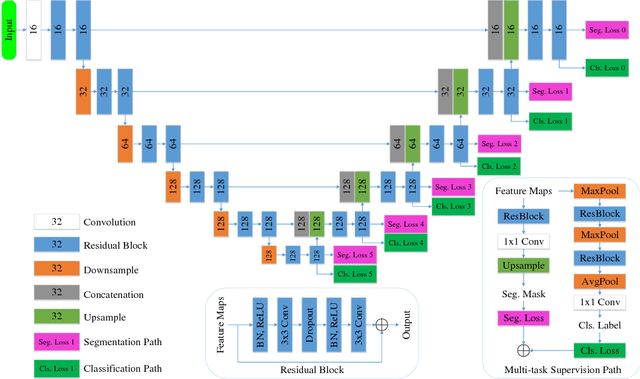
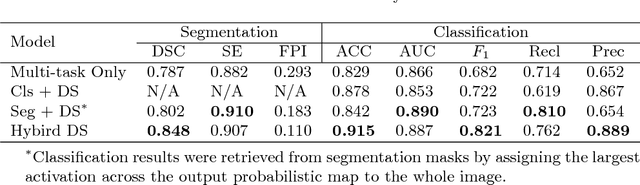
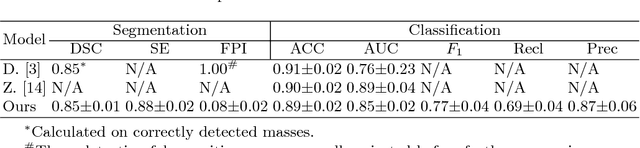
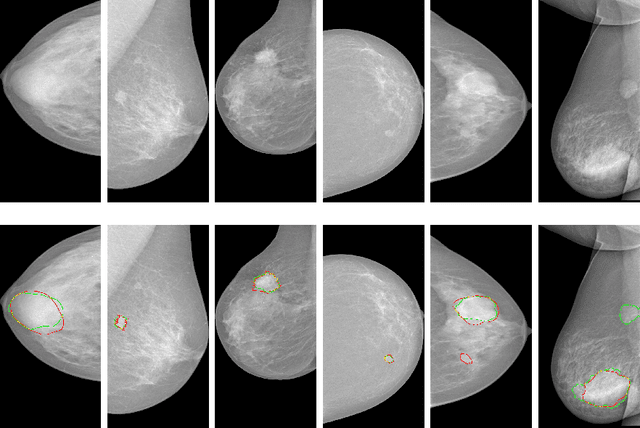
Abstract:Automatic mammogram classification and mass segmentation play a critical role in a computer-aided mammogram screening system. In this work, we present a unified mammogram analysis framework for both whole-mammogram classification and segmentation. Our model is designed based on a deep U-Net with residual connections, and equipped with the novel hybrid deep supervision (HDS) scheme for end-to-end multi-task learning. As an extension of deep supervision (DS), HDS not only can force the model to learn more discriminative features like DS, but also seamlessly integrates segmentation and classification tasks into one model, thus the model can benefit from both pixel-wise and image-wise supervisions. We extensively validate the proposed method on the widely-used INbreast dataset. Ablation study corroborates that pixel-wise and image-wise supervisions are mutually beneficial, evidencing the efficacy of HDS. The results of 5-fold cross validation indicate that our unified model matches state-of-the-art performance on both mammogram segmentation and classification tasks, which achieves an average segmentation Dice similarity coefficient (DSC) of 0.85 and a classification accuracy of 0.89. The code is available at https://github.com/angrypudding/hybrid-ds.
Deep supervision with additional labels for retinal vessel segmentation task
Jun 07, 2018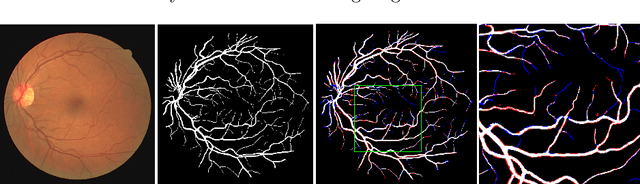

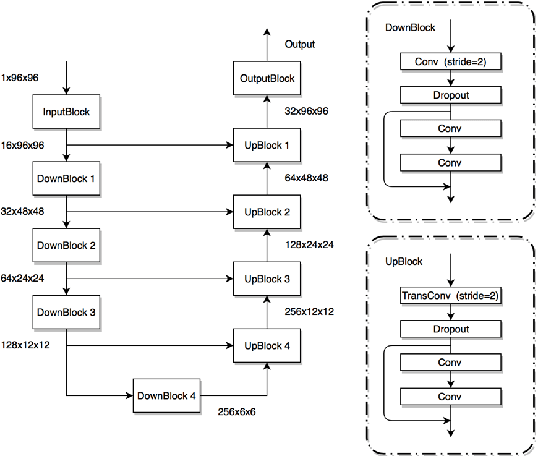
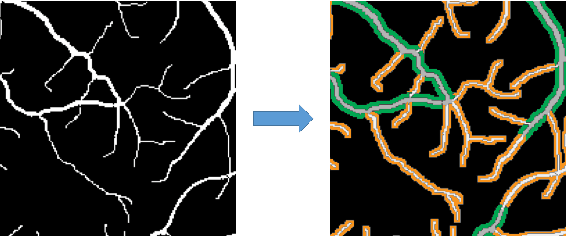
Abstract:Automatic analysis of retinal blood images is of vital importance in diagnosis tasks of retinopathy. Segmenting vessels accurately is a fundamental step in analysing retinal images. However, it is usually difficult due to various imaging conditions, low image contrast and the appearance of pathologies such as micro-aneurysms. In this paper, we propose a novel method with deep neural networks to solve this problem. We utilize U-net with residual connection to detect vessels. To achieve better accuracy, we introduce an edge-aware mechanism, in which we convert the original task into a multi-class task by adding additional labels on boundary areas. In this way, the network will pay more attention to the boundary areas of vessels and achieve a better performance, especially in tiny vessels detecting. Besides, side output layers are applied in order to give deep supervision and therefore help convergence. We train and evaluate our model on three databases: DRIVE, STARE, and CHASEDB1. Experimental results show that our method has a comparable performance with AUC of 97.99% on DRIVE and an efficient running time compared to the state-of-the-art methods.
3D Randomized Connection Network with Graph-based Label Inference
Nov 11, 2017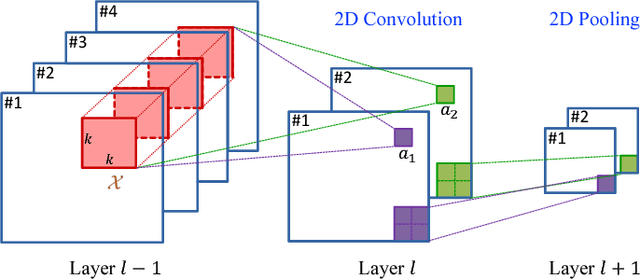
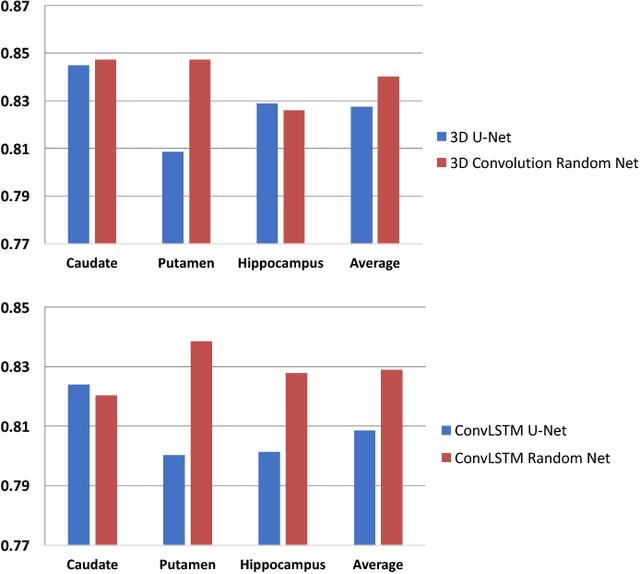
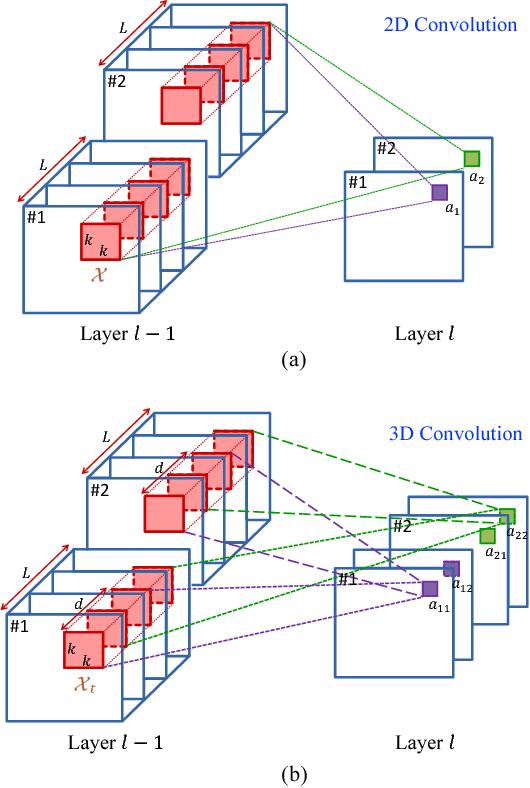
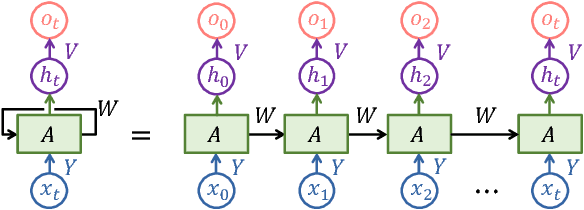
Abstract:In this paper, a novel 3D deep learning network is proposed for brain MR image segmentation with randomized connection, which can decrease the dependency between layers and increase the network capacity. The convolutional LSTM and 3D convolution are employed as network units to capture the long-term and short-term 3D properties respectively. To assemble these two kinds of spatial-temporal information and refine the deep learning outcomes, we further introduce an efficient graph-based node selection and label inference method. Experiments have been carried out on two publicly available databases and results demonstrate that the proposed method can obtain competitive performances as compared with other state-of-the-art methods.
Feature Sensitive Label Fusion with Random Walker for Atlas-based Image Segmentation
Nov 09, 2017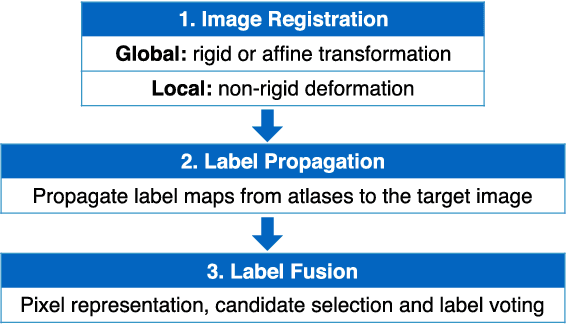
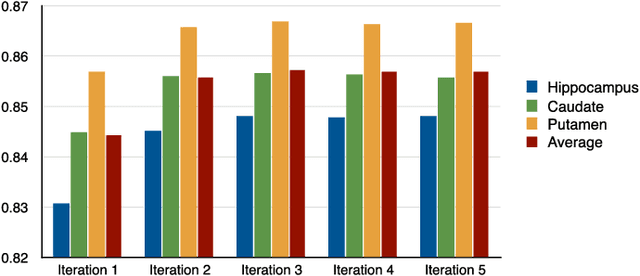
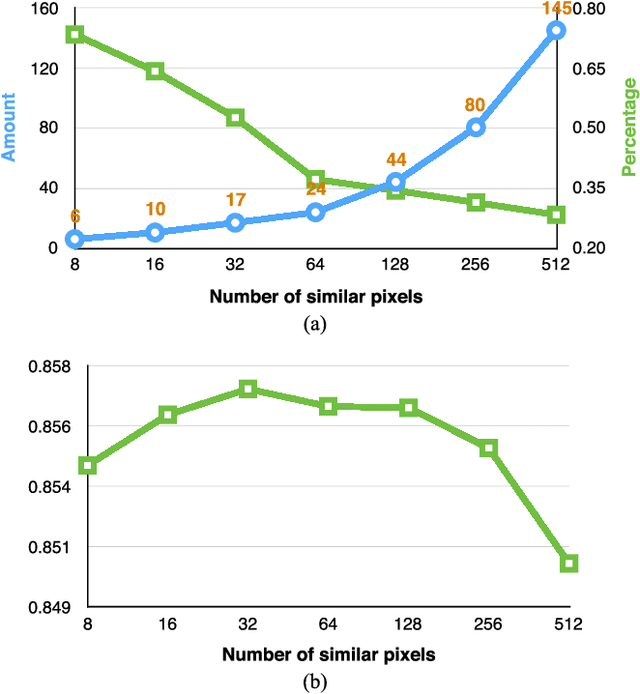
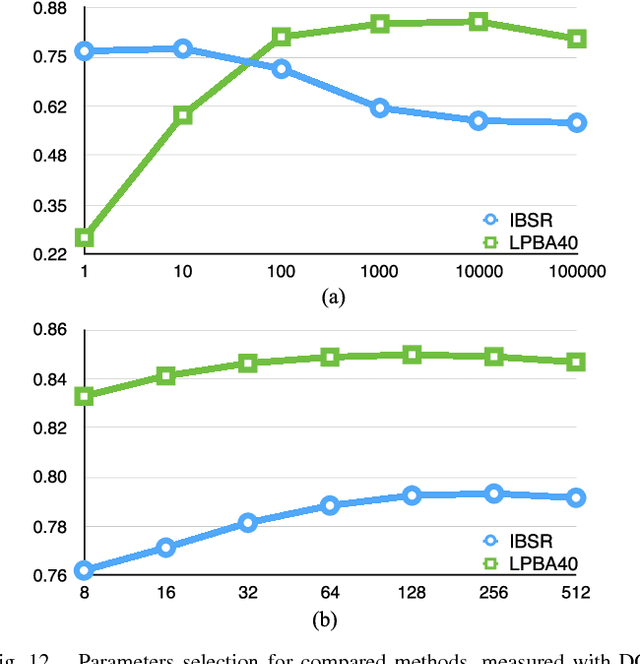
Abstract:In this paper, a novel label fusion method is proposed for brain magnetic resonance image segmentation. This label fusion method is formulated on a graph, which embraces both label priors from atlases and anatomical priors from target image. To represent a pixel in a comprehensive way, three kinds of feature vectors are generated, including intensity, gradient and structural signature. To select candidate atlas nodes for fusion, rather than exact searching, randomized k-d tree with spatial constraint is introduced as an efficient approximation for high-dimensional feature matching. Feature Sensitive Label Prior (FSLP), which takes both the consistency and variety of different features into consideration, is proposed to gather atlas priors. As FSLP is a non-convex problem, one heuristic approach is further designed to solve it efficiently. Moreover, based on the anatomical knowledge, parts of the target pixels are also employed as graph seeds to assist the label fusion process and an iterative strategy is utilized to gradually update the label map. The comprehensive experiments carried out on two publicly available databases give results to demonstrate that the proposed method can obtain better segmentation quality.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge