Akchunya Chanchal
A Causal Framework for Mitigating Data Shifts in Healthcare
Mar 13, 2026Abstract:Developing predictive models that perform reliably across diverse patient populations and heterogeneous environments is a core aim of medical research. However, generalization is only possible if the learned model is robust to statistical differences between data used for training and data seen at the time and place of deployment. Domain generalization methods provide strategies to address data shifts, but each method comes with its own set of assumptions and trade-offs. To apply these methods in healthcare, we must understand how domain shifts arise, what assumptions we prefer to make, and what our design constraints are. This article proposes a causal framework for the design of predictive models to improve generalization. Causality provides a powerful language to characterize and understand diverse domain shifts, regardless of data modality. This allows us to pinpoint why models fail to generalize, leading to more principled strategies to prepare for and adapt to shifts. We recommend general mitigation strategies, discussing trade-offs and highlighting existing work. Our causality-based perspective offers a critical foundation for developing robust, interpretable, and clinically relevant AI solutions in healthcare, paving the way for reliable real-world deployment.
SpecReX: Explainable AI for Raman Spectroscopy
Mar 18, 2025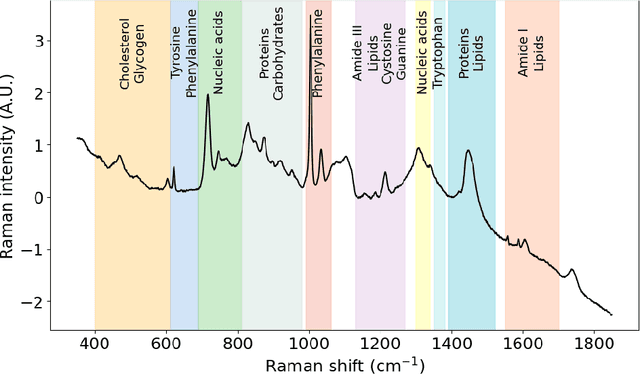
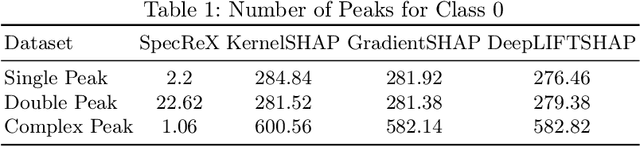
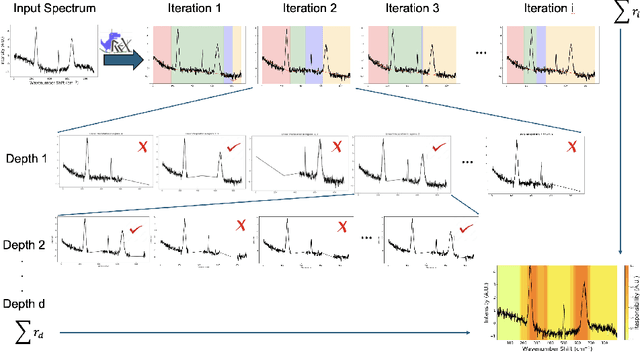
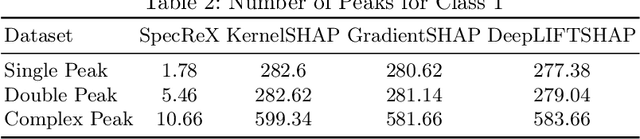
Abstract:Raman spectroscopy is becoming more common for medical diagnostics with deep learning models being increasingly used to leverage its full potential. However, the opaque nature of such models and the sensitivity of medical diagnosis together with regulatory requirements necessitate the need for explainable AI tools. We introduce SpecReX, specifically adapted to explaining Raman spectra. SpecReX uses the theory of actual causality to rank causal responsibility in a spectrum, quantified by iteratively refining mutated versions of the spectrum and testing if it retains the original classification. The explanations provided by SpecReX take the form of a responsibility map, highlighting spectral regions most responsible for the model to make a correct classification. To assess the validity of SpecReX, we create increasingly complex simulated spectra, in which a "ground truth" signal is seeded, to train a classifier. We then obtain SpecReX explanations and compare the results with another explainability tool. By using simulated spectra we establish that SpecReX localizes to the known differences between classes, under a number of conditions. This provides a foundation on which we can find the spectral features which differentiate disease classes. This is an important first step in proving the validity of SpecReX.
MRxaI: Black-Box Explainability for Image Classifiers in a Medical Setting
Nov 24, 2023



Abstract:Existing tools for explaining the output of image classifiers can be divided into white-box, which rely on access to the model internals, and black-box, agnostic to the model. As the usage of AI in the medical domain grows, so too does the usage of explainability tools. Existing work on medical image explanations focuses on white-box tools, such as gradcam. However, there are clear advantages to switching to a black-box tool, including the ability to use it with any classifier and the wide selection of black-box tools available. On standard images, black-box tools are as precise as white-box. In this paper we compare the performance of several black-box methods against gradcam on a brain cancer MRI dataset. We demonstrate that most black-box tools are not suitable for explaining medical image classifications and present a detailed analysis of the reasons for their shortcomings. We also show that one black-box tool, a causal explainability-based rex, performs as well as \gradcam.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge