Aditya Basu
PRISM: Patient Records Interpretation for Semantic Clinical Trial Matching using Large Language Models
Apr 27, 2024



Abstract:Clinical trial matching is the task of identifying trials for which patients may be potentially eligible. Typically, this task is labor-intensive and requires detailed verification of patient electronic health records (EHRs) against the stringent inclusion and exclusion criteria of clinical trials. This process is manual, time-intensive, and challenging to scale up, resulting in many patients missing out on potential therapeutic options. Recent advancements in Large Language Models (LLMs) have made automating patient-trial matching possible, as shown in multiple concurrent research studies. However, the current approaches are confined to constrained, often synthetic datasets that do not adequately mirror the complexities encountered in real-world medical data. In this study, we present the first, end-to-end large-scale empirical evaluation of clinical trial matching using real-world EHRs. Our study showcases the capability of LLMs to accurately match patients with appropriate clinical trials. We perform experiments with proprietary LLMs, including GPT-4 and GPT-3.5, as well as our custom fine-tuned model called OncoLLM and show that OncoLLM, despite its significantly smaller size, not only outperforms GPT-3.5 but also matches the performance of qualified medical doctors. All experiments were carried out on real-world EHRs that include clinical notes and available clinical trials from a single cancer center in the United States.
Onco-Retriever: Generative Classifier for Retrieval of EHR Records in Oncology
Apr 10, 2024



Abstract:Retrieving information from EHR systems is essential for answering specific questions about patient journeys and improving the delivery of clinical care. Despite this fact, most EHR systems still rely on keyword-based searches. With the advent of generative large language models (LLMs), retrieving information can lead to better search and summarization capabilities. Such retrievers can also feed Retrieval-augmented generation (RAG) pipelines to answer any query. However, the task of retrieving information from EHR real-world clinical data contained within EHR systems in order to solve several downstream use cases is challenging due to the difficulty in creating query-document support pairs. We provide a blueprint for creating such datasets in an affordable manner using large language models. Our method results in a retriever that is 30-50 F-1 points better than propriety counterparts such as Ada and Mistral for oncology data elements. We further compare our model, called Onco-Retriever, against fine-tuned PubMedBERT model as well. We conduct an extensive manual evaluation on real-world EHR data along with latency analysis of the different models and provide a path forward for healthcare organizations to build domain-specific retrievers.
Distilling Large Language Models for Matching Patients to Clinical Trials
Dec 15, 2023Abstract:The recent success of large language models (LLMs) has paved the way for their adoption in the high-stakes domain of healthcare. Specifically, the application of LLMs in patient-trial matching, which involves assessing patient eligibility against clinical trial's nuanced inclusion and exclusion criteria, has shown promise. Recent research has shown that GPT-3.5, a widely recognized LLM developed by OpenAI, can outperform existing methods with minimal 'variable engineering' by simply comparing clinical trial information against patient summaries. However, there are significant challenges associated with using closed-source proprietary LLMs like GPT-3.5 in practical healthcare applications, such as cost, privacy and reproducibility concerns. To address these issues, this study presents the first systematic examination of the efficacy of both proprietary (GPT-3.5, and GPT-4) and open-source LLMs (LLAMA 7B,13B, and 70B) for the task of patient-trial matching. Employing a multifaceted evaluation framework, we conducted extensive automated and human-centric assessments coupled with a detailed error analysis for each model. To enhance the adaptability of open-source LLMs, we have created a specialized synthetic dataset utilizing GPT-4, enabling effective fine-tuning under constrained data conditions. Our findings reveal that open-source LLMs, when fine-tuned on this limited and synthetic dataset, demonstrate performance parity with their proprietary counterparts. This presents a massive opportunity for their deployment in real-world healthcare applications. To foster further research and applications in this field, we release both the annotated evaluation dataset along with the fine-tuned LLM -- Trial-LLAMA -- for public use.
Placing Facts on a Timeline: A Classification cum Coref Resolution Approach
Jun 28, 2022
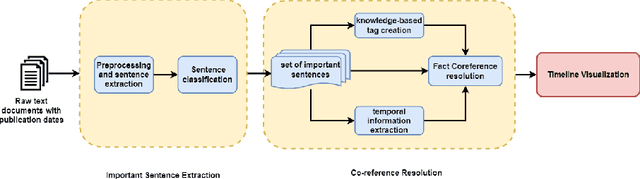
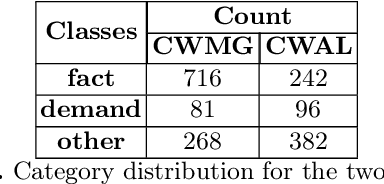
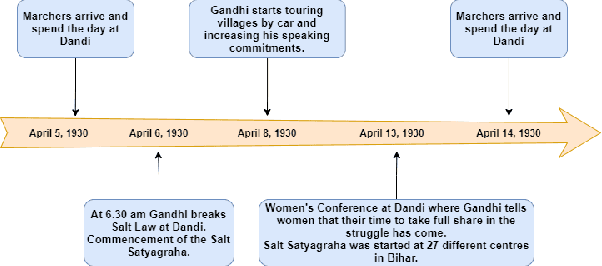
Abstract:A timeline provides one of the most effective ways to visualize the important historical facts that occurred over a period of time, presenting the insights that may not be so apparent from reading the equivalent information in textual form. By leveraging generative adversarial learning for important sentence classification and by assimilating knowledge based tags for improving the performance of event coreference resolution we introduce a two staged system for event timeline generation from multiple (historical) text documents. We demonstrate our results on two manually annotated historical text documents. Our results can be extremely helpful for historians, in advancing research in history and in understanding the socio-political landscape of a country as reflected in the writings of famous personas.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge