Yoshinobu Kawahara
Physically-interpretable classification of network dynamics for complex collective motions
May 13, 2019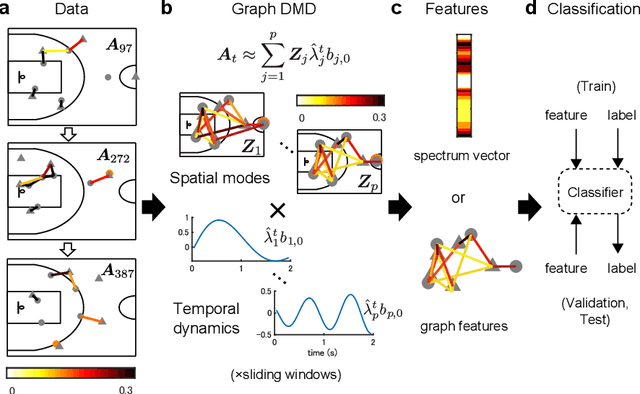
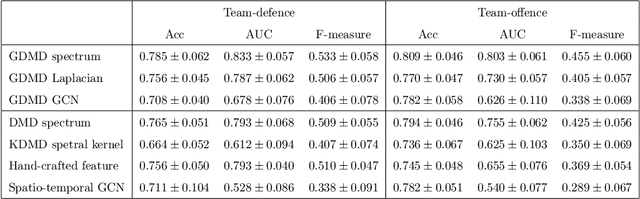
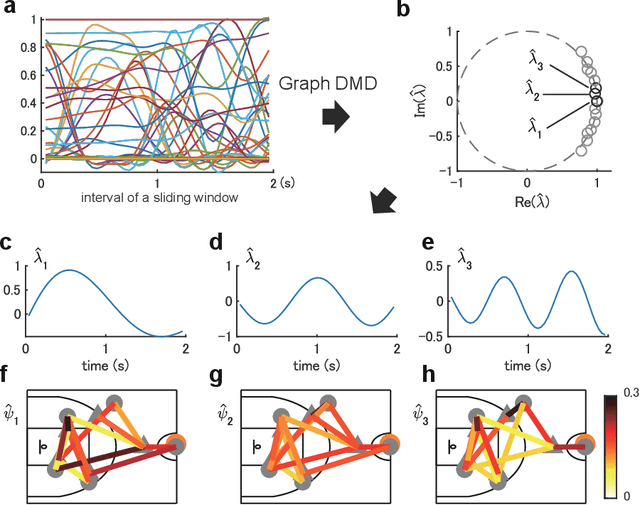
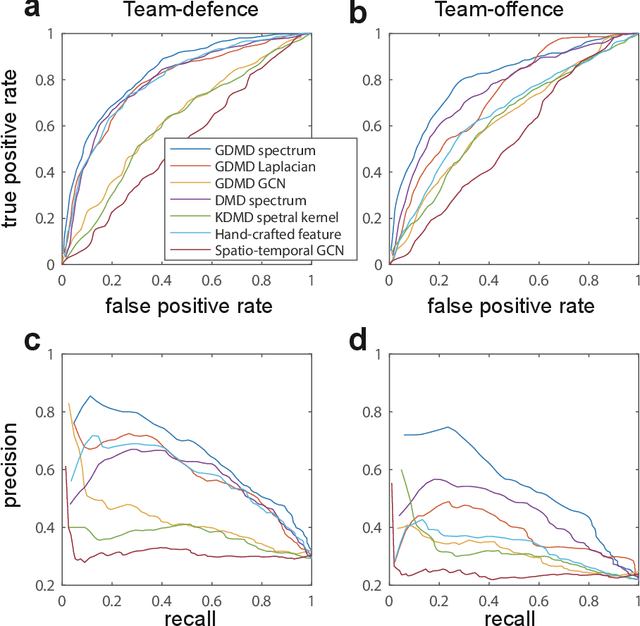
Abstract:Understanding complex network dynamics is a fundamental issue in various scientific and engineering fields. Network theory is capable of revealing the relationship between elements and their propagation; however, for complex collective motions, the network properties often transiently and complexly change. A fundamental question addressed here pertains to the classification of collective motion network based on physically-interpretable dynamical properties. Here we apply a data-driven spectral analysis called graph dynamic mode decomposition, which obtains the dynamical properties for collective motion classification. Using a ballgame as an example, we classified the strategic collective motions in different global behaviours and discovered that, in addition to the physical properties, the contextual node information was critical for classification. Furthermore, we discovered the label-specific stronger spectra in the relationship among the nearest agents, providing physical and semantic interpretations. Our approach contributes to the understanding of complex networks involving collective motions from the perspective of nonlinear dynamical systems.
An efficient branch-and-cut algorithm for approximately submodular function maximization
Apr 26, 2019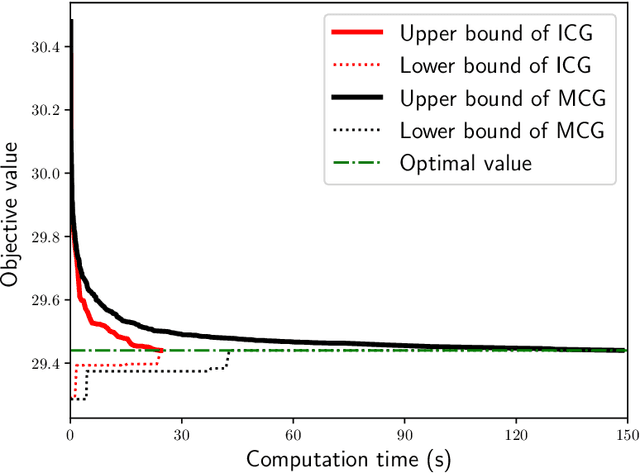
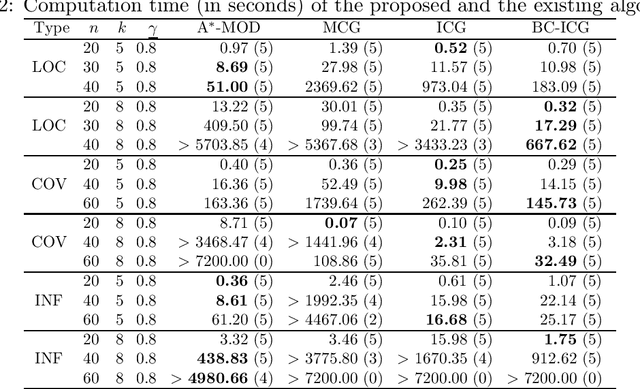
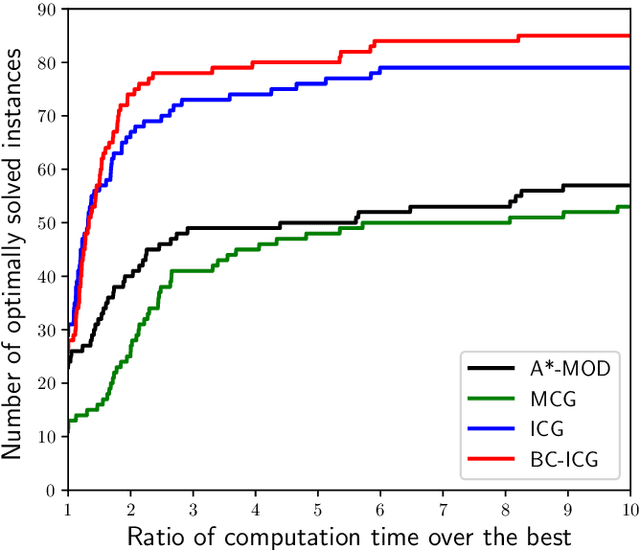
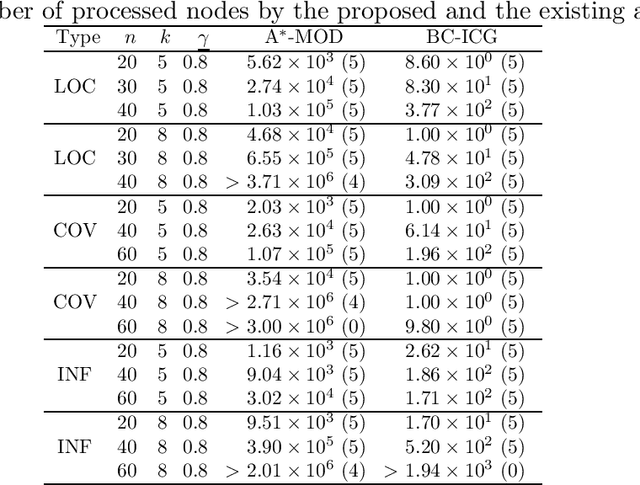
Abstract:When approaching to problems in computer science, we often encounter situations where a subset of a finite set maximizing some utility function needs to be selected. Some of such utility functions are known to be approximately submodular. For the problem of maximizing an approximately submodular function (ASFM problem), a greedy algorithm quickly finds good feasible solutions for many instances while guaranteeing ($1-e^{-\gamma}$)-approximation ratio for a given submodular ratio $\gamma$. However, we still encounter its applications that ask more accurate or exactly optimal solutions within a reasonable computation time. In this paper, we present an efficient branch-and-cut algorithm for the non-decreasing ASFM problem based on its binary integer programming (BIP) formulation with an exponential number of constraints. To this end, we first derive a BIP formulation of the ASFM problem and then, develop an improved constraint generation algorithm that starts from a reduced BIP problem with a small subset of constraints and repeats solving the reduced BIP problem while adding a promising set of constraints at each iteration. Moreover, we incorporate it into a branch-and-cut algorithm to attain good upper bounds while solving a smaller number of nodes of a search tree. The computational results for three types of well-known benchmark instances show that our algorithm performs better than the conventional exact algorithms.
Regularizing Generative Models Using Knowledge of Feature Dependence
Feb 06, 2019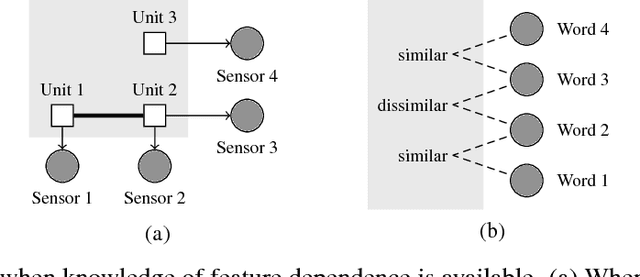
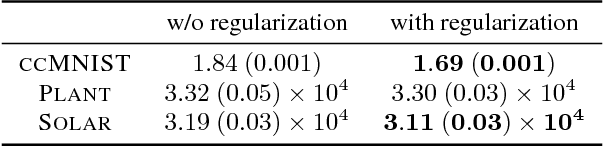
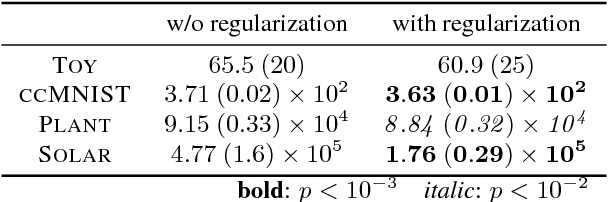


Abstract:Generative modeling is a fundamental problem in machine learning with many potential applications. Efficient learning of generative models requires available prior knowledge to be exploited as much as possible. In this paper, we propose a method to exploit prior knowledge of relative dependence between features for learning generative models. Such knowledge is available, for example, when side-information on features is present. We incorporate the prior knowledge by forcing marginals of the learned generative model to follow a prescribed relative feature dependence. To this end, we formulate a regularization term using a kernel-based dependence criterion. The proposed method can be incorporated straightforwardly into many optimization-based learning schemes of generative models, including variational autoencoders and generative adversarial networks. We show the effectiveness of the proposed method in experiments with multiple types of datasets and models.
An efficient branch-and-bound algorithm for submodular function maximization
Nov 10, 2018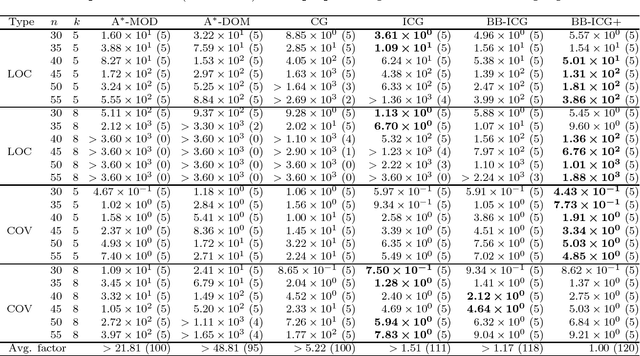
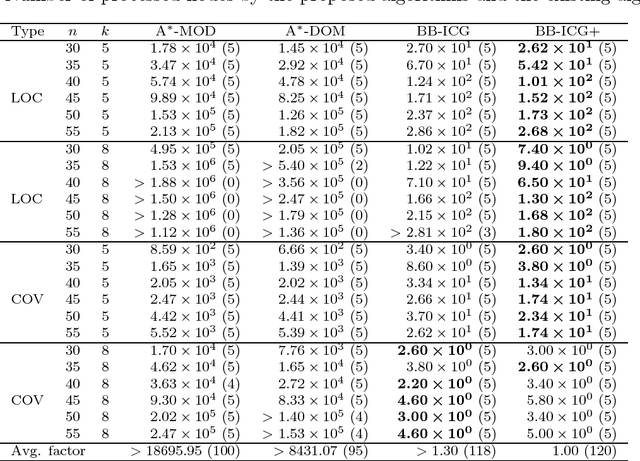
Abstract:The submodular function maximization is an attractive optimization model that appears in many real applications. Although a variety of greedy algorithms quickly find good feasible solutions for many instances while guaranteeing (1-1/e)-approximation ratio, we still encounter many real applications that ask optimal or better feasible solutions within reasonable computation time. In this paper, we present an efficient branch-and-bound algorithm for the non-decreasing submodular function maximization problem based on its binary integer programming (BIP) formulation with a huge number of constraints. Nemhauser and Wolsey developed an exact algorithm called the constraint generation algorithm that starts from a reduced BIP problem with a small subset of constraints taken from the constraints and repeats solving a reduced BIP problem while adding a new constraint at each iteration. However, their algorithm is still computationally expensive due to many reduced BIP problems to be solved. To overcome this, we propose an improved constraint generation algorithm to add a promising set of constraints at each iteration. We incorporate it into a branch-and-bound algorithm to attain good upper bounds while solving a smaller number of reduced BIP problems. According to computational results for well-known benchmark instances, our algorithm achieved better performance than the state-of-the-art exact algorithms.
Metric on Nonlinear Dynamical Systems with Perron-Frobenius Operators
Oct 31, 2018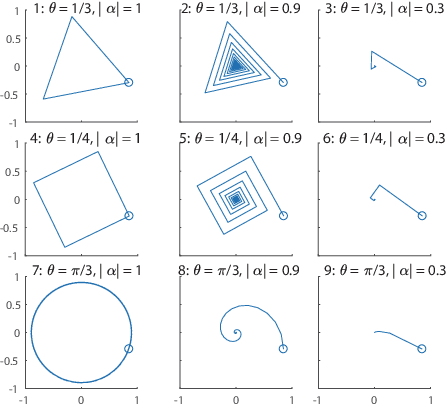
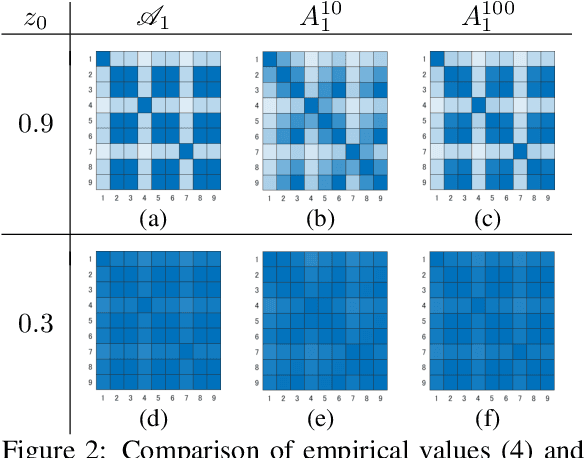
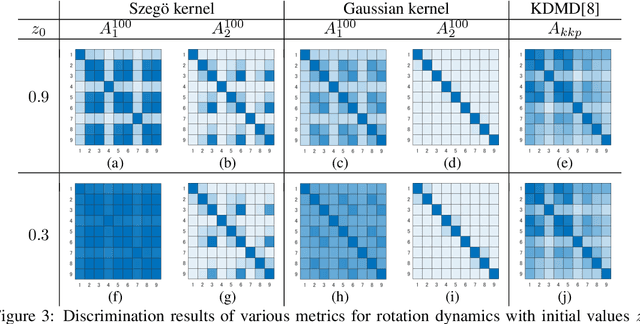
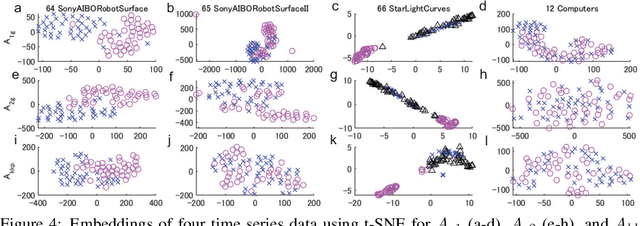
Abstract:The development of a metric for structural data is a long-term problem in pattern recognition and machine learning. In this paper, we develop a general metric for comparing nonlinear dynamical systems that is defined with Perron-Frobenius operators in reproducing kernel Hilbert spaces. Our metric includes the existing fundamental metrics for dynamical systems, which are basically defined with principal angles between some appropriately-chosen subspaces, as its special cases. We also describe the estimation of our metric from finite data. We empirically illustrate our metric with an example of rotation dynamics in a unit disk in a complex plane, and evaluate the performance with real-world time-series data.
Dynamic mode decomposition in vector-valued reproducing kernel Hilbert spaces for extracting dynamical structure among observables
Oct 24, 2018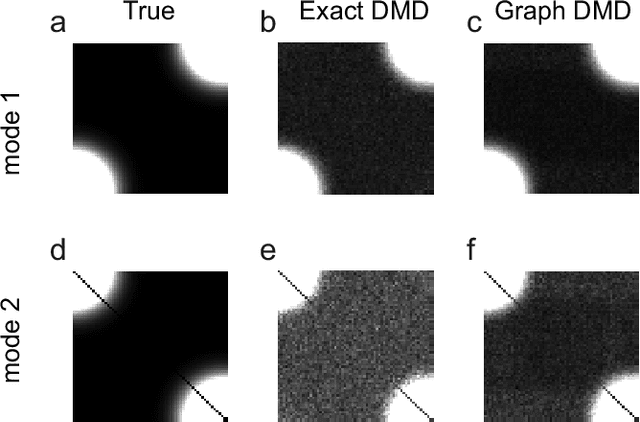
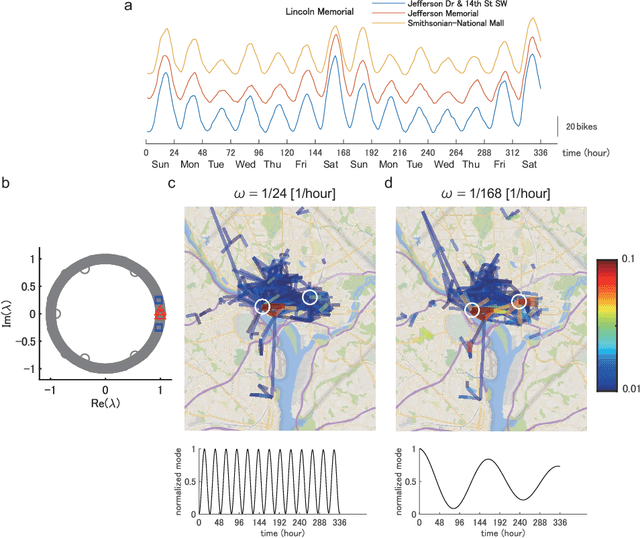
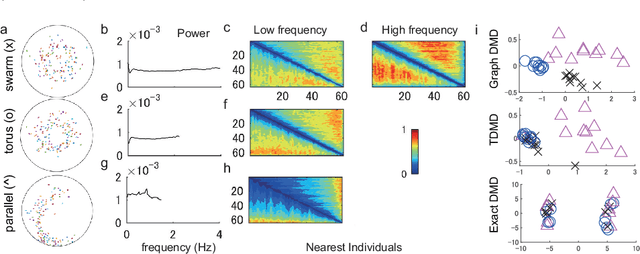
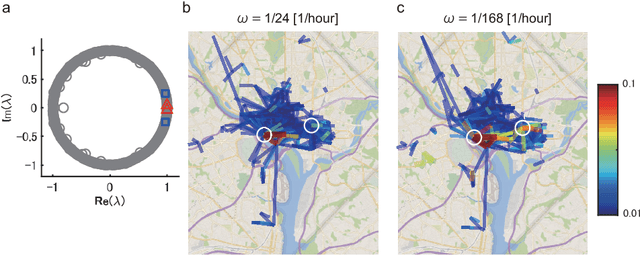
Abstract:Understanding nonlinear dynamical systems (NLDSs) is challenging in a variety of engineering and scientific fields. Dynamic mode decomposition (DMD), which is a numerical algorithm for the spectral analysis of Koopman operators, has been attracting attention as a way of obtaining global modal descriptions of NLDSs without requiring explicit prior knowledge. However, since existing DMD algorithms are in principle formulated based on the concatenation of scalar observables, it is not directly applicable to data with dependent structures among observables, which take, for example, the form of a sequence of graphs. In this paper, we formulate Koopman spectral analysis for NLDSs with structures among observables and propose an estimation algorithm for this problem. This method can extract and visualize the underlying low-dimensional global dynamics of NLDSs with structures among observables from data, which can be useful in understanding the underlying dynamics of such NLDSs. To this end, we first formulate the problem of estimating spectra of the Koopman operator defined in vector-valued reproducing kernel Hilbert spaces, and then develop an estimation procedure for this problem by reformulating tensor-based DMD. As a special case of our method, we propose the method named as Graph DMD, which is a numerical algorithm for Koopman spectral analysis of graph dynamical systems, using a sequence of adjacency matrices. We investigate the empirical performance of our method by using synthetic and real-world data.
Learning Koopman Invariant Subspaces for Dynamic Mode Decomposition
Jan 30, 2018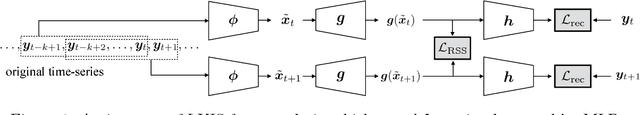
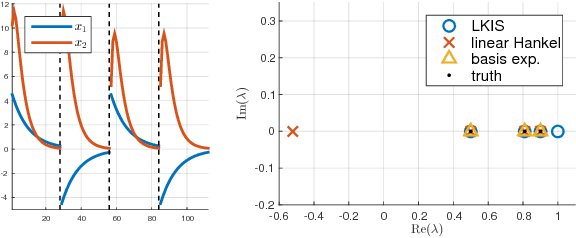
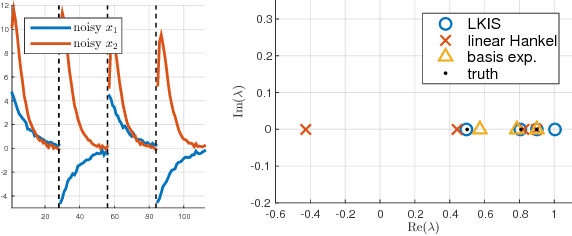

Abstract:Spectral decomposition of the Koopman operator is attracting attention as a tool for the analysis of nonlinear dynamical systems. Dynamic mode decomposition is a popular numerical algorithm for Koopman spectral analysis; however, we often need to prepare nonlinear observables manually according to the underlying dynamics, which is not always possible since we may not have any a priori knowledge about them. In this paper, we propose a fully data-driven method for Koopman spectral analysis based on the principle of learning Koopman invariant subspaces from observed data. To this end, we propose minimization of the residual sum of squares of linear least-squares regression to estimate a set of functions that transforms data into a form in which the linear regression fits well. We introduce an implementation with neural networks and evaluate performance empirically using nonlinear dynamical systems and applications.
Parametric Maxflows for Structured Sparse Learning with Convex Relaxations of Submodular Functions
Sep 14, 2015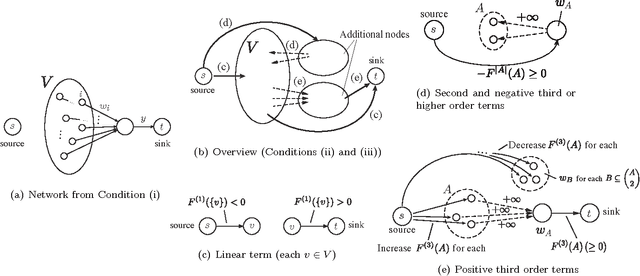
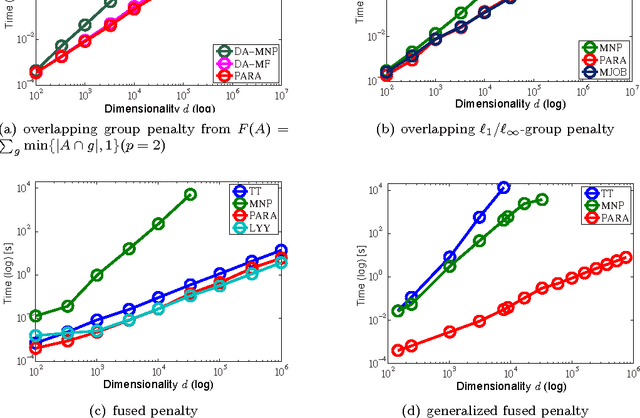
Abstract:The proximal problem for structured penalties obtained via convex relaxations of submodular functions is known to be equivalent to minimizing separable convex functions over the corresponding submodular polyhedra. In this paper, we reveal a comprehensive class of structured penalties for which penalties this problem can be solved via an efficiently solvable class of parametric maxflow optimization. We then show that the parametric maxflow algorithm proposed by Gallo et al. and its variants, which runs, in the worst-case, at the cost of only a constant factor of a single computation of the corresponding maxflow optimization, can be adapted to solve the proximal problems for those penalties. Several existing structured penalties satisfy these conditions; thus, regularized learning with these penalties is solvable quickly using the parametric maxflow algorithm. We also investigate the empirical runtime performance of the proposed framework.
A direct method for estimating a causal ordering in a linear non-Gaussian acyclic model
Aug 09, 2014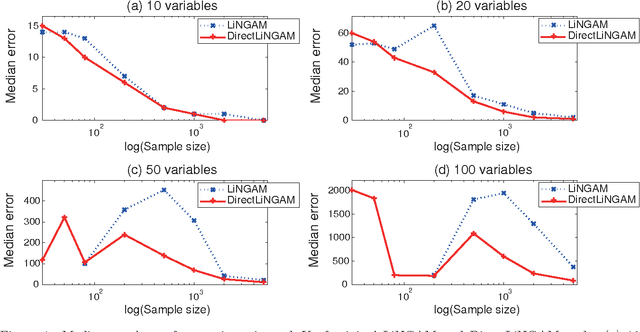
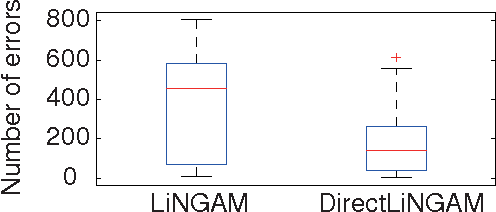
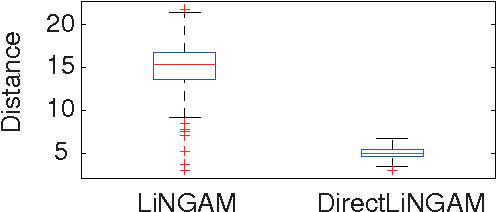
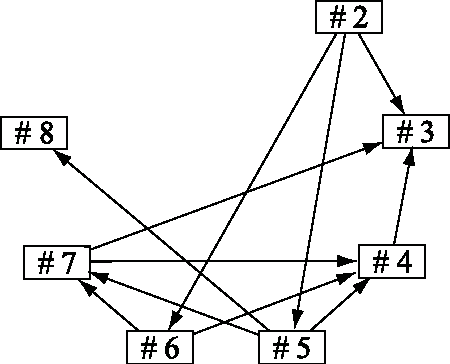
Abstract:Structural equation models and Bayesian networks have been widely used to analyze causal relations between continuous variables. In such frameworks, linear acyclic models are typically used to model the datagenerating process of variables. Recently, it was shown that use of non-Gaussianity identifies a causal ordering of variables in a linear acyclic model without using any prior knowledge on the network structure, which is not the case with conventional methods. However, existing estimation methods are based on iterative search algorithms and may not converge to a correct solution in a finite number of steps. In this paper, we propose a new direct method to estimate a causal ordering based on non-Gaussianity. In contrast to the previous methods, our algorithm requires no algorithmic parameters and is guaranteed to converge to the right solution within a small fixed number of steps if the data strictly follows the model.
Causal Discovery in a Binary Exclusive-or Skew Acyclic Model: BExSAM
Jan 22, 2014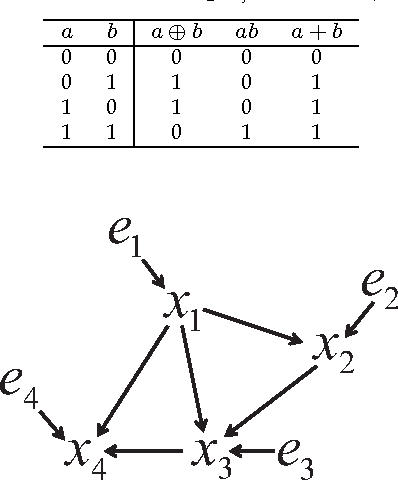
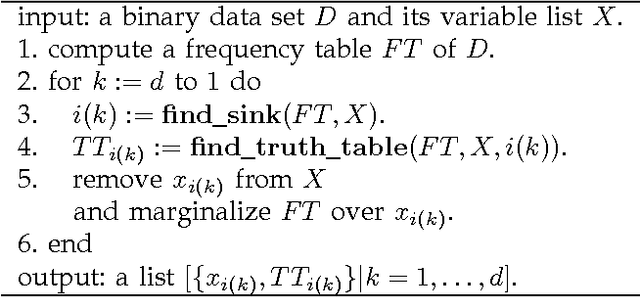
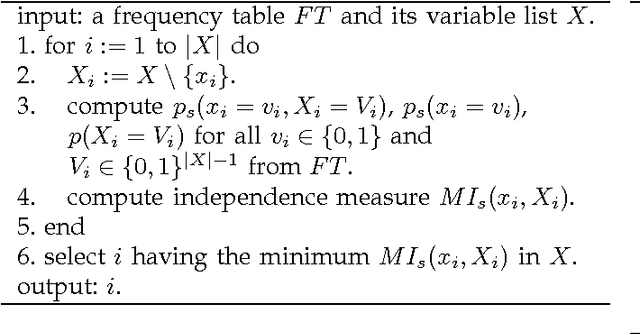

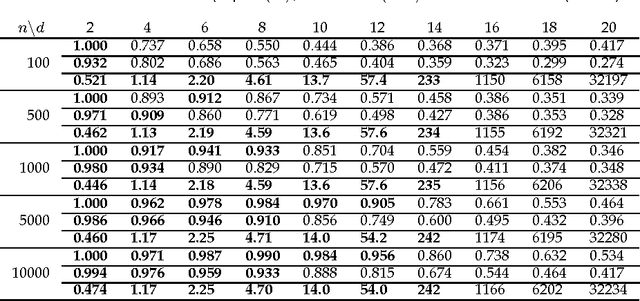

Abstract:Discovering causal relations among observed variables in a given data set is a major objective in studies of statistics and artificial intelligence. Recently, some techniques to discover a unique causal model have been explored based on non-Gaussianity of the observed data distribution. However, most of these are limited to continuous data. In this paper, we present a novel causal model for binary data and propose an efficient new approach to deriving the unique causal model governing a given binary data set under skew distributions of external binary noises. Experimental evaluation shows excellent performance for both artificial and real world data sets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge