Ying Bai
Vector Representations of Vessel Trees
Jun 11, 2025Abstract:We introduce a novel framework for learning vector representations of tree-structured geometric data focusing on 3D vascular networks. Our approach employs two sequentially trained Transformer-based autoencoders. In the first stage, the Vessel Autoencoder captures continuous geometric details of individual vessel segments by learning embeddings from sampled points along each curve. In the second stage, the Vessel Tree Autoencoder encodes the topology of the vascular network as a single vector representation, leveraging the segment-level embeddings from the first model. A recursive decoding process ensures that the reconstructed topology is a valid tree structure. Compared to 3D convolutional models, this proposed approach substantially lowers GPU memory requirements, facilitating large-scale training. Experimental results on a 2D synthetic tree dataset and a 3D coronary artery dataset demonstrate superior reconstruction fidelity, accurate topology preservation, and realistic interpolations in latent space. Our scalable framework, named VeTTA, offers precise, flexible, and topologically consistent modeling of anatomical tree structures in medical imaging.
Atlas-ISTN: Joint Segmentation, Registration and Atlas Construction with Image-and-Spatial Transformer Networks
Dec 18, 2020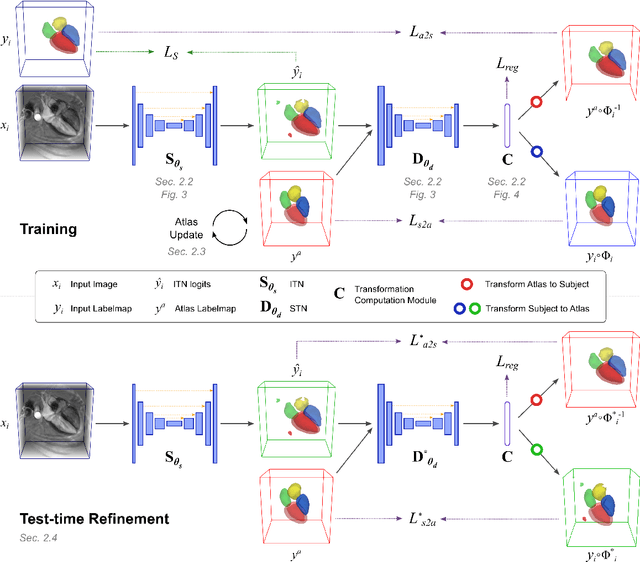
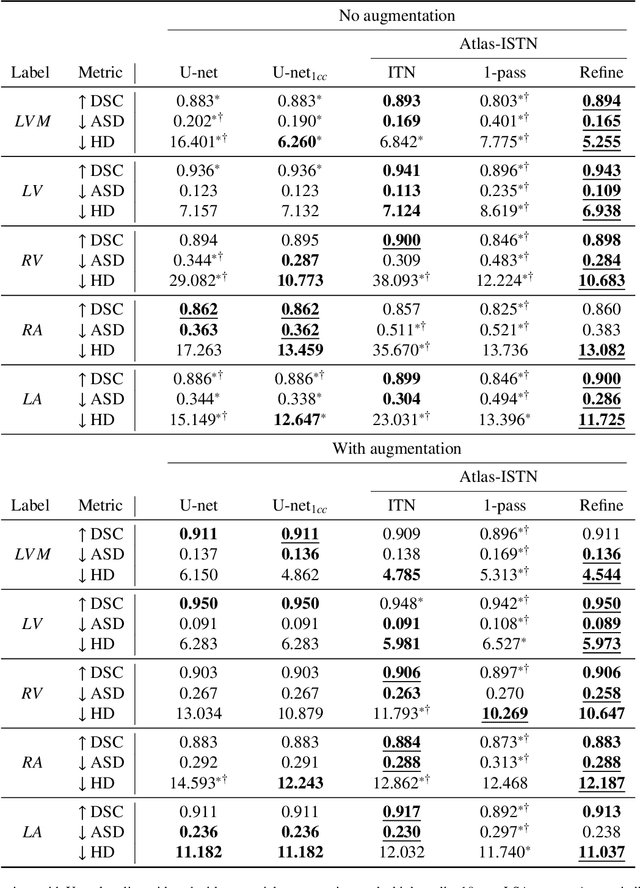
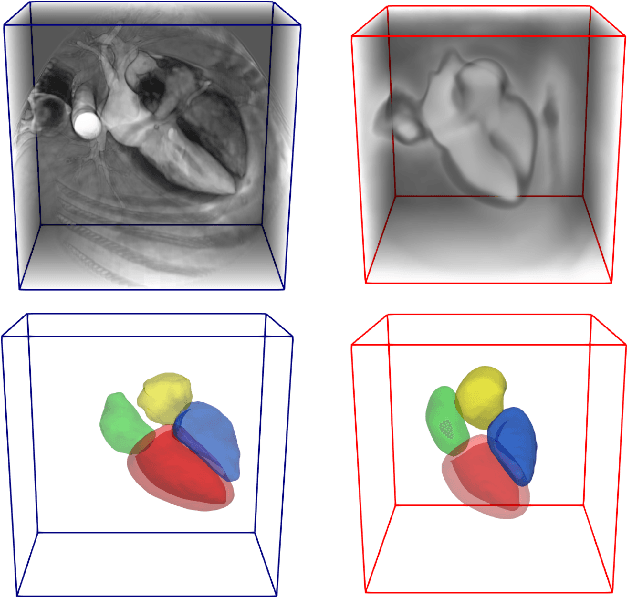
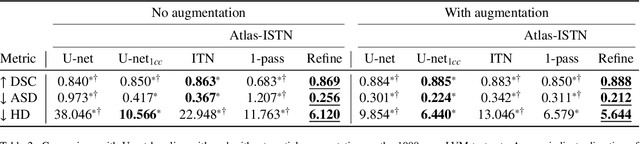
Abstract:Deep learning models for semantic segmentation are able to learn powerful representations for pixel-wise predictions, but are sensitive to noise at test time and do not guarantee a plausible topology. Image registration models on the other hand are able to warp known topologies to target images as a means of segmentation, but typically require large amounts of training data, and have not widely been benchmarked against pixel-wise segmentation models. We propose Atlas-ISTN, a framework that jointly learns segmentation and registration on 2D and 3D image data, and constructs a population-derived atlas in the process. Atlas-ISTN learns to segment multiple structures of interest and to register the constructed, topologically consistent atlas labelmap to an intermediate pixel-wise segmentation. Additionally, Atlas-ISTN allows for test time refinement of the model's parameters to optimize the alignment of the atlas labelmap to an intermediate pixel-wise segmentation. This process both mitigates for noise in the target image that can result in spurious pixel-wise predictions, as well as improves upon the one-pass prediction of the model. Benefits of the Atlas-ISTN framework are demonstrated qualitatively and quantitatively on 2D synthetic data and 3D cardiac computed tomography and brain magnetic resonance image data, out-performing both segmentation and registration baseline models. Atlas-ISTN also provides inter-subject correspondence of the structures of interest, enabling population-level shape and motion analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge