Winston Wang
DiffQRCoder: Diffusion-based Aesthetic QR Code Generation with Scanning Robustness Guided Iterative Refinement
Sep 10, 2024



Abstract:With the success of Diffusion Models for image generation, the technologies also have revolutionized the aesthetic Quick Response (QR) code generation. Despite significant improvements in visual attractiveness for the beautified codes, their scannabilities are usually sacrificed and thus hinder their practical uses in real-world scenarios. To address this issue, we propose a novel Diffusion-based QR Code generator (DiffQRCoder) to effectively craft both scannable and visually pleasing QR codes. The proposed approach introduces Scanning-Robust Perceptual Guidance (SRPG), a new diffusion guidance for Diffusion Models to guarantee the generated aesthetic codes to obey the ground-truth QR codes while maintaining their attractiveness during the denoising process. Additionally, we present another post-processing technique, Scanning Robust Manifold Projected Gradient Descent (SR-MPGD), to further enhance their scanning robustness through iterative latent space optimization. With extensive experiments, the results demonstrate that our approach not only outperforms other compared methods in Scanning Success Rate (SSR) with better or comparable CLIP aesthetic score (CLIP-aes.) but also significantly improves the SSR of the ControlNet-only approach from 60% to 99%. The subjective evaluation indicates that our approach achieves promising visual attractiveness to users as well. Finally, even with different scanning angles and the most rigorous error tolerance settings, our approach robustly achieves over 95% SSR, demonstrating its capability for real-world applications.
Diffusion-based Aesthetic QR Code Generation via Scanning-Robust Perceptual Guidance
Mar 23, 2024



Abstract:QR codes, prevalent in daily applications, lack visual appeal due to their conventional black-and-white design. Integrating aesthetics while maintaining scannability poses a challenge. In this paper, we introduce a novel diffusion-model-based aesthetic QR code generation pipeline, utilizing pre-trained ControlNet and guided iterative refinement via a novel classifier guidance (SRG) based on the proposed Scanning-Robust Loss (SRL) tailored with QR code mechanisms, which ensures both aesthetics and scannability. To further improve the scannability while preserving aesthetics, we propose a two-stage pipeline with Scanning-Robust Perceptual Guidance (SRPG). Moreover, we can further enhance the scannability of the generated QR code by post-processing it through the proposed Scanning-Robust Projected Gradient Descent (SRPGD) post-processing technique based on SRL with proven convergence. With extensive quantitative, qualitative, and subjective experiments, the results demonstrate that the proposed approach can generate diverse aesthetic QR codes with flexibility in detail. In addition, our pipelines outperforming existing models in terms of Scanning Success Rate (SSR) 86.67% (+40%) with comparable aesthetic scores. The pipeline combined with SRPGD further achieves 96.67% (+50%). Our code will be available https://github.com/jwliao1209/DiffQRCode.
Prediction of Small Molecule Kinase Inhibitors for Chemotherapy Using Deep Learning
Jun 30, 2019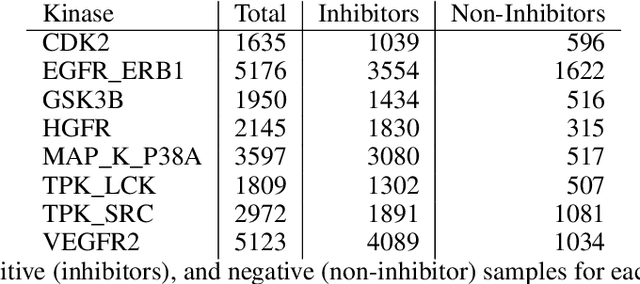
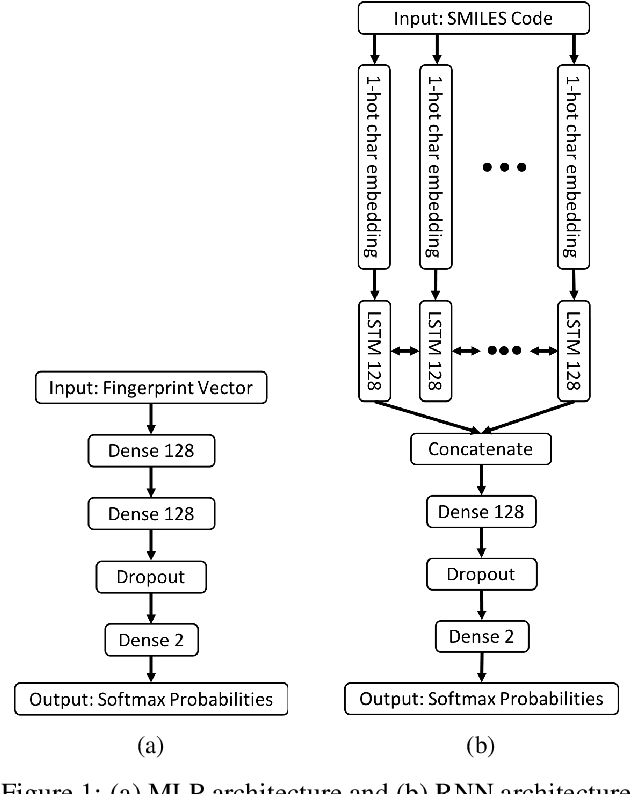
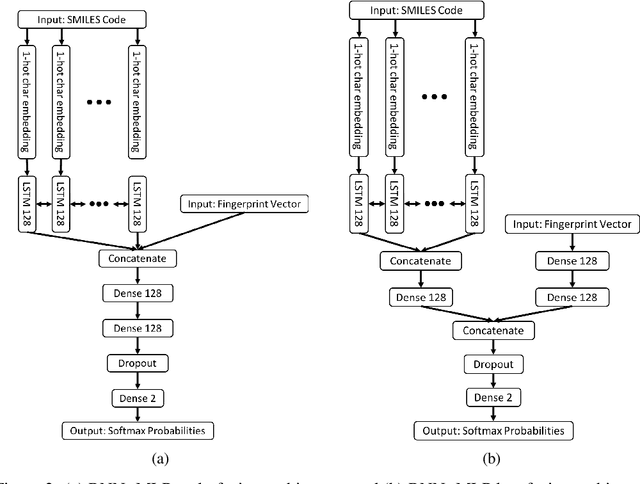
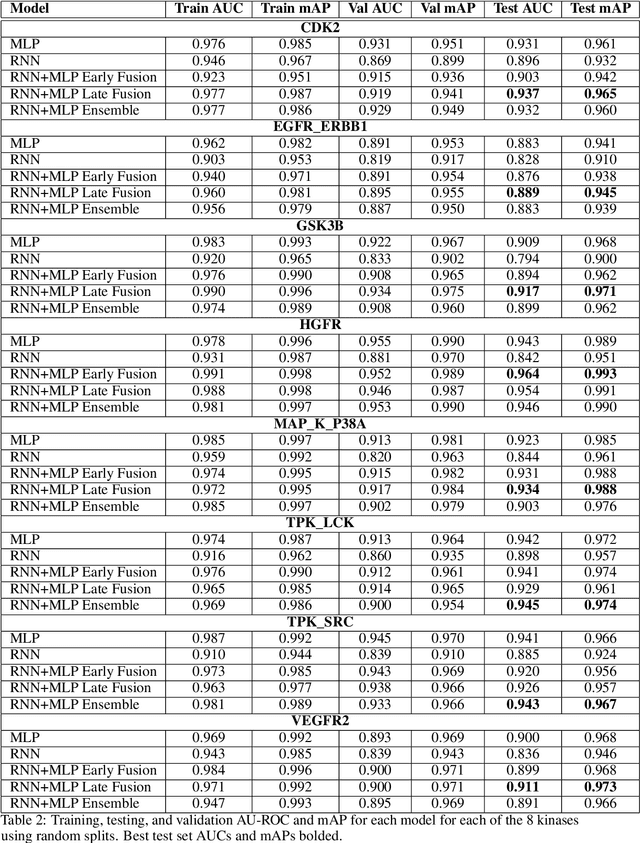
Abstract:The current state of cancer therapeutics has been moving away from one-size-fits-all cytotoxic chemotherapy, and towards a more individualized and specific approach involving the targeting of each tumor's genetic vulnerabilities. Different tumors, even of the same type, may be more reliant on certain cellular pathways more than others. With modern advancements in our understanding of cancer genome sequencing, these pathways can be discovered. Investigating each of the millions of possible small molecule inhibitors for each kinase in vitro, however, would be extremely expensive and time consuming. This project focuses on predicting the inhibition activity of small molecules targeting 8 different kinases using multiple deep learning models. We trained fingerprint-based MLPs and simplified molecular-input line-entry specification (SMILES)-based recurrent neural networks (RNNs) and molecular graph convolutional networks (GCNs) to accurately predict inhibitory activity targeting these 8 kinases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge