Venkataraman Subramanian
A comprehensive multimodal dataset and benchmark for ulcerative colitis scoring in endoscopy
Mar 15, 2026Abstract:Ulcerative colitis (UC) is a chronic mucosal inflammatory condition that places patients at increased risk of colorectal cancer. Colonoscopic surveillance remains the gold standard for assessing disease activity, and reporting typically relies on standardised endoscopic scoring metrics. The most widely used is the Mayo Endoscopic Score (MES), with some centres also adopting the Ulcerative Colitis Endoscopic Index of Severity (UCEIS). Both are descriptive assessments of mucosal inflammation (MES: 0 to 3; UCEIS: 0 to 8), where higher values indicate more severe disease. However, computational methods for automatically predicting these scores remain limited, largely due to the lack of publicly available expert-annotated datasets and the absence of robust benchmarking. There is also a significant research gap in generating clinically meaningful descriptions of UC images, despite image captioning being a well-established computer vision task. Variability in endoscopic systems and procedural workflows across centres further highlights the need for multi-centre datasets to ensure algorithmic robustness and generalisability. In this work, we introduce a curated multi-centre, multi-resolution dataset that includes expert-validated MES and UCEIS labels, alongside detailed clinical descriptions. To our knowledge, this is the first comprehensive dataset that combines dual scoring metrics for classification tasks with expert-generated captions describing mucosal appearance and clinically accepted reasoning for image captioning. This resource opens new opportunities for developing clinically meaningful multimodal algorithms. In addition to the dataset, we also provide benchmarking using convolutional neural networks, vision transformers, hybrid models, and widely used multimodal vision-language captioning algorithms.
Multi-task learning with cross-task consistency for improved depth estimation in colonoscopy
Nov 30, 2023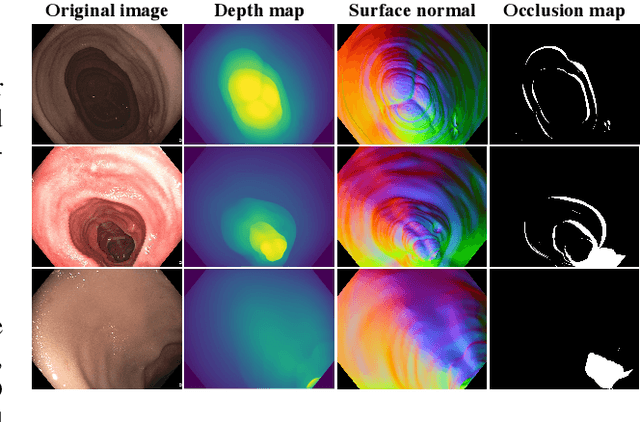
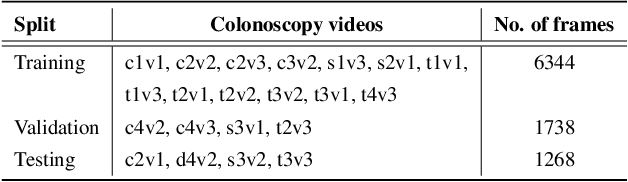
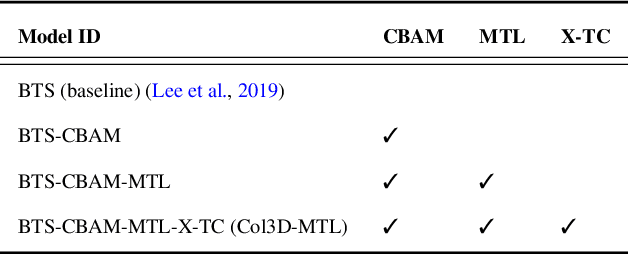

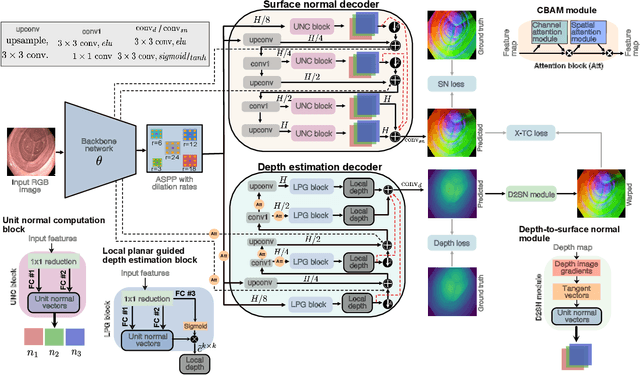

Abstract:Colonoscopy screening is the gold standard procedure for assessing abnormalities in the colon and rectum, such as ulcers and cancerous polyps. Measuring the abnormal mucosal area and its 3D reconstruction can help quantify the surveyed area and objectively evaluate disease burden. However, due to the complex topology of these organs and variable physical conditions, for example, lighting, large homogeneous texture, and image modality estimating distance from the camera aka depth) is highly challenging. Moreover, most colonoscopic video acquisition is monocular, making the depth estimation a non-trivial problem. While methods in computer vision for depth estimation have been proposed and advanced on natural scene datasets, the efficacy of these techniques has not been widely quantified on colonoscopy datasets. As the colonic mucosa has several low-texture regions that are not well pronounced, learning representations from an auxiliary task can improve salient feature extraction, allowing estimation of accurate camera depths. In this work, we propose to develop a novel multi-task learning (MTL) approach with a shared encoder and two decoders, namely a surface normal decoder and a depth estimator decoder. Our depth estimator incorporates attention mechanisms to enhance global context awareness. We leverage the surface normal prediction to improve geometric feature extraction. Also, we apply a cross-task consistency loss among the two geometrically related tasks, surface normal and camera depth. We demonstrate an improvement of 14.17% on relative error and 10.4% improvement on $\delta_{1}$ accuracy over the most accurate baseline state-of-the-art BTS approach. All experiments are conducted on a recently released C3VD dataset; thus, we provide a first benchmark of state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge