Vassilis N. Ioannidis
Unveiling Anomalous Edges and Nominal Connectivity of Attributed Networks
Apr 17, 2021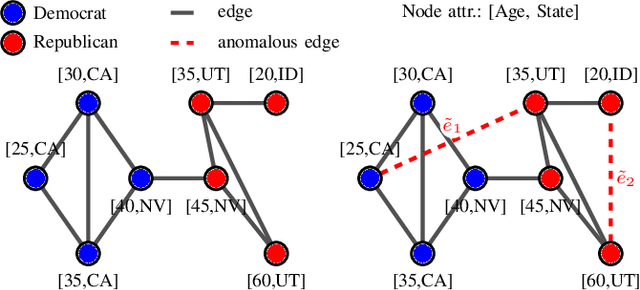
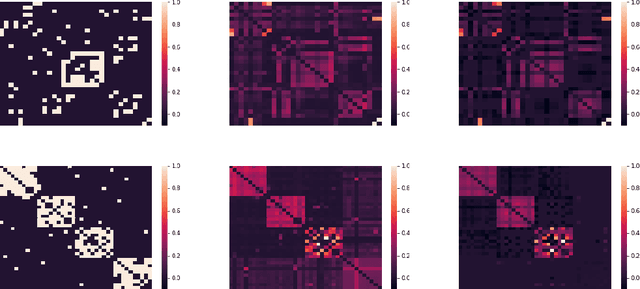
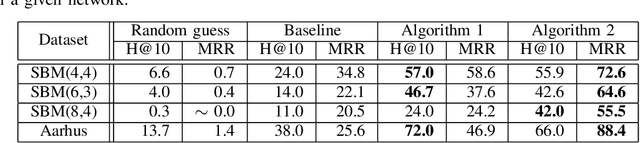
Abstract:Uncovering anomalies in attributed networks has recently gained popularity due to its importance in unveiling outliers and flagging adversarial behavior in a gamut of data and network science applications including {the Internet of Things (IoT)}, finance, security, to list a few. The present work deals with uncovering anomalous edges in attributed graphs using two distinct formulations with complementary strengths, which can be easily distributed, and hence efficient. The first relies on decomposing the graph data matrix into low rank plus sparse components to markedly improve performance. The second broadens the scope of the first by performing robust recovery of the unperturbed graph, which enhances the anomaly identification performance. The novel methods not only capture anomalous edges linking nodes of different communities, but also spurious connections between any two nodes with different features. Experiments conducted on real and synthetic data corroborate the effectiveness of both methods in the anomaly identification task.
COVID-19 Knowledge Graph: Accelerating Information Retrieval and Discovery for Scientific Literature
Jul 24, 2020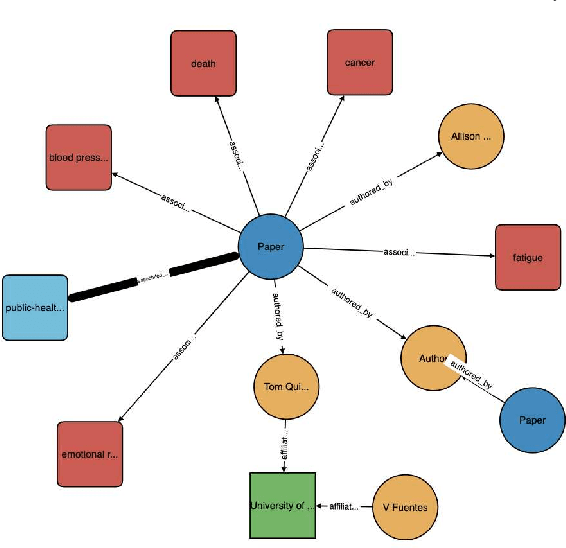
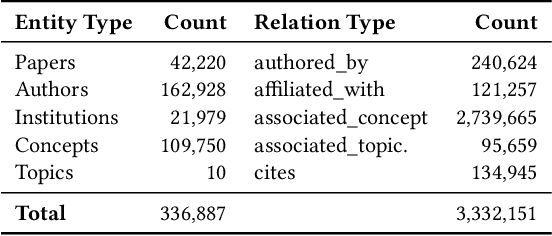
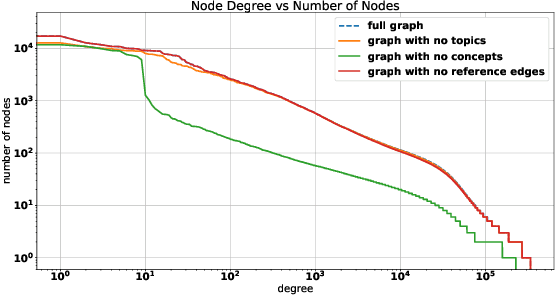

Abstract:The coronavirus disease (COVID-19) has claimed the lives of over 350,000 people and infected more than 6 million people worldwide. Several search engines have surfaced to provide researchers with additional tools to find and retrieve information from the rapidly growing corpora on COVID-19. These engines lack extraction and visualization tools necessary to retrieve and interpret complex relations inherent to scientific literature. Moreover, because these engines mainly rely upon semantic information, their ability to capture complex global relationships across documents is limited, which reduces the quality of similarity-based article recommendations for users. In this work, we present the COVID-19 Knowledge Graph (CKG), a heterogeneous graph for extracting and visualizing complex relationships between COVID-19 scientific articles. The CKG combines semantic information with document topological information for the application of similar document retrieval. The CKG is constructed using the latent schema of the data, and then enriched with biomedical entity information extracted from the unstructured text of articles using scalable AWS technologies to form relations in the graph. Finally, we propose a document similarity engine that leverages low-dimensional graph embeddings from the CKG with semantic embeddings for similar article retrieval. Analysis demonstrates the quality of relationships in the CKG and shows that it can be used to uncover meaningful information in COVID-19 scientific articles. The CKG helps power www.cord19.aws and is publicly available.
PanRep: Universal node embeddings for heterogeneous graphs
Jul 20, 2020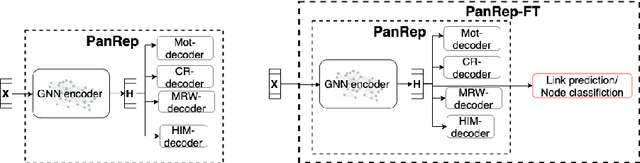
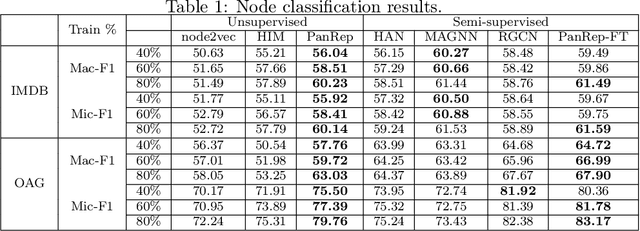
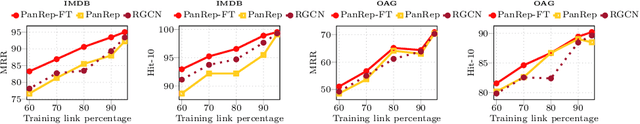
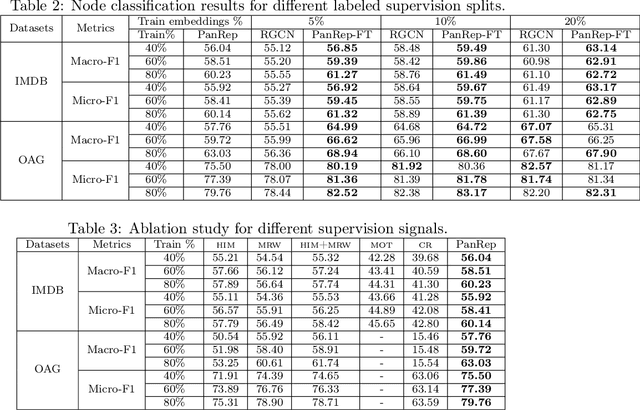
Abstract:Learning unsupervised node embeddings facilitates several downstream tasks such as node classification and link prediction. A node embedding is universal if it is designed to be used by and benefit various downstream tasks. This work introduces PanRep, a graph neural network (GNN) model, for unsupervised learning of universal node representations for heterogenous graphs. PanRep consists of a GNN encoder that obtains node embeddings and four decoders, each capturing different topological and node feature properties. Abiding to these properties the novel unsupervised framework learns universal embeddings applicable to different downstream tasks. PanRep can be furthered fine-tuned to account for possible limited labels. In this operational setting PanRep is considered as a pretrained model for extracting node embeddings of heterogenous graph data. PanRep outperforms all unsupervised and certain supervised methods in node classification and link prediction, especially when the labeled data for the supervised methods is small. PanRep-FT (with fine-tuning) outperforms all other supervised approaches, which corroborates the merits of pretraining models. Finally, we apply PanRep-FT for discovering novel drugs for Covid-19. We showcase the advantage of universal embeddings in drug repurposing and identify several drugs used in clinical trials as possible drug candidates.
Few-shot link prediction via graph neural networks for Covid-19 drug-repurposing
Jul 20, 2020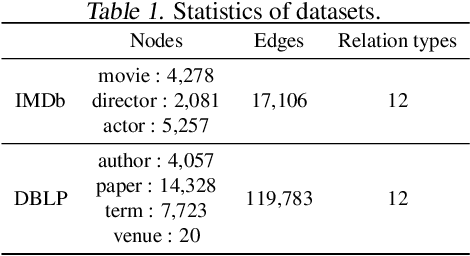
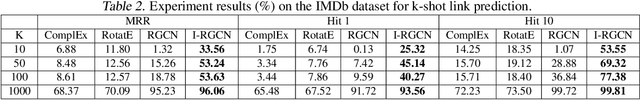
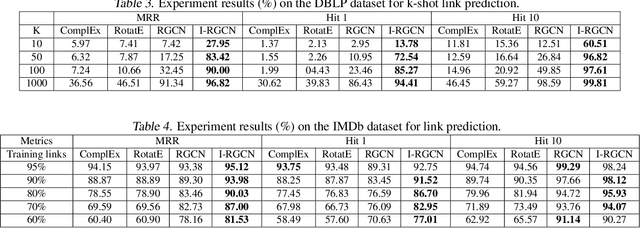
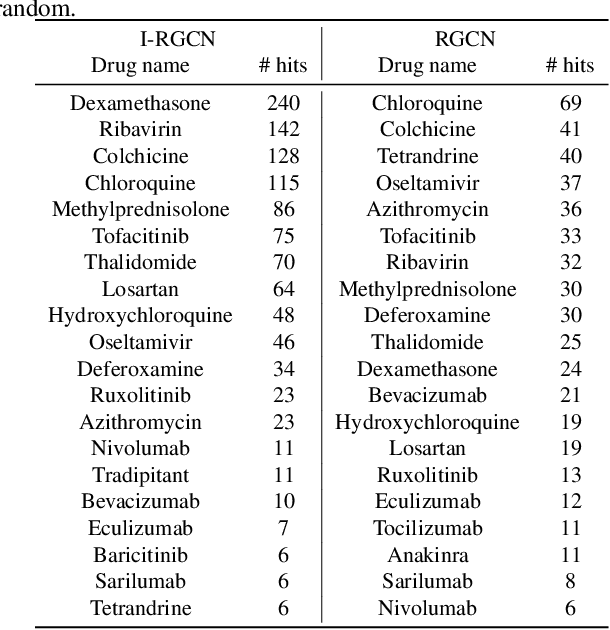
Abstract:Predicting interactions among heterogenous graph structured data has numerous applications such as knowledge graph completion, recommendation systems and drug discovery. Often times, the links to be predicted belong to rare types such as the case in repurposing drugs for novel diseases. This motivates the task of few-shot link prediction. Typically, GCNs are ill-equipped in learning such rare link types since the relation embedding is not learned in an inductive fashion. This paper proposes an inductive RGCN for learning informative relation embeddings even in the few-shot learning regime. The proposed inductive model significantly outperforms the RGCN and state-of-the-art KGE models in few-shot learning tasks. Furthermore, we apply our method on the drug-repurposing knowledge graph (DRKG) for discovering drugs for Covid-19. We pose the drug discovery task as link prediction and learn embeddings for the biological entities that partake in the DRKG. Our initial results corroborate that several drugs used in clinical trials were identified as possible drug candidates. The method in this paper are implemented using the efficient deep graph learning (DGL)
Tensor Graph Convolutional Networks for Multi-relational and Robust Learning
Mar 15, 2020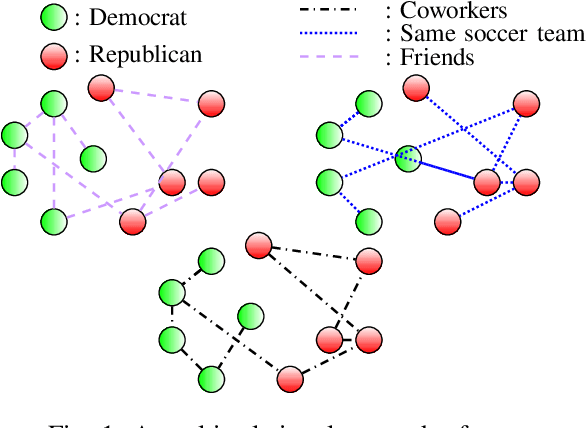
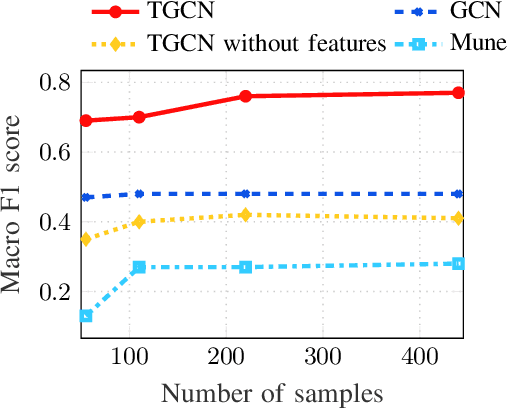
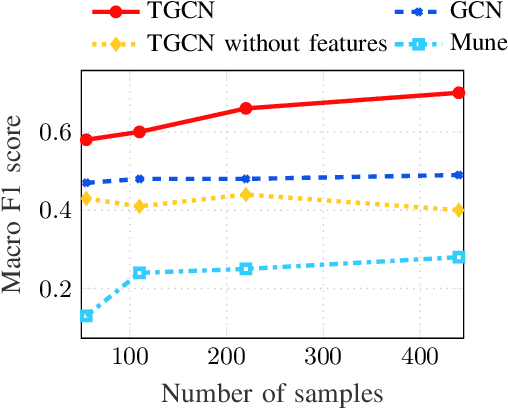
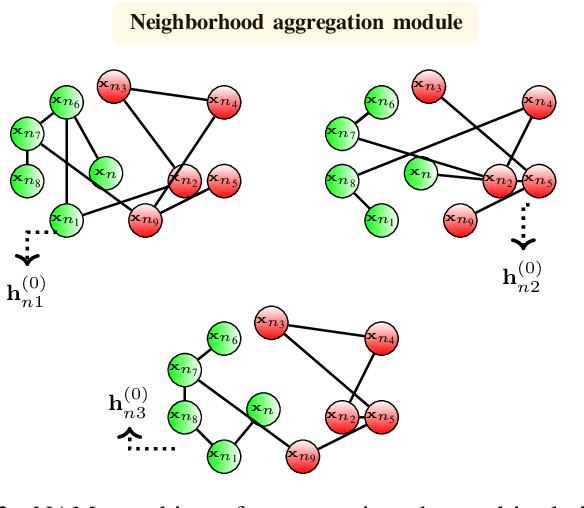
Abstract:The era of "data deluge" has sparked renewed interest in graph-based learning methods and their widespread applications ranging from sociology and biology to transportation and communications. In this context of graph-aware methods, the present paper introduces a tensor-graph convolutional network (TGCN) for scalable semi-supervised learning (SSL) from data associated with a collection of graphs, that are represented by a tensor. Key aspects of the novel TGCN architecture are the dynamic adaptation to different relations in the tensor graph via learnable weights, and the consideration of graph-based regularizers to promote smoothness and alleviate over-parameterization. The ultimate goal is to design a powerful learning architecture able to: discover complex and highly nonlinear data associations, combine (and select) multiple types of relations, scale gracefully with the graph size, and remain robust to perturbations on the graph edges. The proposed architecture is relevant not only in applications where the nodes are naturally involved in different relations (e.g., a multi-relational graph capturing family, friendship and work relations in a social network), but also in robust learning setups where the graph entails a certain level of uncertainty, and the different tensor slabs correspond to different versions (realizations) of the nominal graph. Numerical tests showcase that the proposed architecture achieves markedly improved performance relative to standard GCNs, copes with state-of-the-art adversarial attacks, and leads to remarkable SSL performance over protein-to-protein interaction networks.
Efficient and Stable Graph Scattering Transforms via Pruning
Jan 27, 2020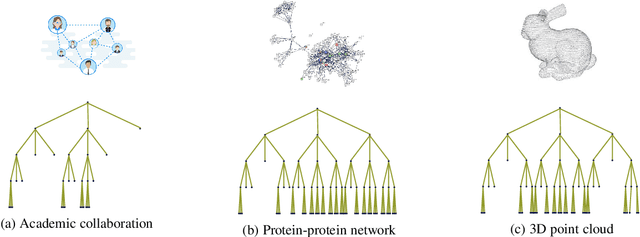
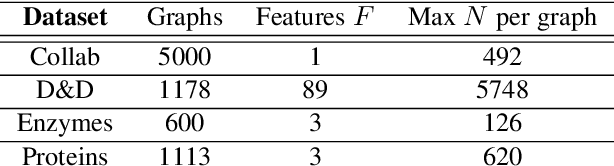
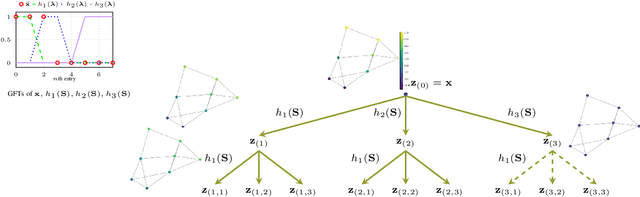
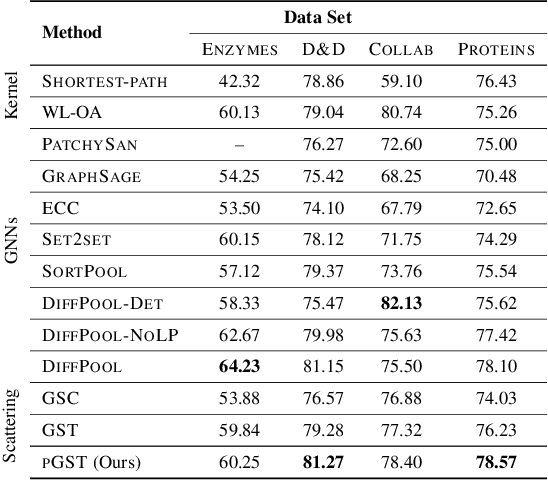
Abstract:Graph convolutional networks (GCNs) have well-documented performance in various graph learning tasks, but their analysis is still at its infancy. Graph scattering transforms (GSTs) offer training-free deep GCN models that extract features from graph data, and are amenable to generalization and stability analyses. The price paid by GSTs is exponential complexity in space and time that increases with the number of layers. This discourages deployment of GSTs when a deep architecture is needed. The present work addresses the complexity limitation of GSTs by introducing an efficient so-termed pruned (p)GST approach. The resultant pruning algorithm is guided by a graph-spectrum-inspired criterion, and retains informative scattering features on-the-fly while bypassing the exponential complexity associated with GSTs. Stability of the novel pGSTs is also established when the input graph data or the network structure are perturbed. Furthermore, the sensitivity of pGST to random and localized signal perturbations is investigated analytically and experimentally. Numerical tests showcase that pGST performs comparably to the baseline GST at considerable computational savings. Furthermore, pGST achieves comparable performance to state-of-the-art GCNs in graph and 3D point cloud classification tasks. Upon analyzing the pGST pruning patterns, it is shown that graph data in different domains call for different network architectures, and that the pruning algorithm may be employed to guide the design choices for contemporary GCNs.
Edge Dithering for Robust Adaptive Graph Convolutional Networks
Oct 21, 2019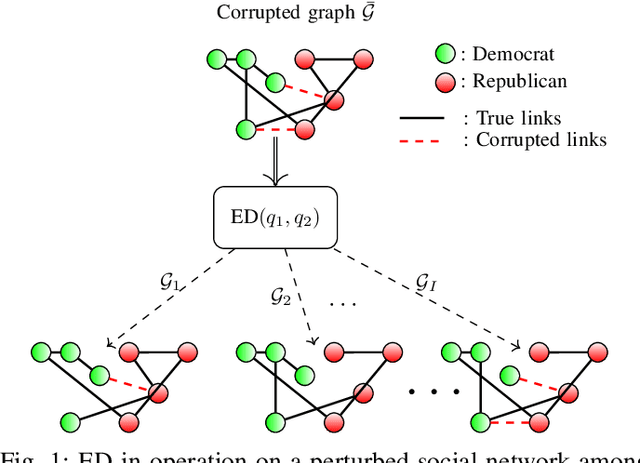
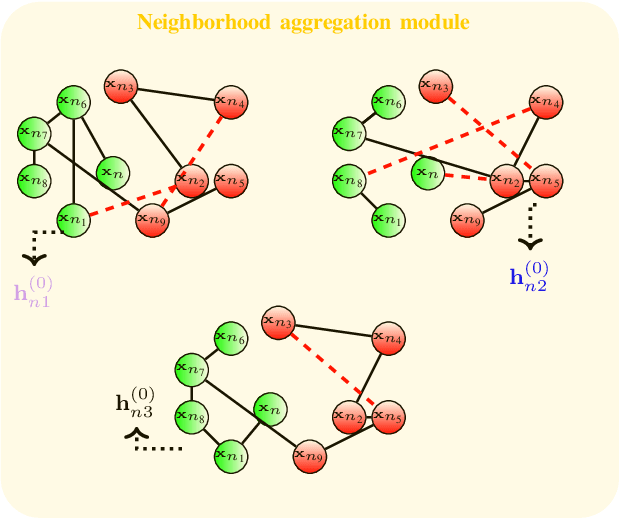
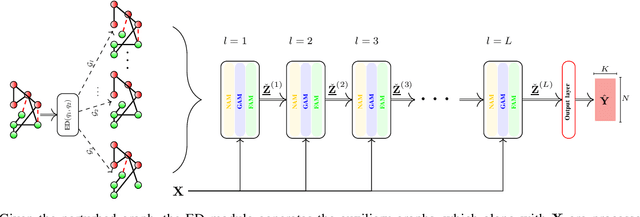
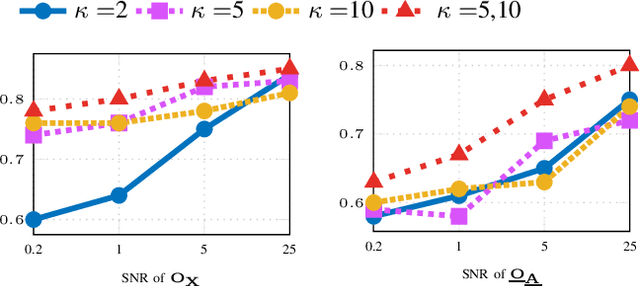
Abstract:Graph convolutional networks (GCNs) are vulnerable to perturbations of the graph structure that are either random, or, adversarially designed. The perturbed links modify the graph neighborhoods, which critically affects the performance of GCNs in semi-supervised learning (SSL) tasks. Aiming at robustifying GCNs conditioned on the perturbed graph, the present paper generates multiple auxiliary graphs, each having its binary 0-1 edge weights flip values with probabilities designed to enhance robustness. The resultant edge-dithered auxiliary graphs are leveraged by an adaptive (A)GCN that performs SSL. Robustness is enabled through learnable graph-combining weights along with suitable regularizers. Relative to GCN, the novel AGCN achieves markedly improved performance in tests with noisy inputs, graph perturbations, and state-of-the-art adversarial attacks. Further experiments with protein interaction networks showcase the competitive performance of AGCN for SSL over multiple graphs.
GraphSAC: Detecting anomalies in large-scale graphs
Oct 21, 2019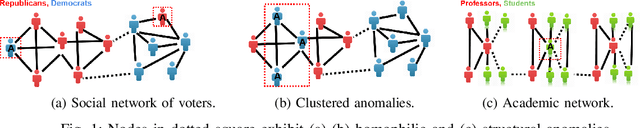
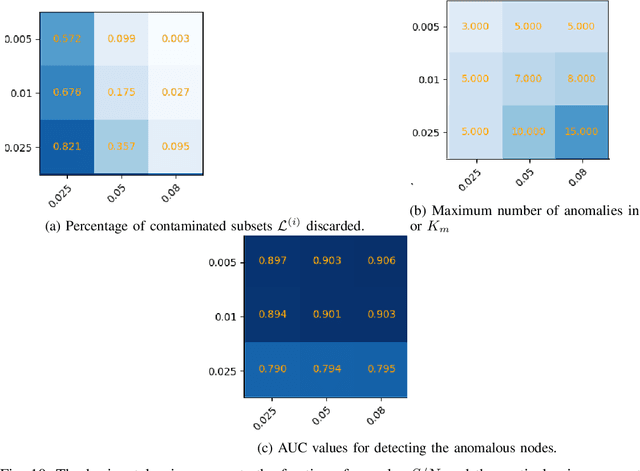
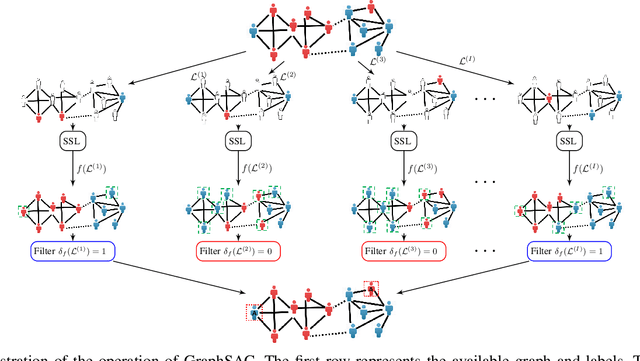
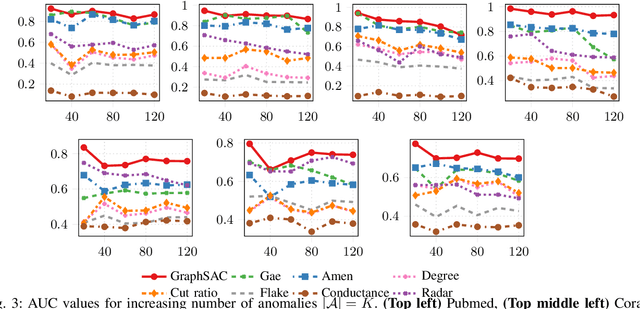
Abstract:A graph-based sampling and consensus (GraphSAC) approach is introduced to effectively detect anomalous nodes in large-scale graphs. Existing approaches rely on connectivity and attributes of all nodes to assign an anomaly score per node. However, nodal attributes and network links might be compromised by adversaries, rendering these holistic approaches vulnerable. Alleviating this limitation, GraphSAC randomly draws subsets of nodes, and relies on graph-aware criteria to judiciously filter out sets contaminated by anomalous nodes, before employing a semi-supervised learning (SSL) module to estimate nominal label distributions per node. These learned nominal distributions are minimally affected by the anomalous nodes, and hence can be directly adopted for anomaly detection. Rigorous analysis provides performance guarantees for GraphSAC, by bounding the required number of draws. The per-draw complexity grows linearly with the number of edges, which implies efficient SSL, while draws can be run in parallel, thereby ensuring scalability to large graphs. GraphSAC is tested under different anomaly generation models based on random walks, clustered anomalies, as well as contemporary adversarial attacks for graph data. Experiments with real-world graphs showcase the advantage of GraphSAC relative to state-of-the-art alternatives.
A Recurrent Graph Neural Network for Multi-Relational Data
Nov 19, 2018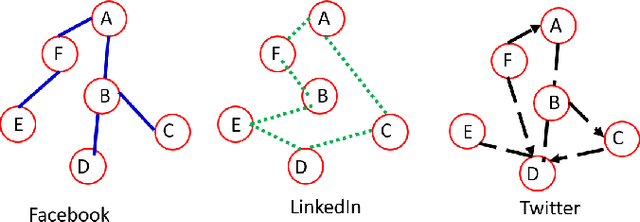
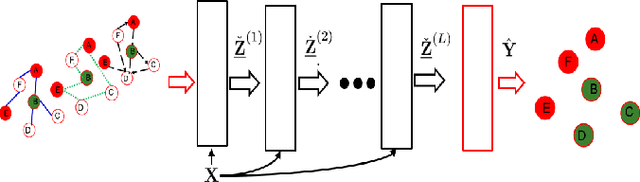
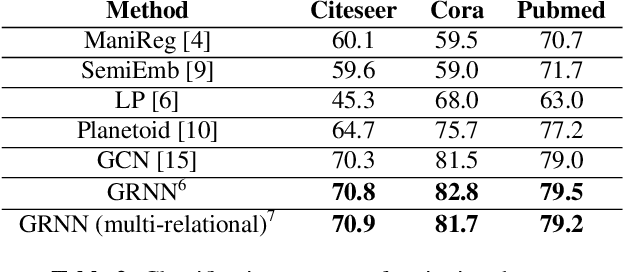
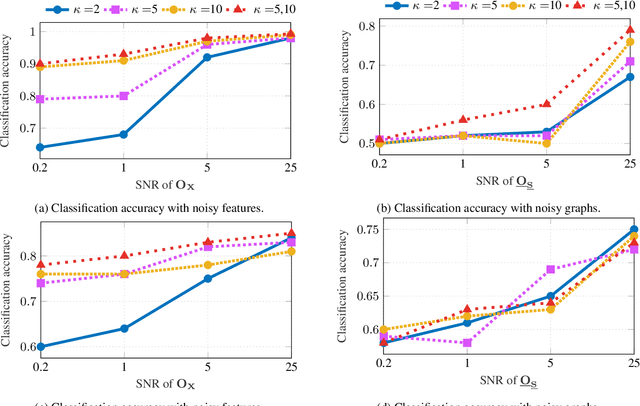
Abstract:The era of data deluge has sparked the interest in graph-based learning methods in a number of disciplines such as sociology, biology, neuroscience, or engineering. In this paper, we introduce a graph recurrent neural network (GRNN) for scalable semi-supervised learning from multi-relational data. Key aspects of the novel GRNN architecture are the use of multi-relational graphs, the dynamic adaptation to the different relations via learnable weights, and the consideration of graph-based regularizers to promote smoothness and alleviate over-parametrization. Our ultimate goal is to design a powerful learning architecture able to: discover complex and highly non-linear data associations, combine (and select) multiple types of relations, and scale gracefully with respect to the size of the graph. Numerical tests with real data sets corroborate the design goals and illustrate the performance gains relative to competing alternatives.
Coupled Graphs and Tensor Factorization for Recommender Systems and Community Detection
Sep 22, 2018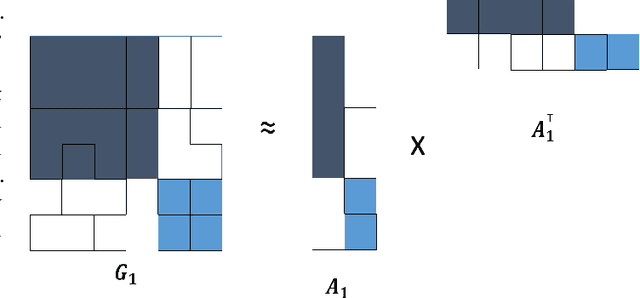
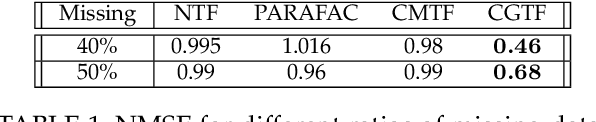
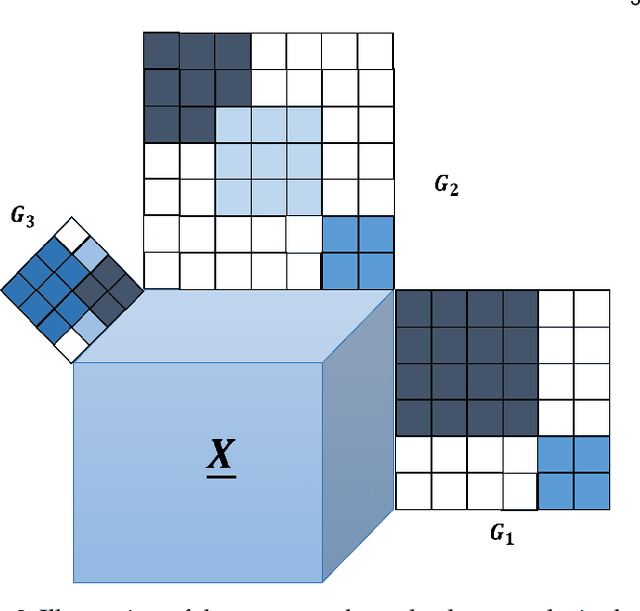
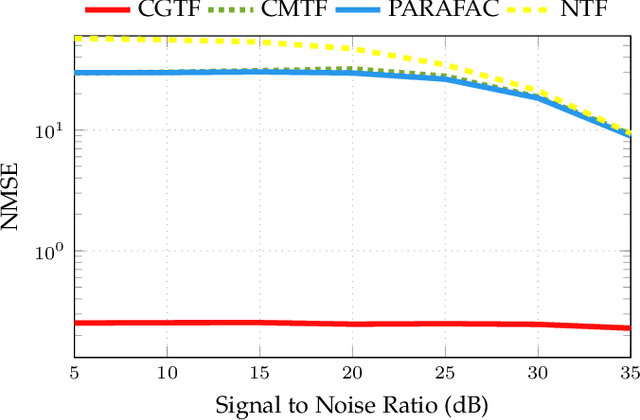
Abstract:Joint analysis of data from multiple information repositories facilitates uncovering the underlying structure in heterogeneous datasets. Single and coupled matrix-tensor factorization (CMTF) has been widely used in this context for imputation-based recommendation from ratings, social network, and other user-item data. When this side information is in the form of item-item correlation matrices or graphs, existing CMTF algorithms may fall short. Alleviating current limitations, we introduce a novel model coined coupled graph-tensor factorization (CGTF) that judiciously accounts for graph-related side information. The CGTF model has the potential to overcome practical challenges, such as missing slabs from the tensor and/or missing rows/columns from the correlation matrices. A novel alternating direction method of multipliers (ADMM) is also developed that recovers the nonnegative factors of CGTF. Our algorithm enjoys closed-form updates that result in reduced computational complexity and allow for convergence claims. A novel direction is further explored by employing the interpretable factors to detect graph communities having the tensor as side information. The resulting community detection approach is successful even when some links in the graphs are missing. Results with real data sets corroborate the merits of the proposed methods relative to state-of-the-art competing factorization techniques in providing recommendations and detecting communities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge