Varun Jog
Graph-Based Ascent Algorithms for Function Maximization
Feb 13, 2018
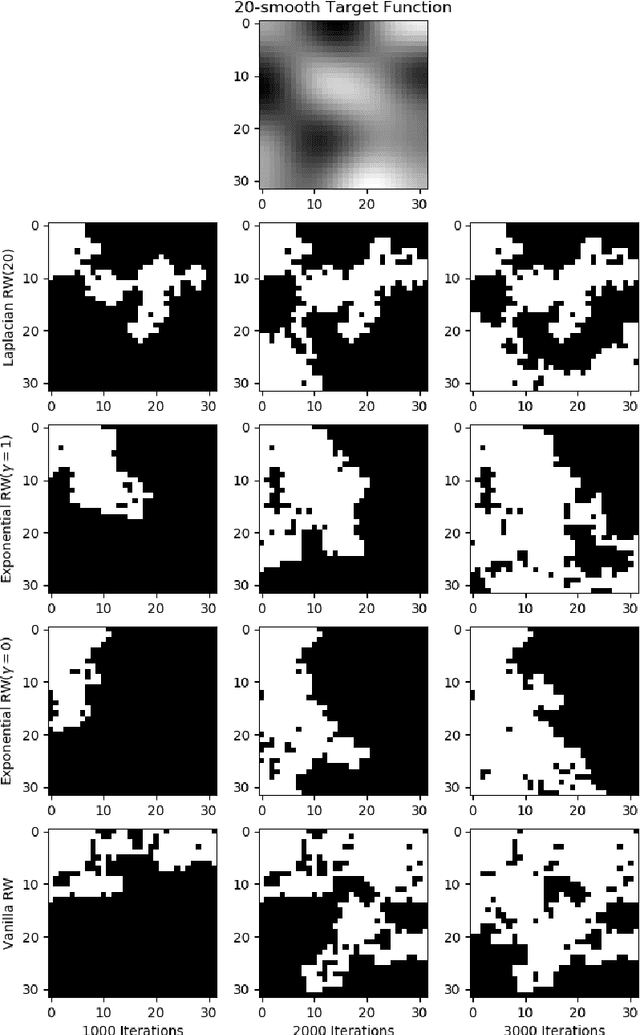
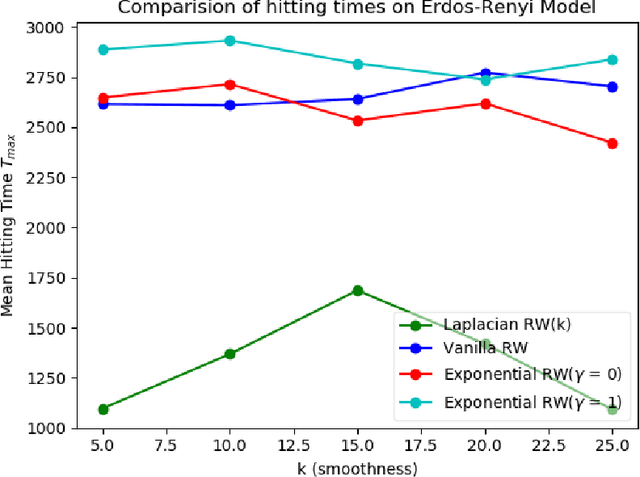
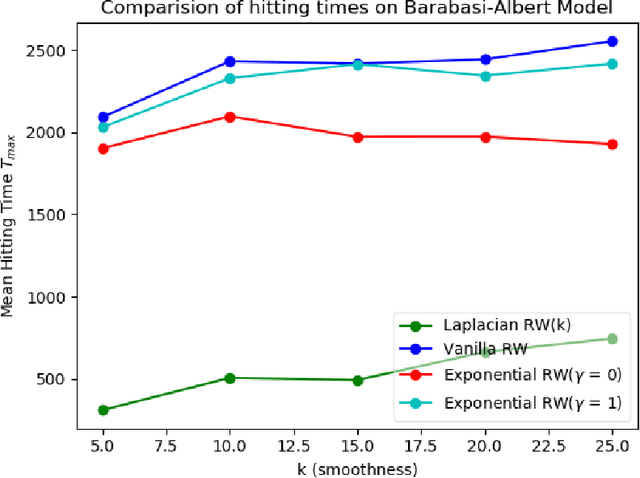
Abstract:We study the problem of finding the maximum of a function defined on the nodes of a connected graph. The goal is to identify a node where the function obtains its maximum. We focus on local iterative algorithms, which traverse the nodes of the graph along a path, and the next iterate is chosen from the neighbors of the current iterate with probability distribution determined by the function values at the current iterate and its neighbors. We study two algorithms corresponding to a Metropolis-Hastings random walk with different transition kernels: (i) The first algorithm is an exponentially weighted random walk governed by a parameter $\gamma$. (ii) The second algorithm is defined with respect to the graph Laplacian and a smoothness parameter $k$. We derive convergence rates for the two algorithms in terms of total variation distance and hitting times. We also provide simulations showing the relative convergence rates of our algorithms in comparison to an unbiased random walk, as a function of the smoothness of the graph function. Our algorithms may be categorized as a new class of "descent-based" methods for function maximization on the nodes of a graph.
Generalization Error Bounds for Noisy, Iterative Algorithms
Jan 12, 2018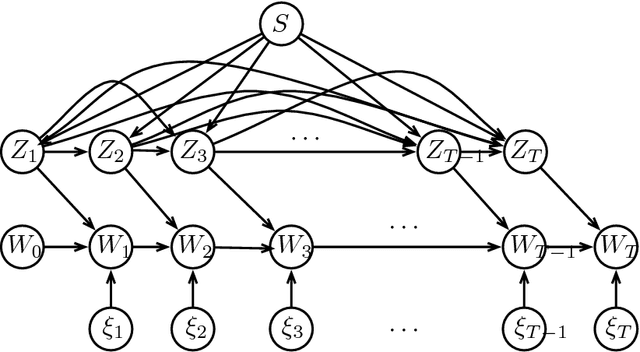
Abstract:In statistical learning theory, generalization error is used to quantify the degree to which a supervised machine learning algorithm may overfit to training data. Recent work [Xu and Raginsky (2017)] has established a bound on the generalization error of empirical risk minimization based on the mutual information $I(S;W)$ between the algorithm input $S$ and the algorithm output $W$, when the loss function is sub-Gaussian. We leverage these results to derive generalization error bounds for a broad class of iterative algorithms that are characterized by bounded, noisy updates with Markovian structure. Our bounds are very general and are applicable to numerous settings of interest, including stochastic gradient Langevin dynamics (SGLD) and variants of the stochastic gradient Hamiltonian Monte Carlo (SGHMC) algorithm. Furthermore, our error bounds hold for any output function computed over the path of iterates, including the last iterate of the algorithm or the average of subsets of iterates, and also allow for non-uniform sampling of data in successive updates of the algorithm.
Computationally Efficient Influence Maximization in Stochastic and Adversarial Models: Algorithms and Analysis
Nov 01, 2016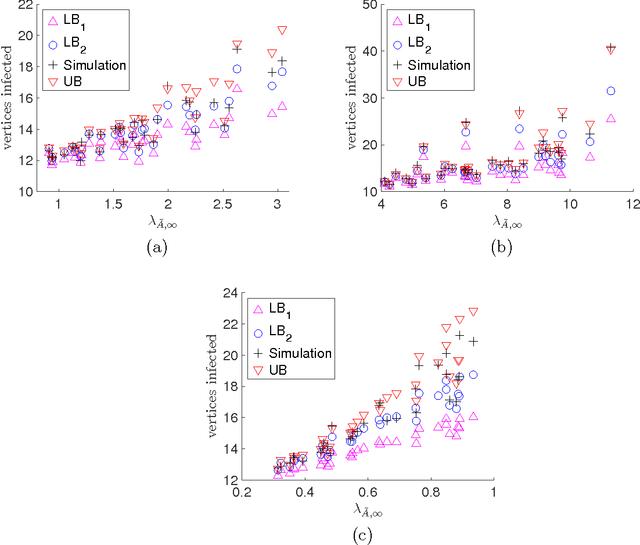
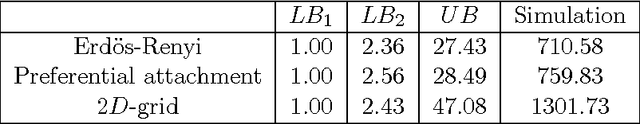
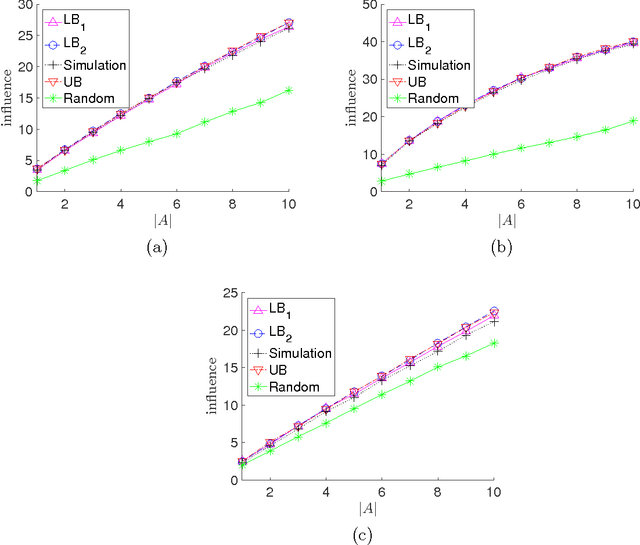
Abstract:We consider the problem of influence maximization in fixed networks, for both stochastic and adversarial contagion models. The common goal is to select a subset of nodes of a specified size to infect so that the number of infected nodes at the conclusion of the epidemic is as large as possible. In the stochastic setting, the epidemic spreads according to a general triggering model, which includes the popular linear threshold and independent cascade models. We establish upper and lower bounds for the influence of an initial subset of nodes in the network, where the influence is defined as the expected number of infected nodes. Although the problem of exact influence computation is NP-hard in general, our bounds may be evaluated efficiently, leading to scalable algorithms for influence maximization with rigorous theoretical guarantees. In the adversarial spreading setting, an adversary is allowed to specify the edges through which contagion may spread, and the player chooses sets of nodes to infect in successive rounds. Both the adversary and player may behave stochastically, but we limit the adversary to strategies that are oblivious of the player's actions. We establish upper and lower bounds on the minimax pseudo-regret in both undirected and directed networks.
On model misspecification and KL separation for Gaussian graphical models
Apr 03, 2015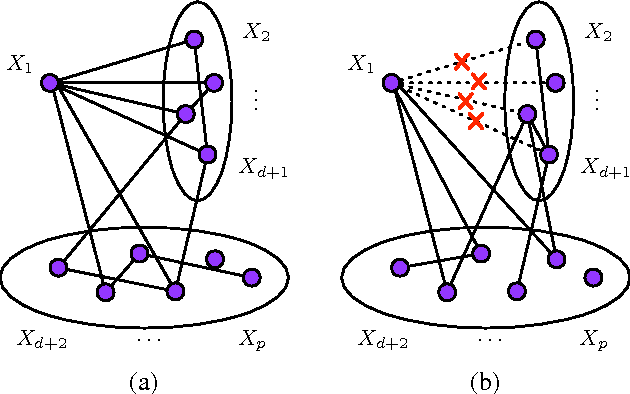
Abstract:We establish bounds on the KL divergence between two multivariate Gaussian distributions in terms of the Hamming distance between the edge sets of the corresponding graphical models. We show that the KL divergence is bounded below by a constant when the graphs differ by at least one edge; this is essentially the tightest possible bound, since classes of graphs exist for which the edge discrepancy increases but the KL divergence remains bounded above by a constant. As a natural corollary to our KL lower bound, we also establish a sample size requirement for correct model selection via maximum likelihood estimation. Our results rigorize the notion that it is essential to estimate the edge structure of a Gaussian graphical model accurately in order to approximate the true distribution to close precision.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge