Uzair Shah
GraPHFormer: A Multimodal Graph Persistent Homology Transformer for the Analysis of Neuroscience Morphologies
Mar 21, 2026Abstract:Neuronal morphology encodes critical information about circuit function, development, and disease, yet current methods analyze topology or graph structure in isolation. We introduce GraPHFormer, a multimodal architecture that unifies these complementary views through CLIP-style contrastive learning. Our vision branch processes a novel three-channel persistence image encoding unweighted, persistence-weighted, and radius-weighted topological densities via DINOv2-ViT-S. In parallel, a TreeLSTM encoder captures geometric and radial attributes from skeleton graphs. Both project to a shared embedding space trained with symmetric InfoNCE loss, augmented by persistence-space transformations that preserve topological semantics. Evaluated on six benchmarks (BIL-6, ACT-4, JML-4, N7, M1-Cell, M1-REG) spanning self-supervised and supervised settings, GraPHFormer achieves state-of-the-art performance on five benchmarks, significantly outperforming topology-only, graph-only, and morphometrics baselines. We demonstrate practical utility by discriminating glial morphologies across cortical regions and species, and detecting signatures of developmental and degenerative processes. Code: https://github.com/Uzshah/GraPHFormer
SAM4EM: Efficient memory-based two stage prompt-free segment anything model adapter for complex 3D neuroscience electron microscopy stacks
Apr 30, 2025Abstract:We present SAM4EM, a novel approach for 3D segmentation of complex neural structures in electron microscopy (EM) data by leveraging the Segment Anything Model (SAM) alongside advanced fine-tuning strategies. Our contributions include the development of a prompt-free adapter for SAM using two stage mask decoding to automatically generate prompt embeddings, a dual-stage fine-tuning method based on Low-Rank Adaptation (LoRA) for enhancing segmentation with limited annotated data, and a 3D memory attention mechanism to ensure segmentation consistency across 3D stacks. We further release a unique benchmark dataset for the segmentation of astrocytic processes and synapses. We evaluated our method on challenging neuroscience segmentation benchmarks, specifically targeting mitochondria, glia, and synapses, with significant accuracy improvements over state-of-the-art (SOTA) methods, including recent SAM-based adapters developed for the medical domain and other vision transformer-based approaches. Experimental results indicate that our approach outperforms existing solutions in the segmentation of complex processes like glia and post-synaptic densities. Our code and models are available at https://github.com/Uzshah/SAM4EM.
Propaganda to Hate: A Multimodal Analysis of Arabic Memes with Multi-Agent LLMs
Sep 11, 2024



Abstract:In the past decade, social media platforms have been used for information dissemination and consumption. While a major portion of the content is posted to promote citizen journalism and public awareness, some content is posted to mislead users. Among different content types such as text, images, and videos, memes (text overlaid on images) are particularly prevalent and can serve as powerful vehicles for propaganda, hate, and humor. In the current literature, there have been efforts to individually detect such content in memes. However, the study of their intersection is very limited. In this study, we explore the intersection between propaganda and hate in memes using a multi-agent LLM-based approach. We extend the propagandistic meme dataset with coarse and fine-grained hate labels. Our finding suggests that there is an association between propaganda and hate in memes. We provide detailed experimental results that can serve as a baseline for future studies. We will make the experimental resources publicly available to the community.
Towards deep observation: A systematic survey on artificial intelligence techniques to monitor fetus via Ultrasound Images
Jan 17, 2022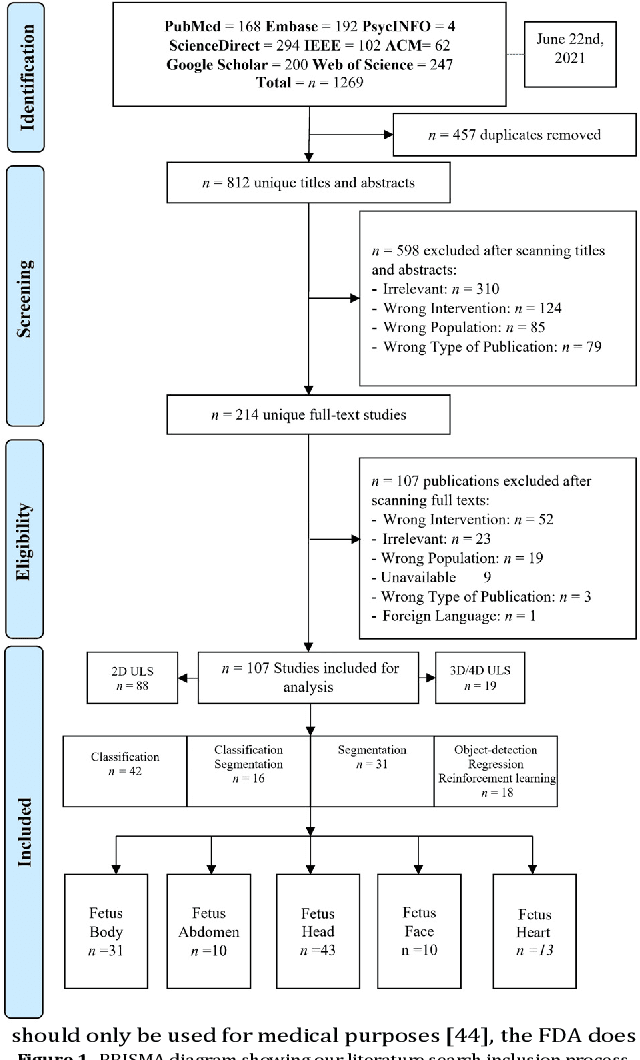
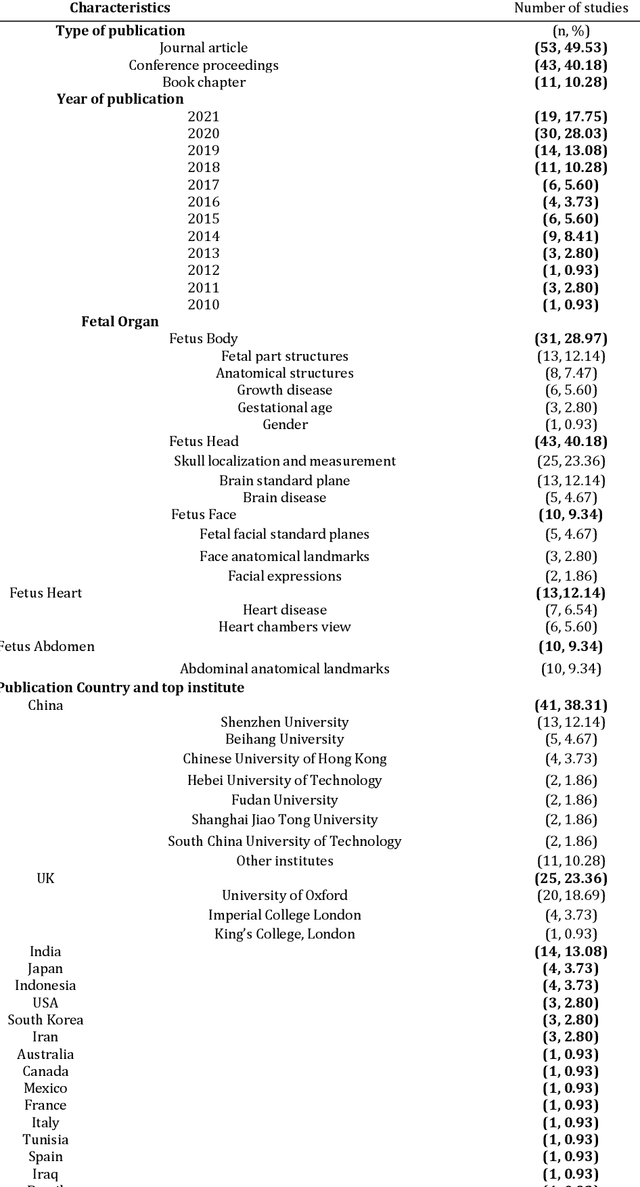
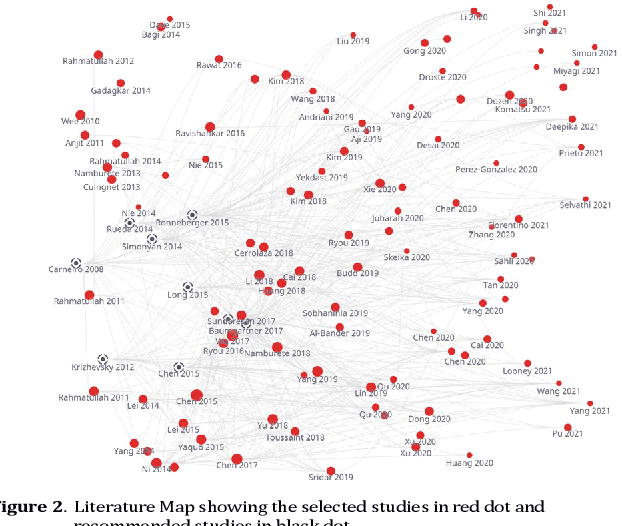
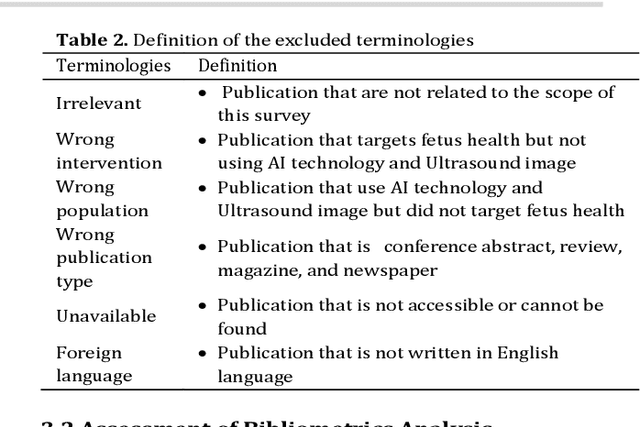

Abstract:Developing innovative informatics approaches aimed to enhance fetal monitoring is a burgeoning field of study in reproductive medicine. Several reviews have been conducted regarding Artificial intelligence (AI) techniques to improve pregnancy outcomes. They are limited by focusing on specific data such as mother's care during pregnancy. This systematic survey aims to explore how artificial intelligence (AI) can assist with fetal growth monitoring via Ultrasound (US) image. We used eight medical and computer science bibliographic databases, including PubMed, Embase, PsycINFO, ScienceDirect, IEEE explore, ACM Library, Google Scholar, and the Web of Science. We retrieved studies published between 2010 to 2021. Data extracted from studies were synthesized using a narrative approach. Out of 1269 retrieved studies, we included 107 distinct studies from queries that were relevant to the topic in the survey. We found that 2D ultrasound images were more popular (n=88) than 3D and 4D ultrasound images (n=19). Classification is the most used method (n=42), followed by segmentation (n=31), classification integrated with segmentation (n=16) and other miscellaneous such as object-detection, regression and reinforcement learning (n=18). The most common areas within the pregnancy domain were the fetus head (n=43), then fetus body (n=31), fetus heart (n=13), fetus abdomen (n=10), and lastly the fetus face (n=10). In the most recent studies, deep learning techniques were primarily used (n=81), followed by machine learning (n=16), artificial neural network (n=7), and reinforcement learning (n=2). AI techniques played a crucial role in predicting fetal diseases and identifying fetus anatomy structures during pregnancy. More research is required to validate this technology from a physician's perspective, such as pilot studies and randomized controlled trials on AI and its applications in a hospital setting.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge