Tomaž Curk
Automated Assignment Grading with Large Language Models: Insights From a Bioinformatics Course
Jan 24, 2025Abstract:Providing students with individualized feedback through assignments is a cornerstone of education that supports their learning and development. Studies have shown that timely, high-quality feedback plays a critical role in improving learning outcomes. However, providing personalized feedback on a large scale in classes with large numbers of students is often impractical due to the significant time and effort required. Recent advances in natural language processing and large language models (LLMs) offer a promising solution by enabling the efficient delivery of personalized feedback. These technologies can reduce the workload of course staff while improving student satisfaction and learning outcomes. Their successful implementation, however, requires thorough evaluation and validation in real classrooms. We present the results of a practical evaluation of LLM-based graders for written assignments in the 2024/25 iteration of the Introduction to Bioinformatics course at the University of Ljubljana. Over the course of the semester, more than 100 students answered 36 text-based questions, most of which were automatically graded using LLMs. In a blind study, students received feedback from both LLMs and human teaching assistants without knowing the source, and later rated the quality of the feedback. We conducted a systematic evaluation of six commercial and open-source LLMs and compared their grading performance with human teaching assistants. Our results show that with well-designed prompts, LLMs can achieve grading accuracy and feedback quality comparable to human graders. Our results also suggest that open-source LLMs perform as well as commercial LLMs, allowing schools to implement their own grading systems while maintaining privacy.
Matrix tri-factorization over the tropical semiring
May 11, 2023



Abstract:Tropical semiring has proven successful in several research areas, including optimal control, bioinformatics, discrete event systems, or solving a decision problem. In previous studies, a matrix two-factorization algorithm based on the tropical semiring has been applied to investigate bipartite and tripartite networks. Tri-factorization algorithms based on standard linear algebra are used for solving tasks such as data fusion, co-clustering, matrix completion, community detection, and more. However, there is currently no tropical matrix tri-factorization approach, which would allow for the analysis of multipartite networks with a high number of parts. To address this, we propose the triFastSTMF algorithm, which performs tri-factorization over the tropical semiring. We apply it to analyze a four-partition network structure and recover the edge lengths of the network. We show that triFastSTMF performs similarly to Fast-NMTF in terms of approximation and prediction performance when fitted on the whole network. When trained on a specific subnetwork and used to predict the whole network, triFastSTMF outperforms Fast-NMTF by several orders of magnitude smaller error. The robustness of triFastSTMF is due to tropical operations, which are less prone to predict large values compared to standard operations.
FastSTMF: Efficient tropical matrix factorization algorithm for sparse data
May 13, 2022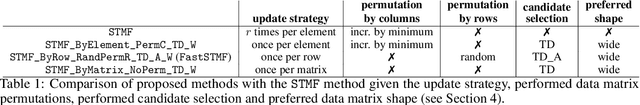
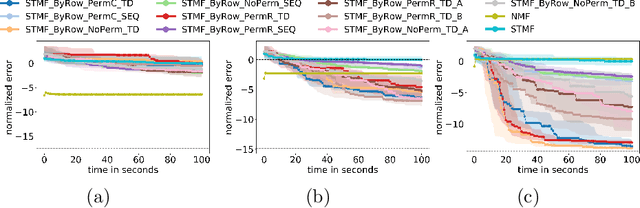
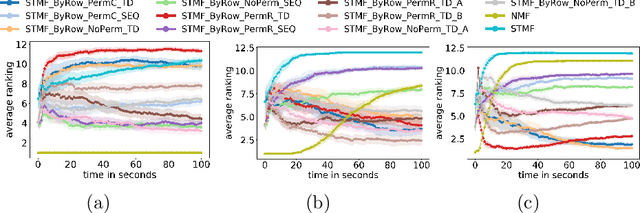
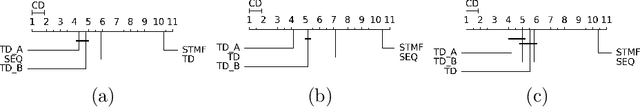
Abstract:Matrix factorization, one of the most popular methods in machine learning, has recently benefited from introducing non-linearity in prediction tasks using tropical semiring. The non-linearity enables a better fit to extreme values and distributions, thus discovering high-variance patterns that differ from those found by standard linear algebra. However, the optimization process of various tropical matrix factorization methods is slow. In our work, we propose a new method FastSTMF based on Sparse Tropical Matrix Factorization (STMF), which introduces a novel strategy for updating factor matrices that results in efficient computational performance. We evaluated the efficiency of FastSTMF on synthetic and real gene expression data from the TCGA database, and the results show that FastSTMF outperforms STMF in both accuracy and running time. Compared to NMF, we show that FastSTMF performs better on some datasets and is not prone to overfitting as NMF. This work sets the basis for developing other matrix factorization techniques based on many other semirings using a new proposed optimization process.
Data embedding and prediction by sparse tropical matrix factorization
Dec 09, 2020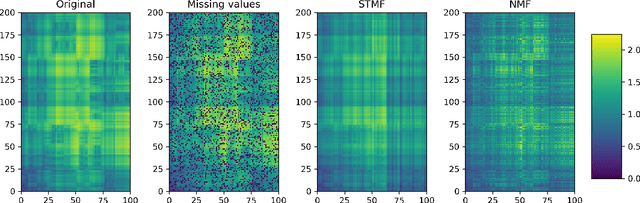
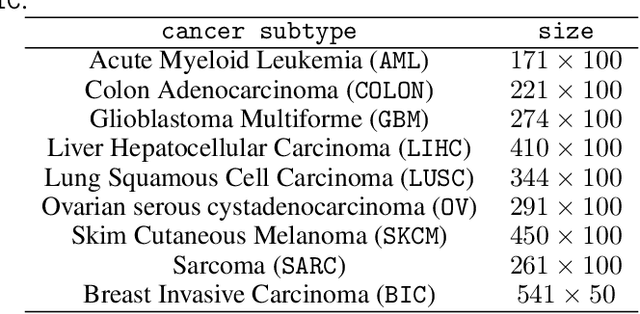

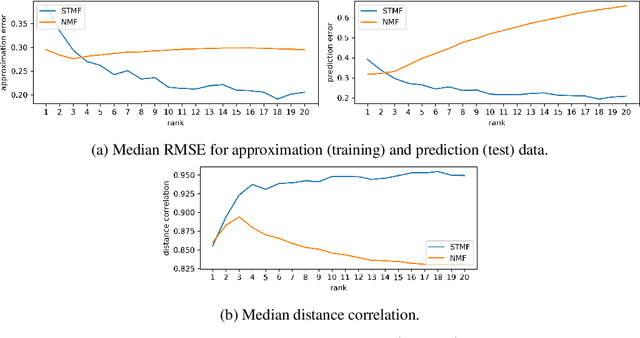
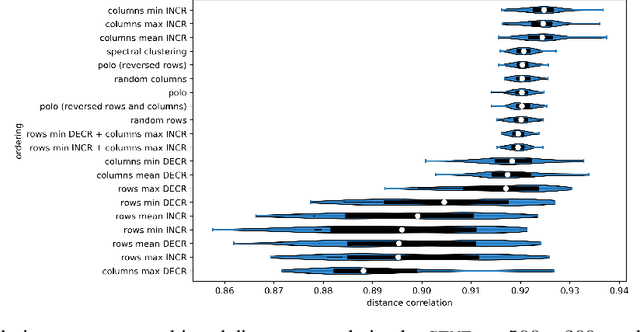
Abstract:Matrix factorization methods are linear models, with limited capability to model complex relations. In our work, we use tropical semiring to introduce non-linearity into matrix factorization models. We propose a method called Sparse Tropical Matrix Factorization (STMF) for the estimation of missing (unknown) values. We evaluate the efficiency of the STMF method on both synthetic data and biological data in the form of gene expression measurements downloaded from The Cancer Genome Atlas (TCGA) database. Tests on unique synthetic data showed that STMF approximation achieves a higher correlation than non-negative matrix factorization (NMF), which is unable to recover patterns effectively. On real data, STMF outperforms NMF on six out of nine gene expression datasets. While NMF assumes normal distribution and tends toward the mean value, STMF can better fit to extreme values and distributions. STMF is the first work that uses tropical semiring on sparse data. We show that in certain cases semirings are useful because they consider the structure, which is different and simpler to understand than it is with standard linear algebra.
Learning the kernel matrix via predictive low-rank approximations
May 09, 2016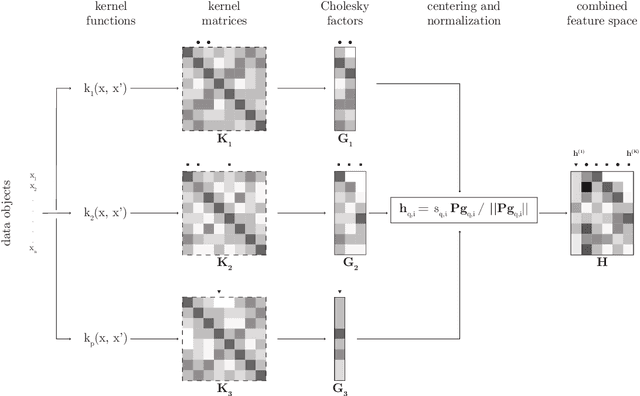
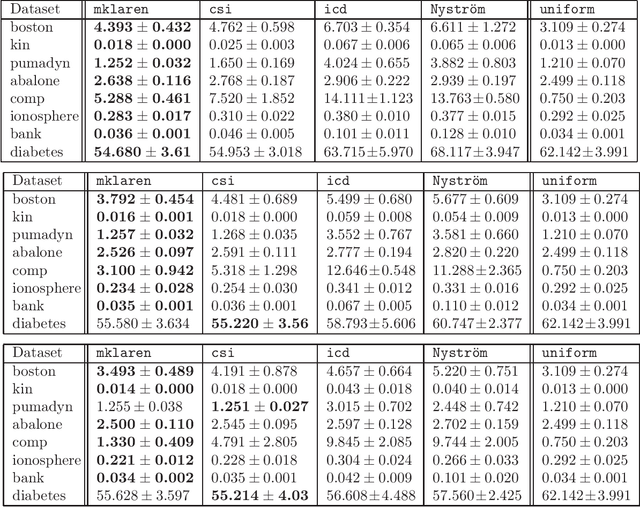
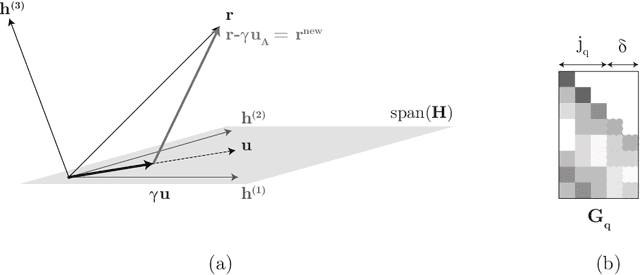
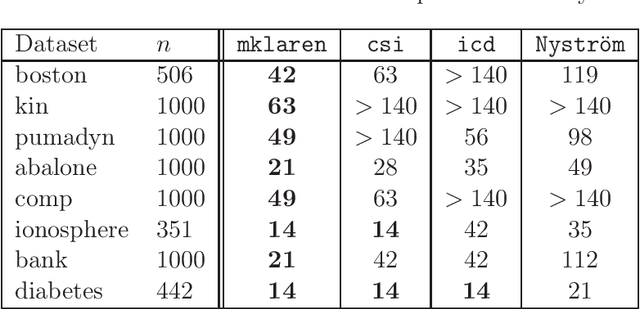
Abstract:Efficient and accurate low-rank approximations of multiple data sources are essential in the era of big data. The scaling of kernel-based learning algorithms to large datasets is limited by the O(n^2) computation and storage complexity of the full kernel matrix, which is required by most of the recent kernel learning algorithms. We present the Mklaren algorithm to approximate multiple kernel matrices learn a regression model, which is entirely based on geometrical concepts. The algorithm does not require access to full kernel matrices yet it accounts for the correlations between all kernels. It uses Incomplete Cholesky decomposition, where pivot selection is based on least-angle regression in the combined, low-dimensional feature space. The algorithm has linear complexity in the number of data points and kernels. When explicit feature space induced by the kernel can be constructed, a mapping from the dual to the primal Ridge regression weights is used for model interpretation. The Mklaren algorithm was tested on eight standard regression datasets. It outperforms contemporary kernel matrix approximation approaches when learning with multiple kernels. It identifies relevant kernels, achieving highest explained variance than other multiple kernel learning methods for the same number of iterations. Test accuracy, equivalent to the one using full kernel matrices, was achieved with at significantly lower approximation ranks. A difference in run times of two orders of magnitude was observed when either the number of samples or kernels exceeds 3000.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge