Blaž Zupan
Inferring Chronic Treatment Onset from ePrescription Data: A Renewal Process Approach
Feb 27, 2026Abstract:Longitudinal electronic health record (EHR) data are often left-censored, making diagnosis records incomplete and unreliable for determining disease onset. In contrast, outpatient prescriptions form renewal-based trajectories that provide a continuous signal of disease management. We propose a probabilistic framework to infer chronic treatment onset by modeling prescription dynamics as a renewal process and detecting transitions from sporadic to sustained therapy via change-point detection between a baseline Poisson (sporadic prescribing) regime and a regime-specific Weibull (sustained therapy) renewal model. Using a nationwide ePrescription dataset of 2.4 million individuals, we show that the approach yields more temporally plausible onset estimates than naive rule-based triggering, substantially reducing implausible early detections under strong left censoring. Detection performance varies across diseases and is strongly associated with prescription density, highlighting both the strengths and limits of treatment-based onset inference.
Automated Assignment Grading with Large Language Models: Insights From a Bioinformatics Course
Jan 24, 2025Abstract:Providing students with individualized feedback through assignments is a cornerstone of education that supports their learning and development. Studies have shown that timely, high-quality feedback plays a critical role in improving learning outcomes. However, providing personalized feedback on a large scale in classes with large numbers of students is often impractical due to the significant time and effort required. Recent advances in natural language processing and large language models (LLMs) offer a promising solution by enabling the efficient delivery of personalized feedback. These technologies can reduce the workload of course staff while improving student satisfaction and learning outcomes. Their successful implementation, however, requires thorough evaluation and validation in real classrooms. We present the results of a practical evaluation of LLM-based graders for written assignments in the 2024/25 iteration of the Introduction to Bioinformatics course at the University of Ljubljana. Over the course of the semester, more than 100 students answered 36 text-based questions, most of which were automatically graded using LLMs. In a blind study, students received feedback from both LLMs and human teaching assistants without knowing the source, and later rated the quality of the feedback. We conducted a systematic evaluation of six commercial and open-source LLMs and compared their grading performance with human teaching assistants. Our results show that with well-designed prompts, LLMs can achieve grading accuracy and feedback quality comparable to human graders. Our results also suggest that open-source LLMs perform as well as commercial LLMs, allowing schools to implement their own grading systems while maintaining privacy.
VERA: Generating Visual Explanations of Two-Dimensional Embeddings via Region Annotation
Jun 07, 2024Abstract:Two-dimensional embeddings obtained from dimensionality reduction techniques, such as MDS, t-SNE, and UMAP, are widely used across various disciplines to visualize high-dimensional data. These visualizations provide a valuable tool for exploratory data analysis, allowing researchers to visually identify clusters, outliers, and other interesting patterns in the data. However, interpreting the resulting visualizations can be challenging, as it often requires additional manual inspection to understand the differences between data points in different regions of the embedding space. To address this issue, we propose Visual Explanations via Region Annotation (VERA), an automatic embedding-annotation approach that generates visual explanations for any two-dimensional embedding. VERA produces informative explanations that characterize distinct regions in the embedding space, allowing users to gain an overview of the embedding landscape at a glance. Unlike most existing approaches, which typically require some degree of manual user intervention, VERA produces static explanations, automatically identifying and selecting the most informative visual explanations to show to the user. We illustrate the usage of VERA on a real-world data set and validate the utility of our approach with a comparative user study. Our results demonstrate that the explanations generated by VERA are as useful as fully-fledged interactive tools on typical exploratory data analysis tasks but require significantly less time and effort from the user.
Visualizing High-Dimensional Temporal Data Using Direction-Aware t-SNE
Mar 27, 2024Abstract:Many real-world data sets contain a temporal component or involve transitions from state to state. For exploratory data analysis, we can represent these high-dimensional data sets in two-dimensional maps, using embeddings of the data objects under exploration and representing their temporal relationships with directed edges. Most existing dimensionality reduction techniques, such as t-SNE and UMAP, do not take into account the temporal or relational nature of the data when constructing the embeddings, resulting in temporally cluttered visualizations that obscure potentially interesting patterns. To address this problem, we propose two complementary, direction-aware loss terms in the optimization function of t-SNE that emphasize the temporal aspects of the data, guiding the optimization and the resulting embedding to reveal temporal patterns that might otherwise go unnoticed. The Directional Coherence Loss (DCL) encourages nearby arrows connecting two adjacent time series points to point in the same direction, while the Edge Length Loss (ELL) penalizes arrows - which effectively represent time gaps in the visualized embedding - based on their length. Both loss terms are differentiable and can be easily incorporated into existing dimensionality reduction techniques. By promoting local directionality of the directed edges, our procedure produces more temporally meaningful and less cluttered visualizations. We demonstrate the effectiveness of our approach on a toy dataset and two real-world datasets.
Explainable Point-Based Document Visualizations
Sep 28, 2021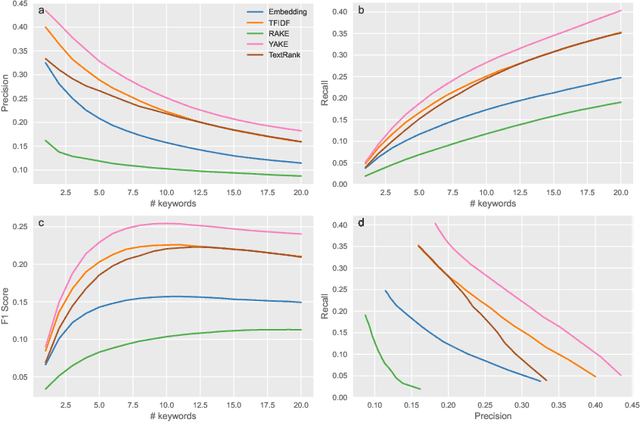
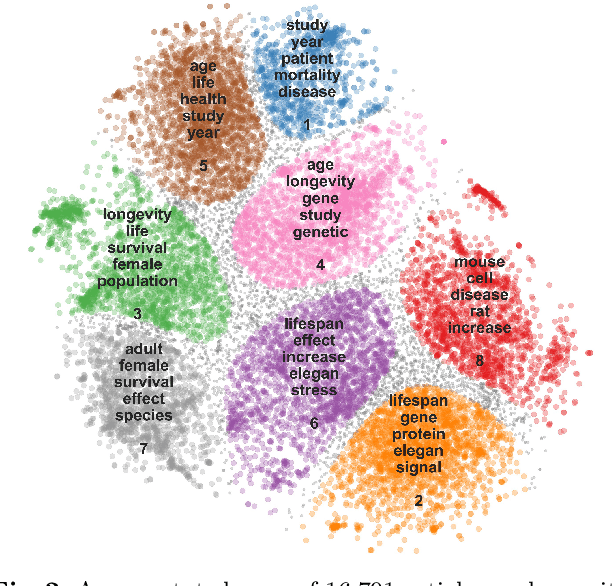
Abstract:Two-dimensional data maps can visually reveal information about the relations between data instances. Popular techniques to construct data maps are t-SNE and UMAP. The resulting point-based visualizations, though, provide information only through their interpretation. We here consider a set of abstracts from the articles on longevity to argue for using keyword extraction methods to label clusters of documents in the map. Among the considered approaches, the best results were obtained by recently proposed YAKE!. Surprisingly, a classical TF-IDF term ranking outperformed graph and embedding-based techniques.
Data Fusion by Matrix Factorization
Feb 06, 2015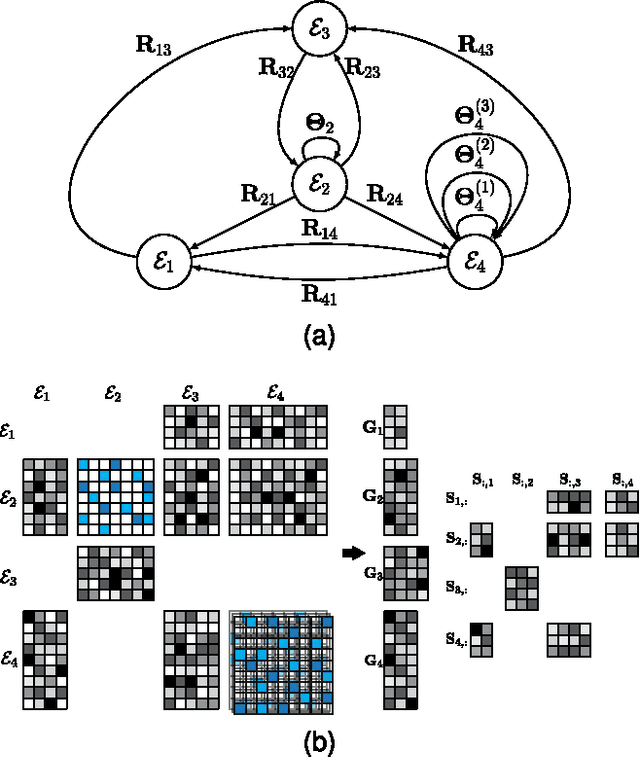
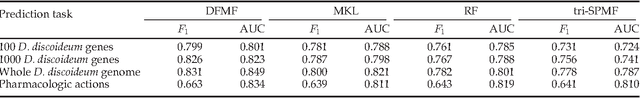
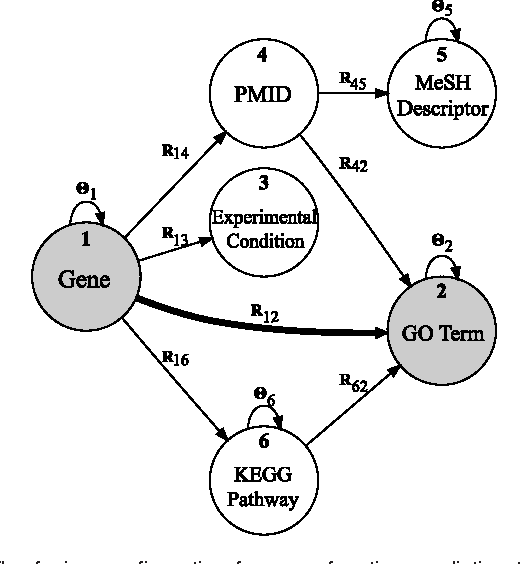
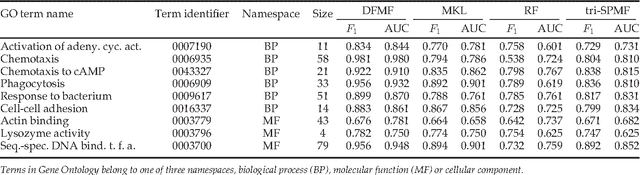
Abstract:For most problems in science and engineering we can obtain data sets that describe the observed system from various perspectives and record the behavior of its individual components. Heterogeneous data sets can be collectively mined by data fusion. Fusion can focus on a specific target relation and exploit directly associated data together with contextual data and data about system's constraints. In the paper we describe a data fusion approach with penalized matrix tri-factorization (DFMF) that simultaneously factorizes data matrices to reveal hidden associations. The approach can directly consider any data that can be expressed in a matrix, including those from feature-based representations, ontologies, associations and networks. We demonstrate the utility of DFMF for gene function prediction task with eleven different data sources and for prediction of pharmacologic actions by fusing six data sources. Our data fusion algorithm compares favorably to alternative data integration approaches and achieves higher accuracy than can be obtained from any single data source alone.
* Short preprint, 13 pages, 3 Figures, 3 Tables. Full paper in 10.1109/TPAMI.2014.2343973
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge