Polona Oblak
Matrix tri-factorization over the tropical semiring
May 11, 2023



Abstract:Tropical semiring has proven successful in several research areas, including optimal control, bioinformatics, discrete event systems, or solving a decision problem. In previous studies, a matrix two-factorization algorithm based on the tropical semiring has been applied to investigate bipartite and tripartite networks. Tri-factorization algorithms based on standard linear algebra are used for solving tasks such as data fusion, co-clustering, matrix completion, community detection, and more. However, there is currently no tropical matrix tri-factorization approach, which would allow for the analysis of multipartite networks with a high number of parts. To address this, we propose the triFastSTMF algorithm, which performs tri-factorization over the tropical semiring. We apply it to analyze a four-partition network structure and recover the edge lengths of the network. We show that triFastSTMF performs similarly to Fast-NMTF in terms of approximation and prediction performance when fitted on the whole network. When trained on a specific subnetwork and used to predict the whole network, triFastSTMF outperforms Fast-NMTF by several orders of magnitude smaller error. The robustness of triFastSTMF is due to tropical operations, which are less prone to predict large values compared to standard operations.
SMIXS: Novel efficient algorithm for non-parametric mixture regression-based clustering
Sep 19, 2022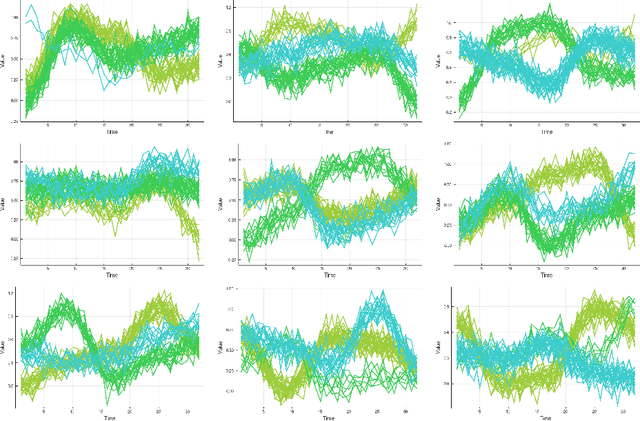
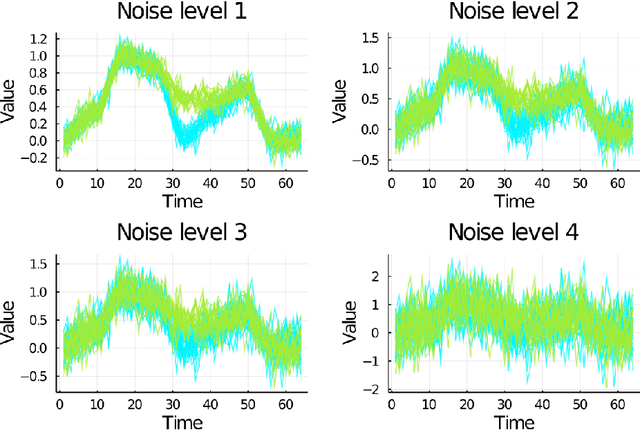
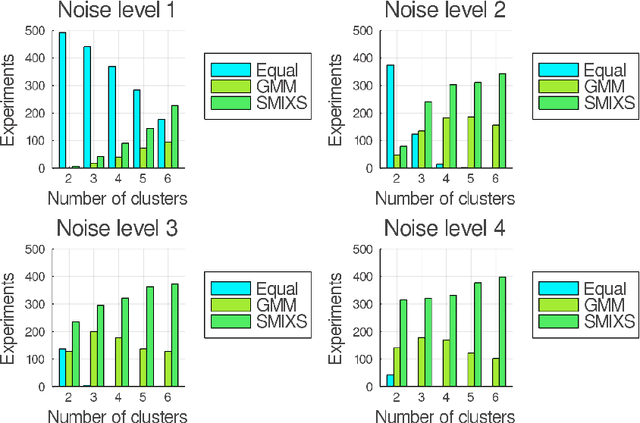
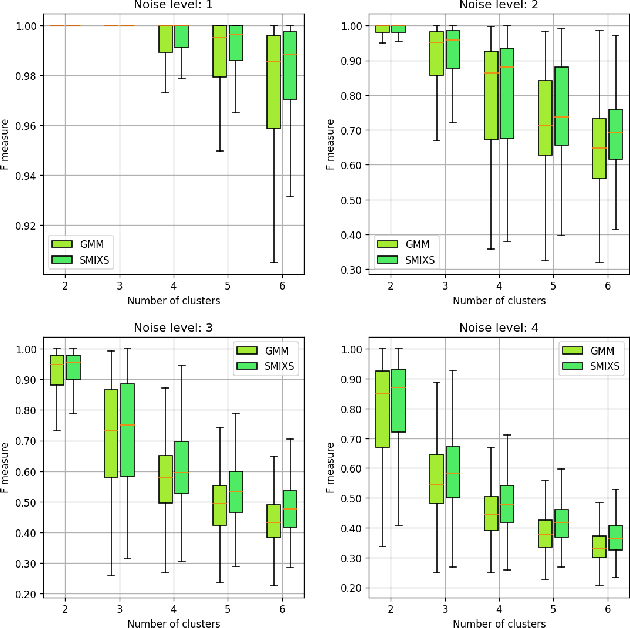
Abstract:We investigate a novel non-parametric regression-based clustering algorithm for longitudinal data analysis. Combining natural cubic splines with Gaussian mixture models (GMM), the algorithm can produce smooth cluster means that describe the underlying data well. However, there are some shortcomings in the algorithm: high computational complexity in the parameter estimation procedure and a numerically unstable variance estimator. Therefore, to further increase the usability of the method, we incorporated approaches to reduce its computational complexity, we developed a new, more stable variance estimator, and we developed a new smoothing parameter estimation procedure. We show that the developed algorithm, SMIXS, performs better than GMM on a synthetic dataset in terms of clustering and regression performance. We demonstrate the impact of the computational speed-ups, which we formally prove in the new framework. Finally, we perform a case study by using SMIXS to cluster vertical atmospheric measurements to determine different weather regimes.
FastSTMF: Efficient tropical matrix factorization algorithm for sparse data
May 13, 2022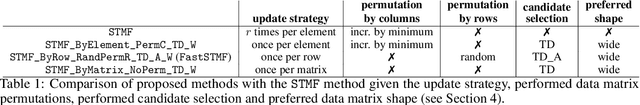
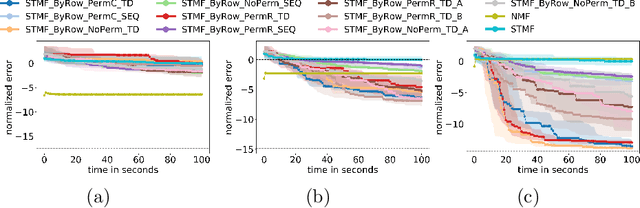
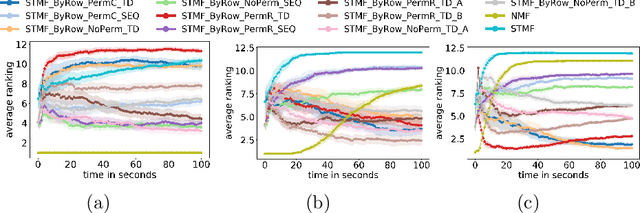
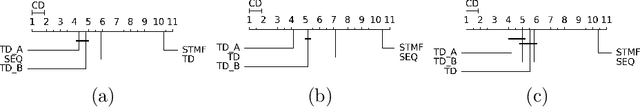
Abstract:Matrix factorization, one of the most popular methods in machine learning, has recently benefited from introducing non-linearity in prediction tasks using tropical semiring. The non-linearity enables a better fit to extreme values and distributions, thus discovering high-variance patterns that differ from those found by standard linear algebra. However, the optimization process of various tropical matrix factorization methods is slow. In our work, we propose a new method FastSTMF based on Sparse Tropical Matrix Factorization (STMF), which introduces a novel strategy for updating factor matrices that results in efficient computational performance. We evaluated the efficiency of FastSTMF on synthetic and real gene expression data from the TCGA database, and the results show that FastSTMF outperforms STMF in both accuracy and running time. Compared to NMF, we show that FastSTMF performs better on some datasets and is not prone to overfitting as NMF. This work sets the basis for developing other matrix factorization techniques based on many other semirings using a new proposed optimization process.
Data embedding and prediction by sparse tropical matrix factorization
Dec 09, 2020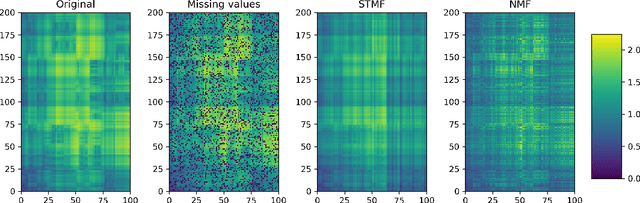
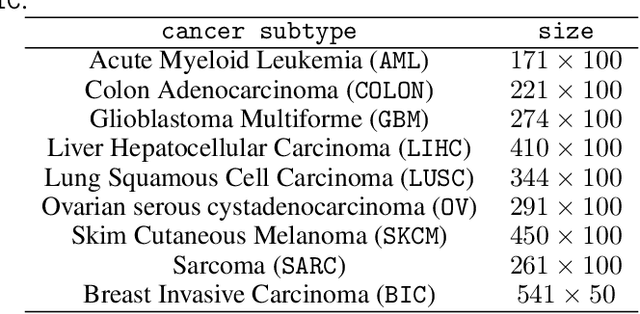

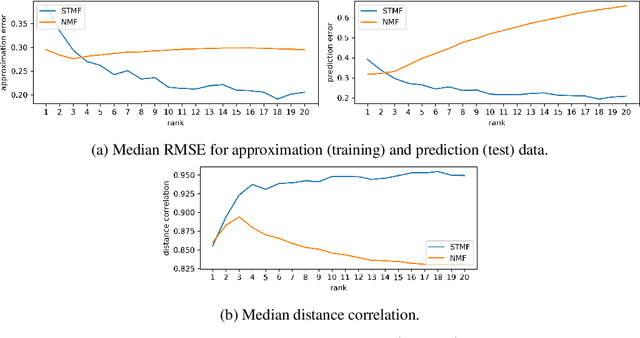
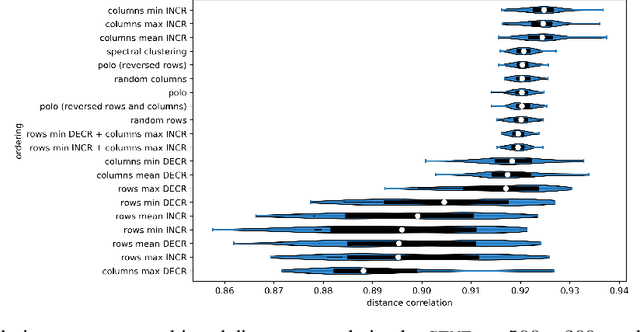
Abstract:Matrix factorization methods are linear models, with limited capability to model complex relations. In our work, we use tropical semiring to introduce non-linearity into matrix factorization models. We propose a method called Sparse Tropical Matrix Factorization (STMF) for the estimation of missing (unknown) values. We evaluate the efficiency of the STMF method on both synthetic data and biological data in the form of gene expression measurements downloaded from The Cancer Genome Atlas (TCGA) database. Tests on unique synthetic data showed that STMF approximation achieves a higher correlation than non-negative matrix factorization (NMF), which is unable to recover patterns effectively. On real data, STMF outperforms NMF on six out of nine gene expression datasets. While NMF assumes normal distribution and tends toward the mean value, STMF can better fit to extreme values and distributions. STMF is the first work that uses tropical semiring on sparse data. We show that in certain cases semirings are useful because they consider the structure, which is different and simpler to understand than it is with standard linear algebra.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge