Tim W. Nattkemper
Self-Supervised Federated Learning under Data Heterogeneity for Label-Scarce Diatom Classification
Mar 31, 2026Abstract:Label-scarce visual classification under decentralized and heterogeneous data is a fundamental challenge in pattern recognition, especially when sites exhibit partially overlapping class sets. While self-supervised federated learning (SSFL) offers a promising solution, existing studies commonly assume the same data heterogeneity pattern throughout pre-training and fine-tuning. Moreover, current partitioning schemes often fail to generate pure partially class-disjoint data settings, limiting controllable simulation of real-world label-space heterogeneity. In this work, we introduce SSFL for diatom classification as a representative real-world instance and systematically investigate stage-specific data heterogeneity. We study cross-site variation in unlabeled data volume during pre-training and label-space misalignment during downstream fine-tuning. To study the latter in a controllable setting, we propose PreDi, a partitioning scheme that disentangles label-space heterogeneity into two orthogonal dimensions, namely class Prevalence and class-set size Disparity, enabling separate analysis of their effects. Guided by the resulting insights, we further propose PreP-WFL (Prevalence-based Personalized Weighted Federated Learning) to adaptively strengthen rare-class representations in low-prevalence scenarios. Extensive experiments show that SSFL consistently outperforms local-only training under both homogeneous and heterogeneous settings. The pronounced heterogeneity in unlabeled data volume is associated with improved representation pre-training, whereas under label-space heterogeneity, prevalence dominates performance and disparity has a smaller effect. PreP-WFL effectively mitigates this degradation, with gains increasing as prevalence decreases. These findings provide a mechanistic basis for characterizing label-space heterogeneity in decentralized recognition systems.
SoRC -- Evaluation of Computational Molecular Co-Localization Analysis in Mass Spectrometry Images
Sep 24, 2020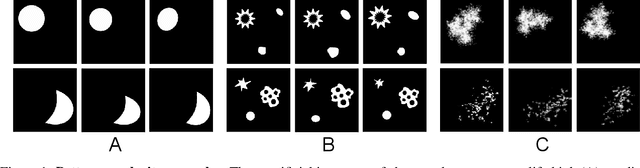
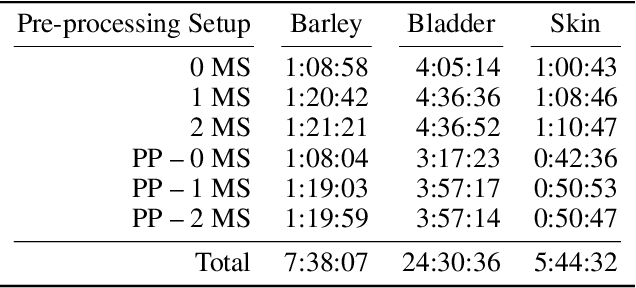
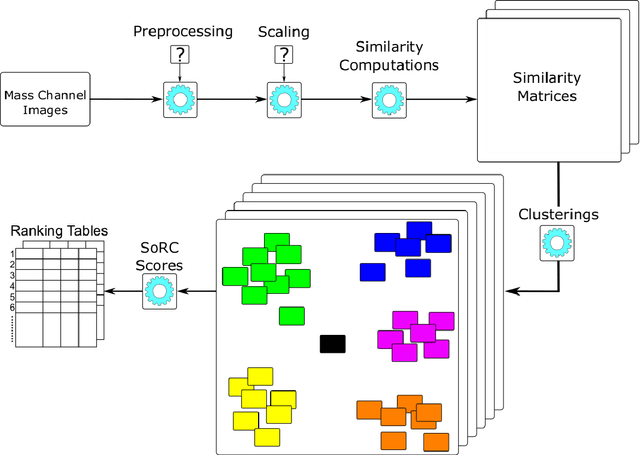
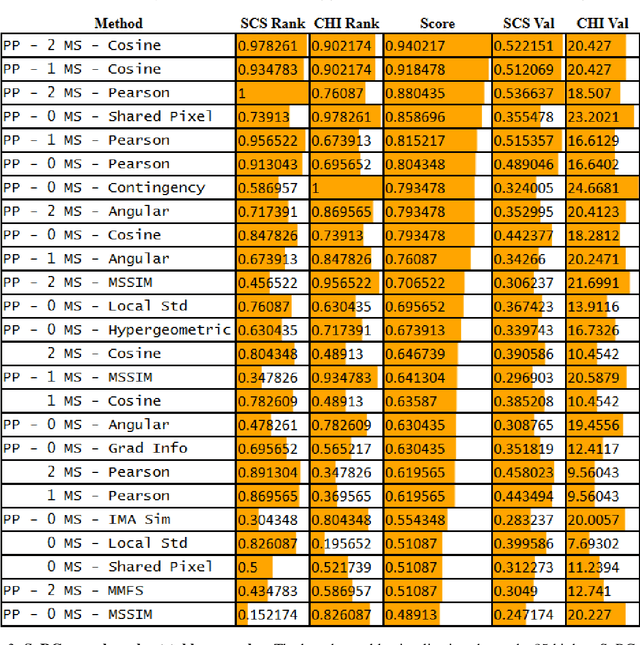
Abstract:The computational analysis of Mass Spectrometry Imaging (MSI) data aims at the identification of interesting mass co-localizations and the visualization of their lateral distribution in the sample, usually a tissue cross section. But as the morphological structure of tissues and the different kinds of mass co-localization naturally show a huge diversity, the selection and tuning of the computational method is a time-consuming effort. In this work we address the special problem of computationally grouping mass channel images according to their similarities in their lateral distribution patterns. Such an analysis is driven by the idea, that groups of molecules that feature a similar distribution pattern may have a functional relation. But the selection of the similarity function and other parameters is often done by a time-consuming and unsatsifactory trial and error. We propose a new flexible workflow scheme called SoRC (sum of ranked cluster indices) for automating this tuning step and making it much more efficient. We test SoRC using three different data sets acquired from the lab for three different kinds of samples (barley seed, mouse bladder tissue, human PXE skin). We show, that SORC can be applied to score and visualize the results obtained with the applied methods in short time without too much effort. In our application example, the SoRC results for the three data sets reveal that a) some well-known similarity functions are suited to achieve good results for all three data sets and b) for the MSI data featuring a higher degree of irregularity improved results can be achieved by applying non-standard similarity functions. The SoRC scores computed with our approach indicate that an automated testing and scoring of different methods for mass channel image grouping can improve the final outcome of a study by finally selecting the methods of the highest scores.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge