Thomas Lukasiewicz
Universal Hopfield Networks: A General Framework for Single-Shot Associative Memory Models
Feb 09, 2022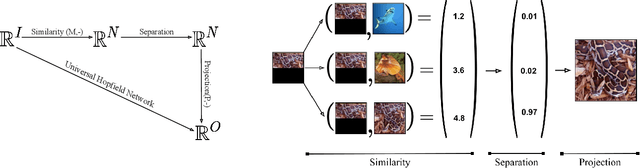
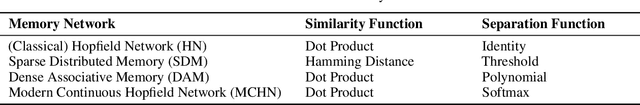
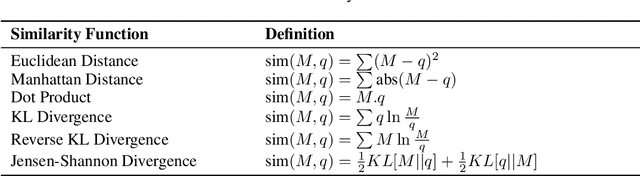

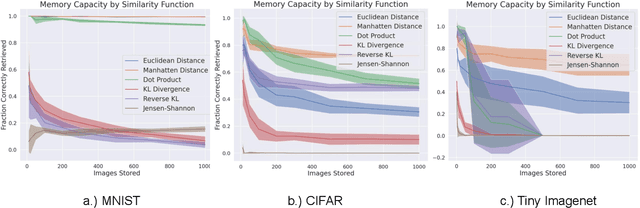

Abstract:A large number of neural network models of associative memory have been proposed in the literature. These include the classical Hopfield networks (HNs), sparse distributed memories (SDMs), and more recently the modern continuous Hopfield networks (MCHNs), which possesses close links with self-attention in machine learning. In this paper, we propose a general framework for understanding the operation of such memory networks as a sequence of three operations: similarity, separation, and projection. We derive all these memory models as instances of our general framework with differing similarity and separation functions. We extend the mathematical framework of Krotov et al (2020) to express general associative memory models using neural network dynamics with only second-order interactions between neurons, and derive a general energy function that is a Lyapunov function of the dynamics. Finally, using our framework, we empirically investigate the capacity of using different similarity functions for these associative memory models, beyond the dot product similarity measure, and demonstrate empirically that Euclidean or Manhattan distance similarity metrics perform substantially better in practice on many tasks, enabling a more robust retrieval and higher memory capacity than existing models.
Learning on Arbitrary Graph Topologies via Predictive Coding
Feb 05, 2022
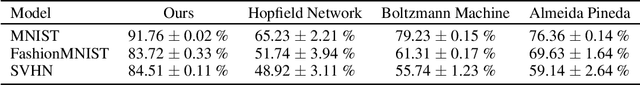
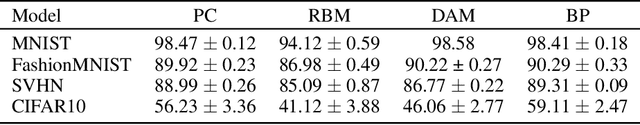

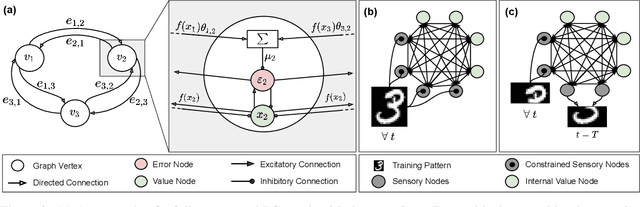

Abstract:Training with backpropagation (BP) in standard deep learning consists of two main steps: a forward pass that maps a data point to its prediction, and a backward pass that propagates the error of this prediction back through the network. This process is highly effective when the goal is to minimize a specific objective function. However, it does not allow training on networks with cyclic or backward connections. This is an obstacle to reaching brain-like capabilities, as the highly complex heterarchical structure of the neural connections in the neocortex are potentially fundamental for its effectiveness. In this paper, we show how predictive coding (PC), a theory of information processing in the cortex, can be used to perform inference and learning on arbitrary graph topologies. We experimentally show how this formulation, called PC graphs, can be used to flexibly perform different tasks with the same network by simply stimulating specific neurons, and investigate how the topology of the graph influences the final performance. We conclude by comparing against simple baselines trained~with~BP.
The Defeat of the Winograd Schema Challenge
Jan 16, 2022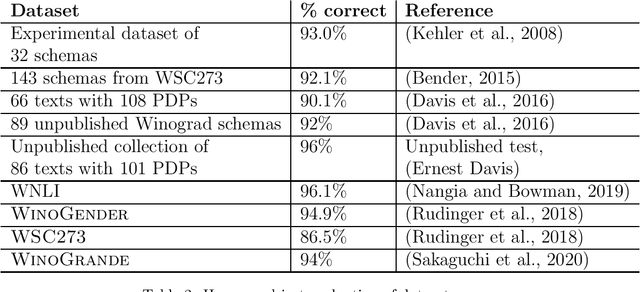

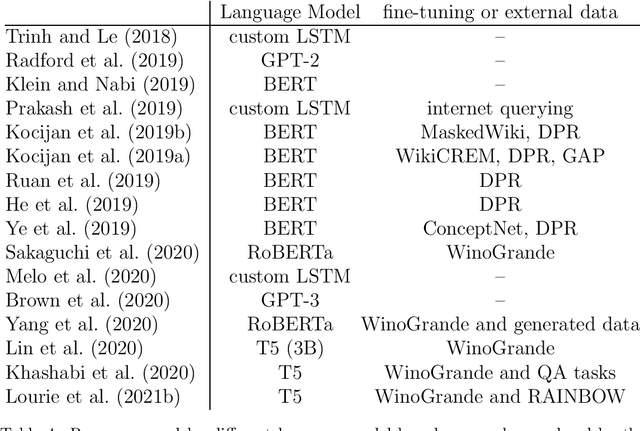
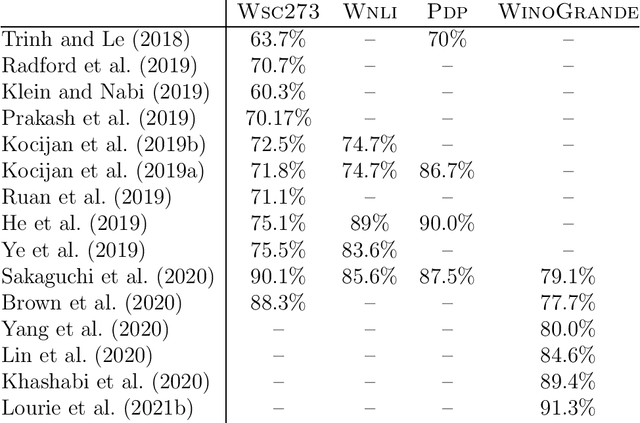
Abstract:The Winograd Schema Challenge -- a set of twin sentences involving pronoun reference disambiguation that seem to require the use of commonsense knowledge -- was proposed by Hector Levesque in 2011. By 2019, a number of AI systems, based on large pre-trained transformer-based language models and fine-tuned on these kinds of problems, achieved better than 90% accuracy. In this paper, we review the history of the Winograd Schema Challenge and assess its significance.
Few-Shot Out-of-Domain Transfer Learning of Natural Language Explanations
Dec 12, 2021
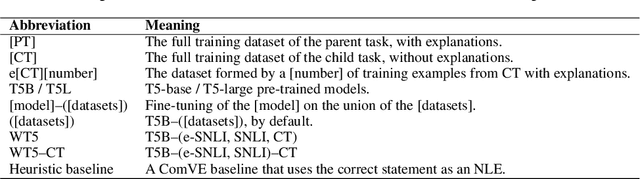
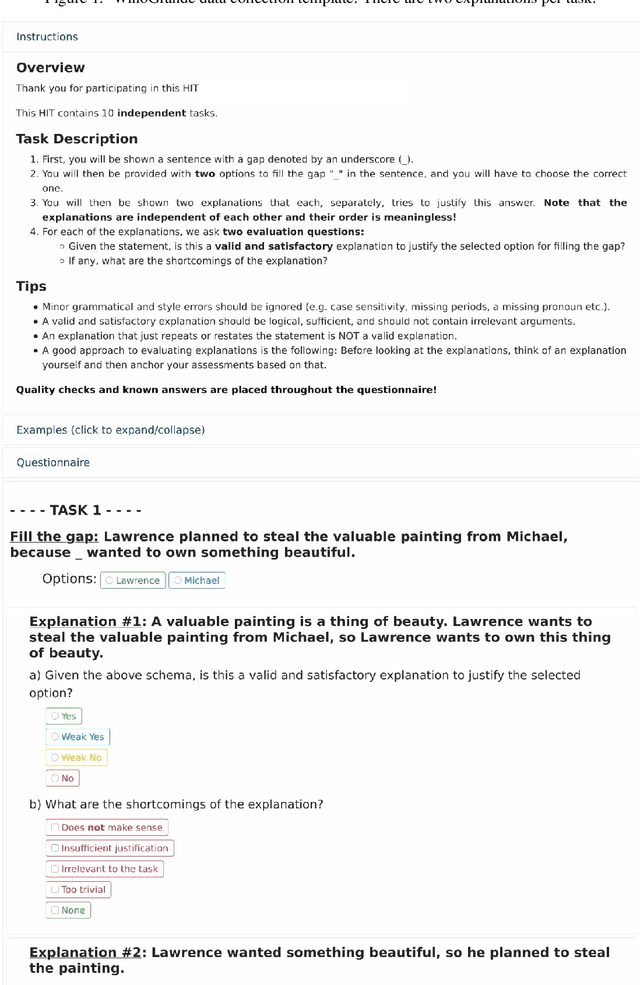

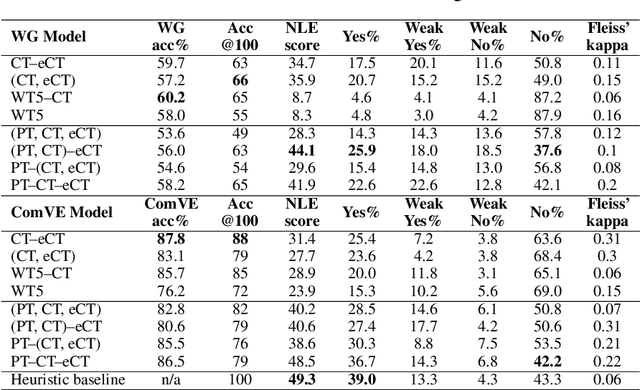
Abstract:Recently, there has been an increasing interest in models that generate natural language explanations (NLEs) for their decisions. However, training a model to provide NLEs requires the acquisition of task-specific NLEs, which is time- and resource-consuming. A potential solution is the out-of-domain transfer of NLEs from a domain with a large number of NLEs to a domain with scarce NLEs but potentially a large number of labels, via few-shot transfer learning. In this work, we introduce three vanilla approaches for few-shot transfer learning of NLEs for the case of few NLEs but abundant labels, along with an adaptation of an existing vanilla fine-tuning approach. We transfer explainability from the natural language inference domain, where a large dataset of human-written NLEs exists (e-SNLI), to the domains of (1) hard cases of pronoun resolution, where we introduce a small dataset of NLEs on top of the WinoGrande dataset (small-e-WinoGrande), and (2) commonsense validation (ComVE). Our results demonstrate that the transfer of NLEs outperforms the single-task methods, and establish the best strategies out of the four identified training regimes. We also investigate the scalability of the best methods, both in terms of training data and model size.
Rationale production to support clinical decision-making
Nov 15, 2021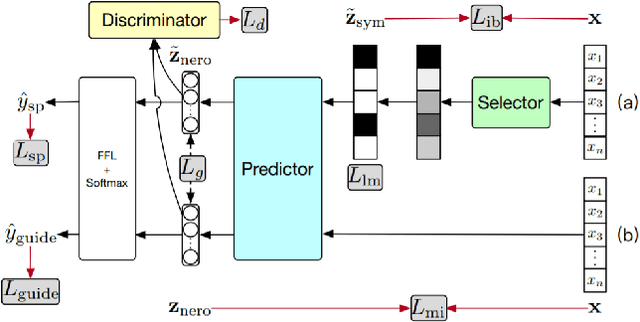
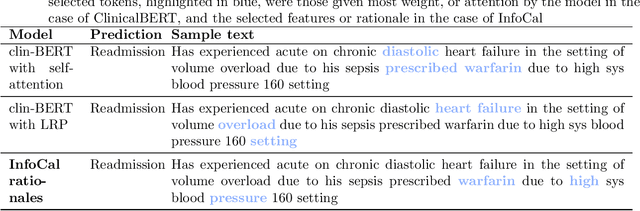
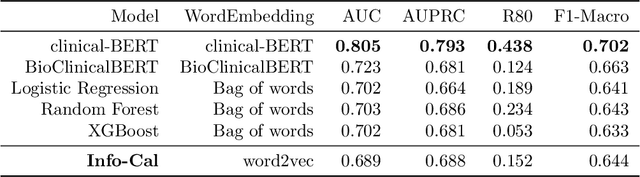

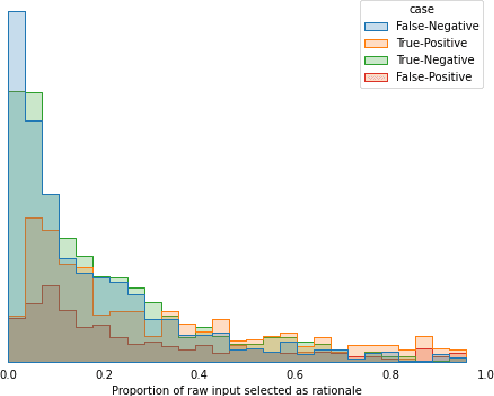
Abstract:The development of neural networks for clinical artificial intelligence (AI) is reliant on interpretability, transparency, and performance. The need to delve into the black-box neural network and derive interpretable explanations of model output is paramount. A task of high clinical importance is predicting the likelihood of a patient being readmitted to hospital in the near future to enable efficient triage. With the increasing adoption of electronic health records (EHRs), there is great interest in applications of natural language processing (NLP) to clinical free-text contained within EHRs. In this work, we apply InfoCal, the current state-of-the-art model that produces extractive rationales for its predictions, to the task of predicting hospital readmission using hospital discharge notes. We compare extractive rationales produced by InfoCal to competitive transformer-based models pretrained on clinical text data and for which the attention mechanism can be used for interpretation. We find each presented model with selected interpretability or feature importance methods yield varying results, with clinical language domain expertise and pretraining critical to performance and subsequent interpretability.
Associative Memories via Predictive Coding
Sep 16, 2021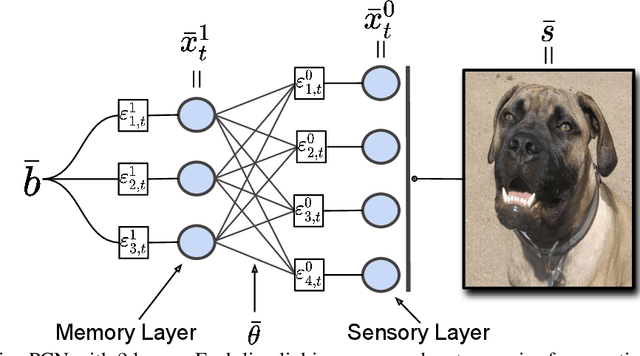
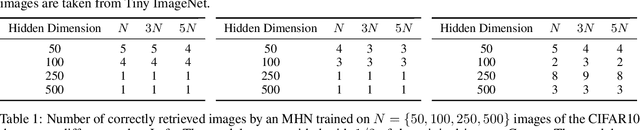
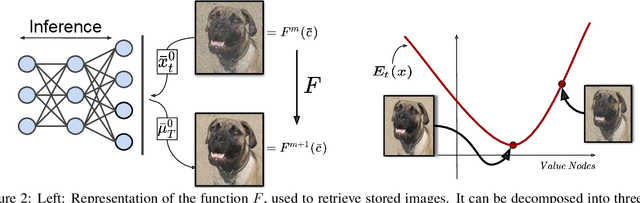
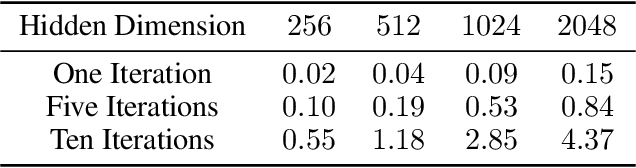
Abstract:Associative memories in the brain receive and store patterns of activity registered by the sensory neurons, and are able to retrieve them when necessary. Due to their importance in human intelligence, computational models of associative memories have been developed for several decades now. They include autoassociative memories, which allow for storing data points and retrieving a stored data point $s$ when provided with a noisy or partial variant of $s$, and heteroassociative memories, able to store and recall multi-modal data. In this paper, we present a novel neural model for realizing associative memories, based on a hierarchical generative network that receives external stimuli via sensory neurons. This model is trained using predictive coding, an error-based learning algorithm inspired by information processing in the cortex. To test the capabilities of this model, we perform multiple retrieval experiments from both corrupted and incomplete data points. In an extensive comparison, we show that this new model outperforms in retrieval accuracy and robustness popular associative memory models, such as autoencoders trained via backpropagation, and modern Hopfield networks. In particular, in completing partial data points, our model achieves remarkable results on natural image datasets, such as ImageNet, with a surprisingly high accuracy, even when only a tiny fraction of pixels of the original images is presented. Furthermore, we show that this method is able to handle multi-modal data, retrieving images from descriptions, and vice versa. We conclude by discussing the possible impact of this work in the neuroscience community, by showing that our model provides a plausible framework to study learning and retrieval of memories in the brain, as it closely mimics the behavior of the hippocampus as a memory index and generative model.
Knowledge Base Completion Meets Transfer Learning
Aug 30, 2021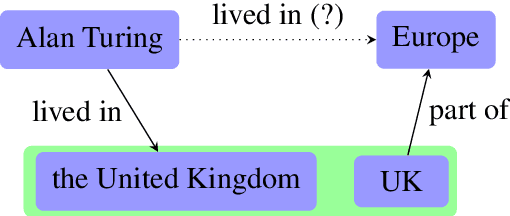

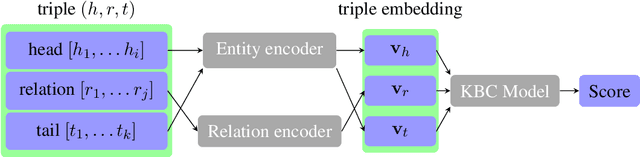
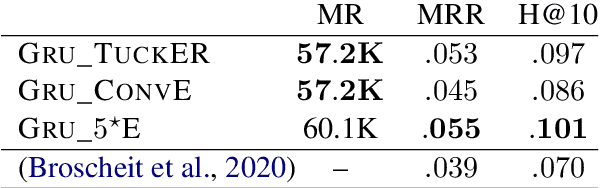
Abstract:The aim of knowledge base completion is to predict unseen facts from existing facts in knowledge bases. In this work, we introduce the first approach for transfer of knowledge from one collection of facts to another without the need for entity or relation matching. The method works for both canonicalized knowledge bases and uncanonicalized or open knowledge bases, i.e., knowledge bases where more than one copy of a real-world entity or relation may exist. Such knowledge bases are a natural output of automated information extraction tools that extract structured data from unstructured text. Our main contribution is a method that can make use of a large-scale pre-training on facts, collected from unstructured text, to improve predictions on structured data from a specific domain. The introduced method is the most impactful on small datasets such as ReVerb20K, where we obtained 6% absolute increase of mean reciprocal rank and 65% relative decrease of mean rank over the previously best method, despite not relying on large pre-trained models like BERT.
* Presented at EMNLP 2021
Accurate, yet inconsistent? Consistency Analysis on Language Understanding Models
Aug 15, 2021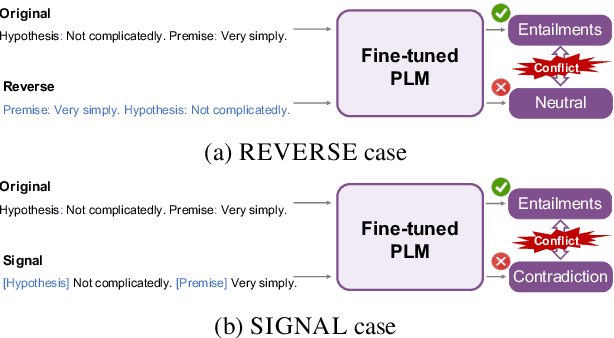
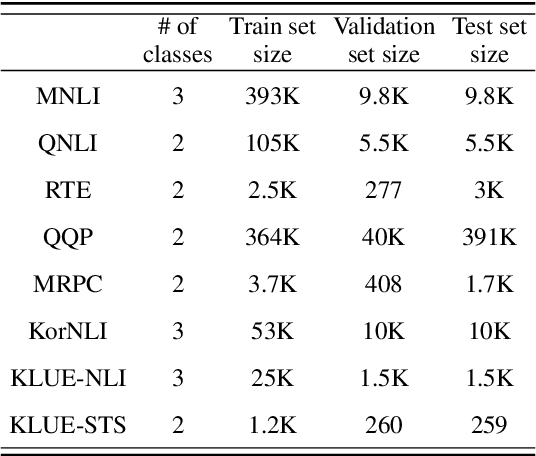
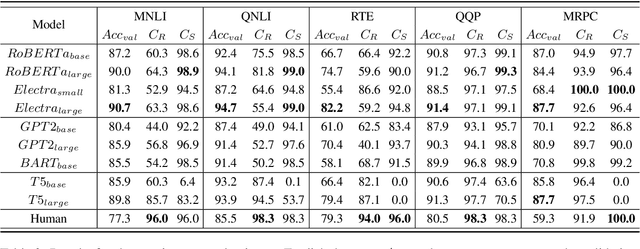
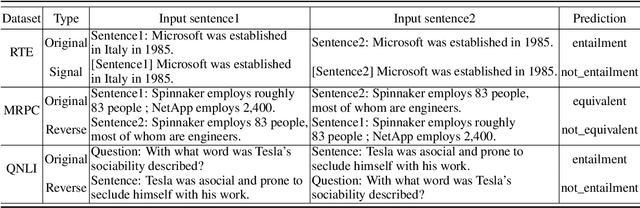
Abstract:Consistency, which refers to the capability of generating the same predictions for semantically similar contexts, is a highly desirable property for a sound language understanding model. Although recent pretrained language models (PLMs) deliver outstanding performance in various downstream tasks, they should exhibit consistent behaviour provided the models truly understand language. In this paper, we propose a simple framework named consistency analysis on language understanding models (CALUM)} to evaluate the model's lower-bound consistency ability. Through experiments, we confirmed that current PLMs are prone to generate inconsistent predictions even for semantically identical inputs. We also observed that multi-task training with paraphrase identification tasks is of benefit to improve consistency, increasing the consistency by 13% on average.
Selective Pseudo-label Clustering
Jul 22, 2021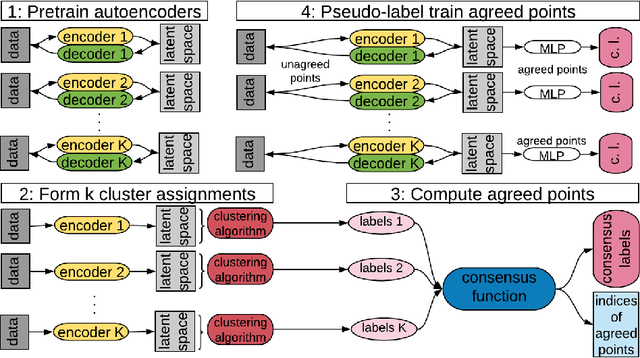
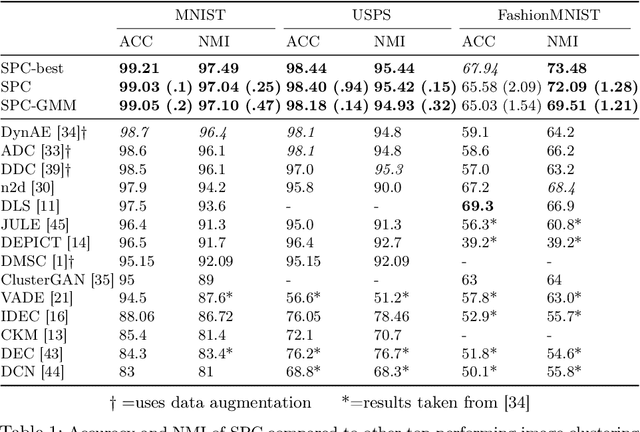
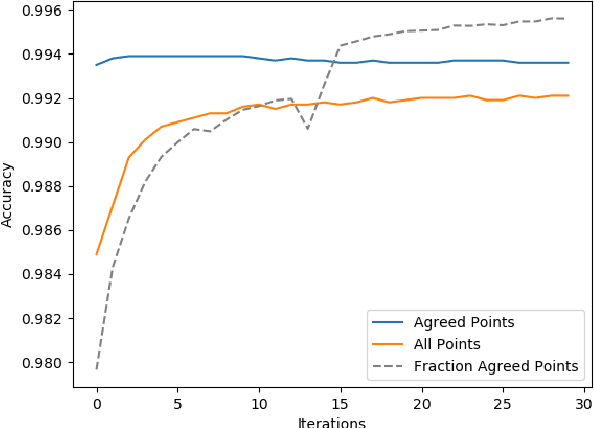
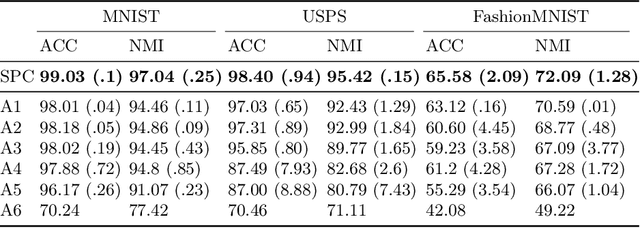
Abstract:Deep neural networks (DNNs) offer a means of addressing the challenging task of clustering high-dimensional data. DNNs can extract useful features, and so produce a lower dimensional representation, which is more amenable to clustering techniques. As clustering is typically performed in a purely unsupervised setting, where no training labels are available, the question then arises as to how the DNN feature extractor can be trained. The most accurate existing approaches combine the training of the DNN with the clustering objective, so that information from the clustering process can be used to update the DNN to produce better features for clustering. One problem with this approach is that these ``pseudo-labels'' produced by the clustering algorithm are noisy, and any errors that they contain will hurt the training of the DNN. In this paper, we propose selective pseudo-label clustering, which uses only the most confident pseudo-labels for training the~DNN. We formally prove the performance gains under certain conditions. Applied to the task of image clustering, the new approach achieves a state-of-the-art performance on three popular image datasets. Code is available at https://github.com/Lou1sM/clustering.
Rationale-Inspired Natural Language Explanations with Commonsense
Jun 25, 2021
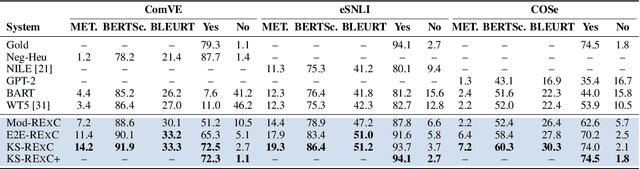
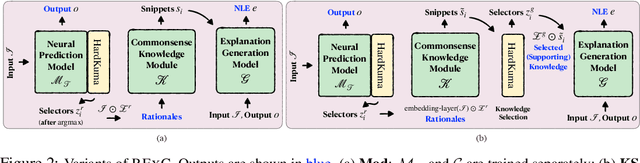
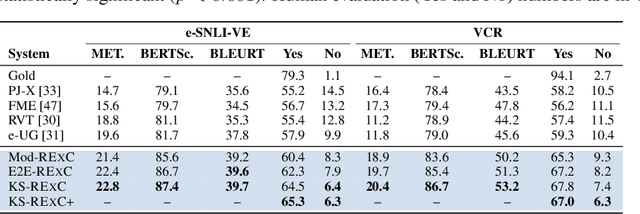
Abstract:Explainable machine learning models primarily justify predicted labels using either extractive rationales (i.e., subsets of input features) or free-text natural language explanations (NLEs) as abstractive justifications. While NLEs can be more comprehensive than extractive rationales, machine-generated NLEs have been shown to sometimes lack commonsense knowledge. Here, we show that commonsense knowledge can act as a bridge between extractive rationales and NLEs, rendering both types of explanations better. More precisely, we introduce a unified framework, called RExC (Rationale-Inspired Explanations with Commonsense), that (1) extracts rationales as a set of features responsible for machine predictions, (2) expands the extractive rationales using available commonsense resources, and (3) uses the expanded knowledge to generate natural language explanations. Our framework surpasses by a large margin the previous state-of-the-art in generating NLEs across five tasks in both natural language processing and vision-language understanding, with human annotators consistently rating the explanations generated by RExC to be more comprehensive, grounded in commonsense, and overall preferred compared to previous state-of-the-art models. Moreover, our work shows that commonsense-grounded explanations can enhance both task performance and rationales extraction capabilities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge