Tamara Broderick
Developing a Series of AI Challenges for the United States Department of the Air Force
Jul 14, 2022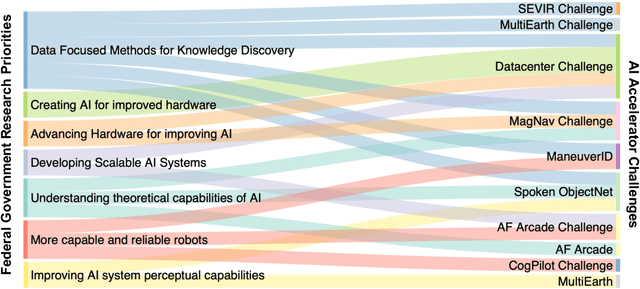
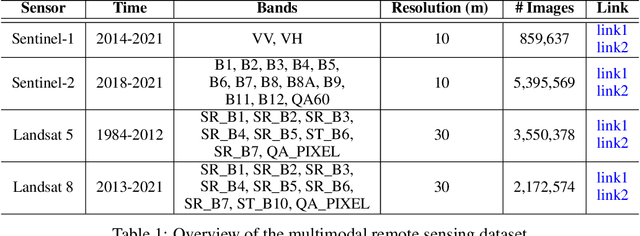

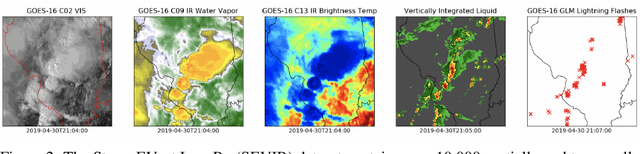
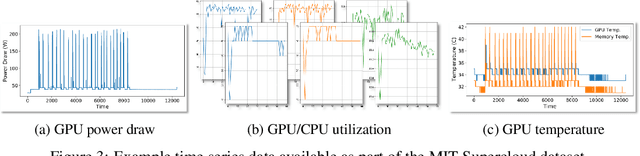
Abstract:Through a series of federal initiatives and orders, the U.S. Government has been making a concerted effort to ensure American leadership in AI. These broad strategy documents have influenced organizations such as the United States Department of the Air Force (DAF). The DAF-MIT AI Accelerator is an initiative between the DAF and MIT to bridge the gap between AI researchers and DAF mission requirements. Several projects supported by the DAF-MIT AI Accelerator are developing public challenge problems that address numerous Federal AI research priorities. These challenges target priorities by making large, AI-ready datasets publicly available, incentivizing open-source solutions, and creating a demand signal for dual use technologies that can stimulate further research. In this article, we describe these public challenges being developed and how their application contributes to scientific advances.
Diffusion probabilistic modeling of protein backbones in 3D for the motif-scaffolding problem
Jun 08, 2022



Abstract:Construction of a scaffold structure that supports a desired motif, conferring protein function, shows promise for the design of vaccines and enzymes. But a general solution to this motif-scaffolding problem remains open. Current machine-learning techniques for scaffold design are either limited to unrealistically small scaffolds (up to length 20) or struggle to produce multiple diverse scaffolds. We propose to learn a distribution over diverse and longer protein backbone structures via an E(3)-equivariant graph neural network. We develop SMCDiff to efficiently sample scaffolds from this distribution conditioned on a given motif; our algorithm is the first to theoretically guarantee conditional samples from a diffusion model in the large-compute limit. We evaluate our designed backbones by how well they align with AlphaFold2-predicted structures. We show that our method can (1) sample scaffolds up to 80 residues and (2) achieve structurally diverse scaffolds for a fixed motif.
Many processors, little time: MCMC for partitions via optimal transport couplings
Feb 23, 2022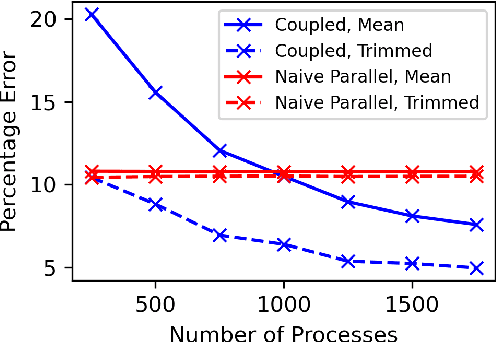
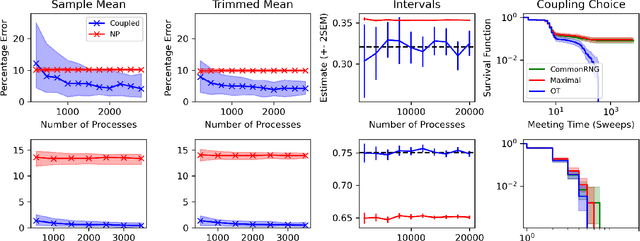
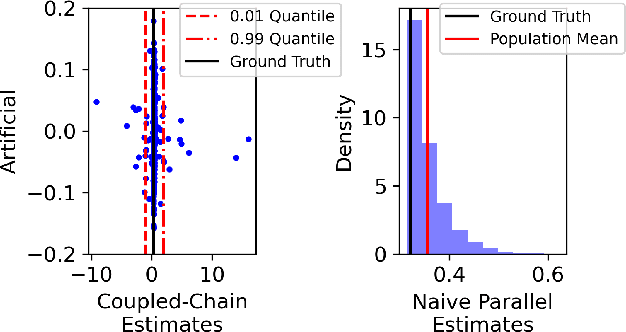
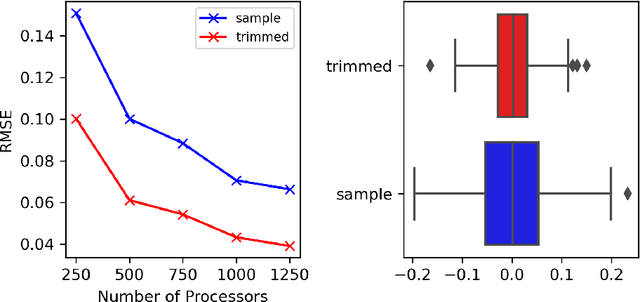
Abstract:Markov chain Monte Carlo (MCMC) methods are often used in clustering since they guarantee asymptotically exact expectations in the infinite-time limit. In finite time, though, slow mixing often leads to poor performance. Modern computing environments offer massive parallelism, but naive implementations of parallel MCMC can exhibit substantial bias. In MCMC samplers of continuous random variables, Markov chain couplings can overcome bias. But these approaches depend crucially on paired chains meetings after a small number of transitions. We show that straightforward applications of existing coupling ideas to discrete clustering variables fail to meet quickly. This failure arises from the "label-switching problem": semantically equivalent cluster relabelings impede fast meeting of coupled chains. We instead consider chains as exploring the space of partitions rather than partitions' (arbitrary) labelings. Using a metric on the partition space, we formulate a practical algorithm using optimal transport couplings. Our theory confirms our method is accurate and efficient. In experiments ranging from clustering of genes or seeds to graph colorings, we show the benefits of our coupling in the highly parallel, time-limited regime.
Toward a Taxonomy of Trust for Probabilistic Machine Learning
Dec 05, 2021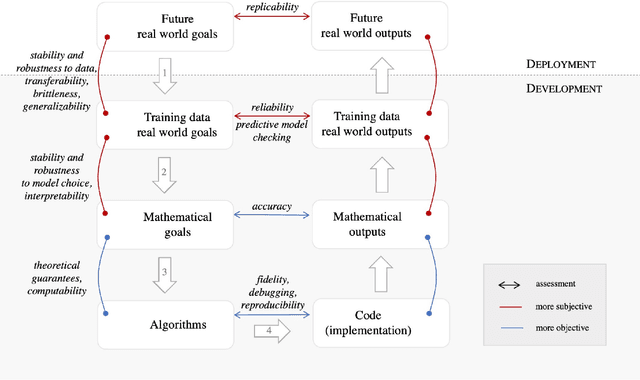
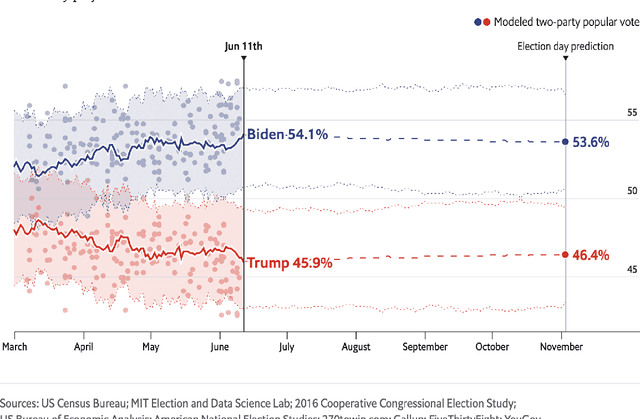
Abstract:Probabilistic machine learning increasingly informs critical decisions in medicine, economics, politics, and beyond. We need evidence to support that the resulting decisions are well-founded. To aid development of trust in these decisions, we develop a taxonomy delineating where trust in an analysis can break down: (1) in the translation of real-world goals to goals on a particular set of available training data, (2) in the translation of abstract goals on the training data to a concrete mathematical problem, (3) in the use of an algorithm to solve the stated mathematical problem, and (4) in the use of a particular code implementation of the chosen algorithm. We detail how trust can fail at each step and illustrate our taxonomy with two case studies: an analysis of the efficacy of microcredit and The Economist's predictions of the 2020 US presidential election. Finally, we describe a wide variety of methods that can be used to increase trust at each step of our taxonomy. The use of our taxonomy highlights steps where existing research work on trust tends to concentrate and also steps where establishing trust is particularly challenging.
Can we globally optimize cross-validation loss? Quasiconvexity in ridge regression
Jul 19, 2021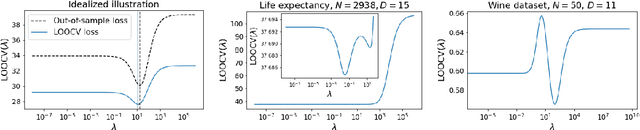
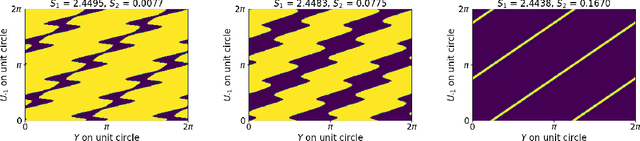
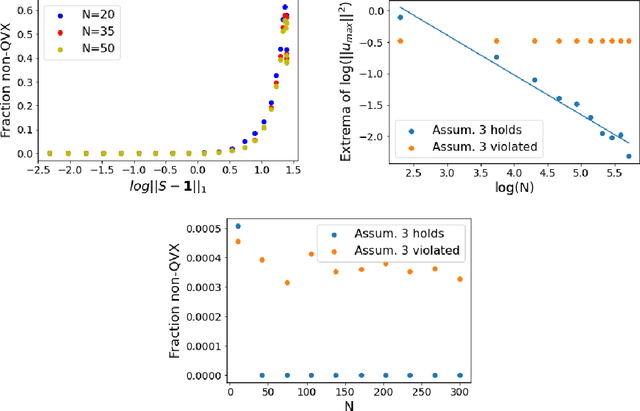
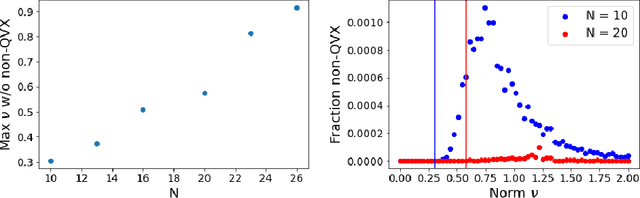
Abstract:Models like LASSO and ridge regression are extensively used in practice due to their interpretability, ease of use, and strong theoretical guarantees. Cross-validation (CV) is widely used for hyperparameter tuning in these models, but do practical optimization methods minimize the true out-of-sample loss? A recent line of research promises to show that the optimum of the CV loss matches the optimum of the out-of-sample loss (possibly after simple corrections). It remains to show how tractable it is to minimize the CV loss. In the present paper, we show that, in the case of ridge regression, the CV loss may fail to be quasiconvex and thus may have multiple local optima. We can guarantee that the CV loss is quasiconvex in at least one case: when the spectrum of the covariate matrix is nearly flat and the noise in the observed responses is not too high. More generally, we show that quasiconvexity status is independent of many properties of the observed data (response norm, covariate-matrix right singular vectors and singular-value scaling) and has a complex dependence on the few that remain. We empirically confirm our theory using simulated experiments.
For high-dimensional hierarchical models, consider exchangeability of effects across covariates instead of across datasets
Jul 13, 2021



Abstract:Hierarchical Bayesian methods enable information sharing across multiple related regression problems. While standard practice is to model regression parameters (effects) as (1) exchangeable across datasets and (2) correlated to differing degrees across covariates, we show that this approach exhibits poor statistical performance when the number of covariates exceeds the number of datasets. For instance, in statistical genetics, we might regress dozens of traits (defining datasets) for thousands of individuals (responses) on up to millions of genetic variants (covariates). When an analyst has more covariates than datasets, we argue that it is often more natural to instead model effects as (1) exchangeable across covariates and (2) correlated to differing degrees across datasets. To this end, we propose a hierarchical model expressing our alternative perspective. We devise an empirical Bayes estimator for learning the degree of correlation between datasets. We develop theory that demonstrates that our method outperforms the classic approach when the number of covariates dominates the number of datasets, and corroborate this result empirically on several high-dimensional multiple regression and classification problems.
Evaluating Sensitivity to the Stick-Breaking Prior in Bayesian Nonparametrics
Jul 12, 2021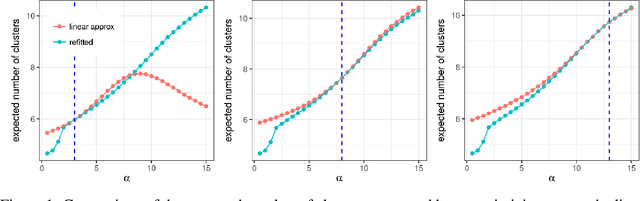
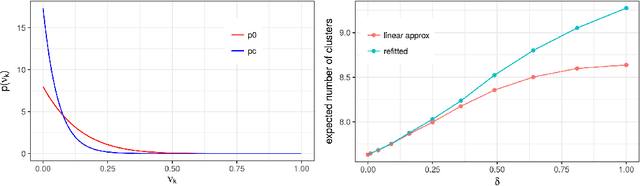
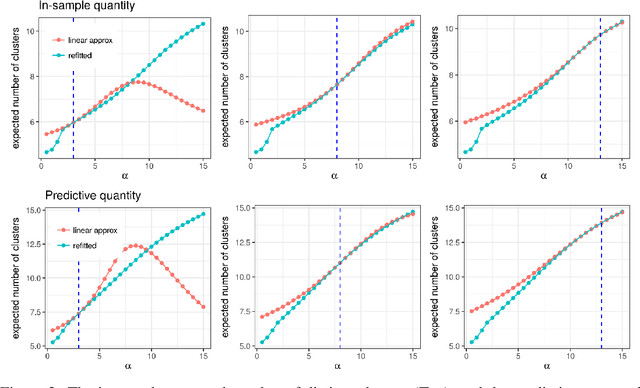
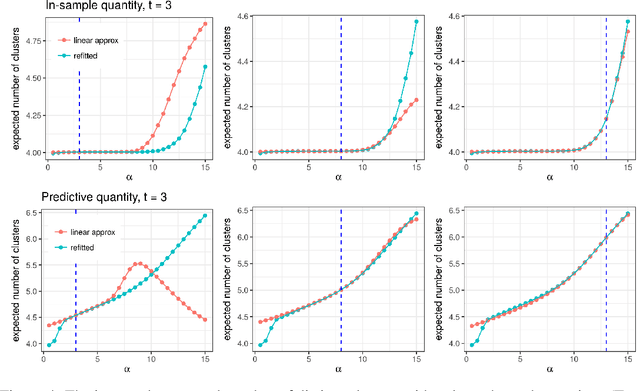
Abstract:Bayesian models based on the Dirichlet process and other stick-breaking priors have been proposed as core ingredients for clustering, topic modeling, and other unsupervised learning tasks. Prior specification is, however, relatively difficult for such models, given that their flexibility implies that the consequences of prior choices are often relatively opaque. Moreover, these choices can have a substantial effect on posterior inferences. Thus, considerations of robustness need to go hand in hand with nonparametric modeling. In the current paper, we tackle this challenge by exploiting the fact that variational Bayesian methods, in addition to having computational advantages in fitting complex nonparametric models, also yield sensitivities with respect to parametric and nonparametric aspects of Bayesian models. In particular, we demonstrate how to assess the sensitivity of conclusions to the choice of concentration parameter and stick-breaking distribution for inferences under Dirichlet process mixtures and related mixture models. We provide both theoretical and empirical support for our variational approach to Bayesian sensitivity analysis.
The SKIM-FA Kernel: High-Dimensional Variable Selection and Nonlinear Interaction Discovery in Linear Time
Jun 23, 2021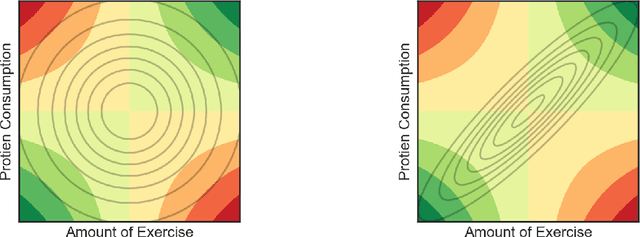
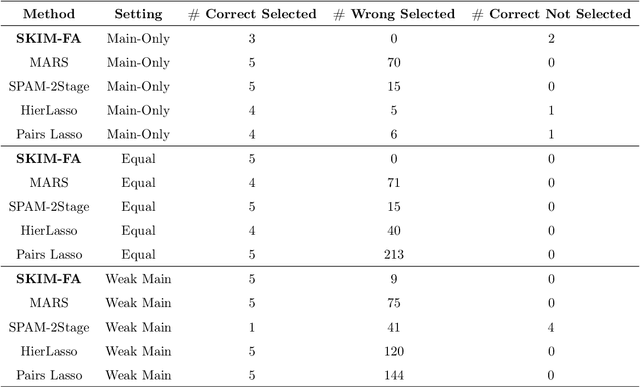
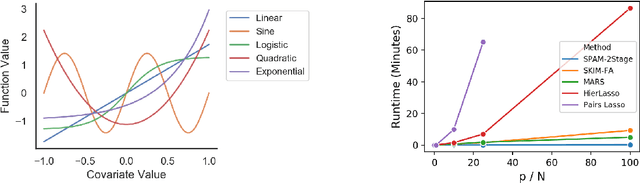
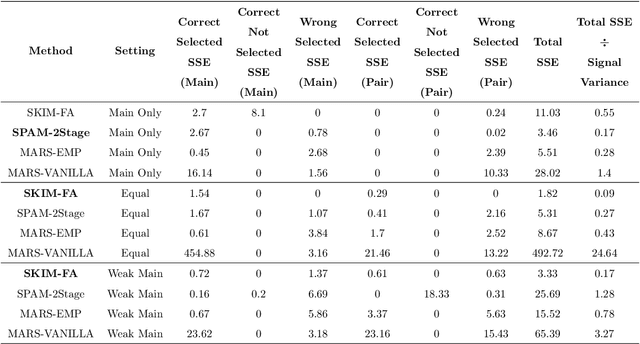
Abstract:Many scientific problems require identifying a small set of covariates that are associated with a target response and estimating their effects. Often, these effects are nonlinear and include interactions, so linear and additive methods can lead to poor estimation and variable selection. The Bayesian framework makes it straightforward to simultaneously express sparsity, nonlinearity, and interactions in a hierarchical model. But, as for the few other methods that handle this trifecta, inference is computationally intractable - with runtime at least quadratic in the number of covariates, and often worse. In the present work, we solve this computational bottleneck. We first show that suitable Bayesian models can be represented as Gaussian processes (GPs). We then demonstrate how a kernel trick can reduce computation with these GPs to O(# covariates) time for both variable selection and estimation. Our resulting fit corresponds to a sparse orthogonal decomposition of the regression function in a Hilbert space (i.e., a functional ANOVA decomposition), where interaction effects represent all variation that cannot be explained by lower-order effects. On a variety of synthetic and real datasets, our approach outperforms existing methods used for large, high-dimensional datasets while remaining competitive (or being orders of magnitude faster) in runtime.
Measuring the sensitivity of Gaussian processes to kernel choice
Jun 11, 2021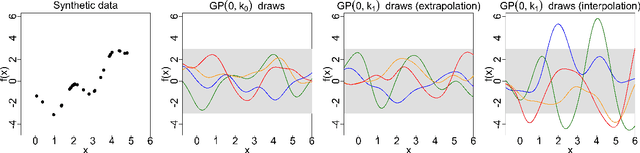
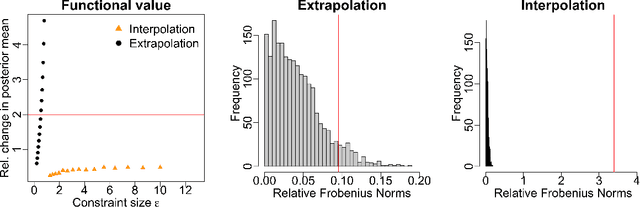
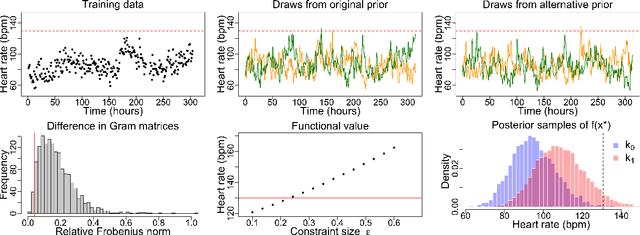
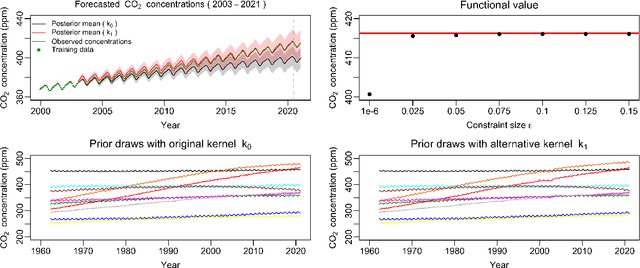
Abstract:Gaussian processes (GPs) are used to make medical and scientific decisions, including in cardiac care and monitoring of carbon dioxide emissions. But the choice of GP kernel is often somewhat arbitrary. In particular, uncountably many kernels typically align with qualitative prior knowledge (e.g. function smoothness or stationarity). But in practice, data analysts choose among a handful of convenient standard kernels (e.g. squared exponential). In the present work, we ask: Would decisions made with a GP differ under other, qualitatively interchangeable kernels? We show how to formulate this sensitivity analysis as a constrained optimization problem over a finite-dimensional space. We can then use standard optimizers to identify substantive changes in relevant decisions made with a GP. We demonstrate in both synthetic and real-world examples that decisions made with a GP can exhibit substantial sensitivity to kernel choice, even when prior draws are qualitatively interchangeable to a user.
Independent finite approximations for Bayesian nonparametric inference: construction, error bounds, and practical implications
Sep 22, 2020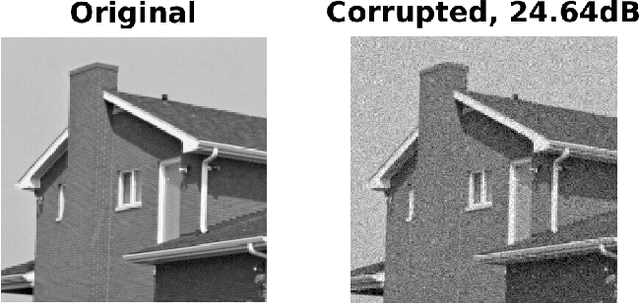
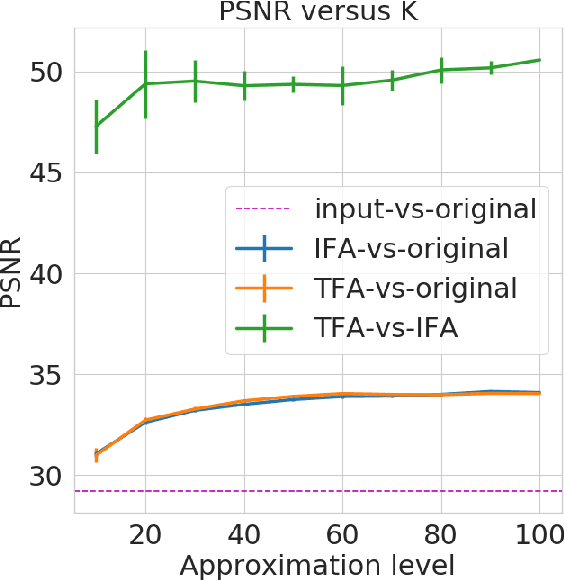
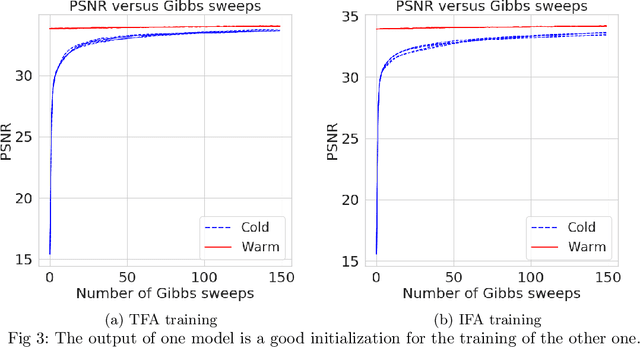
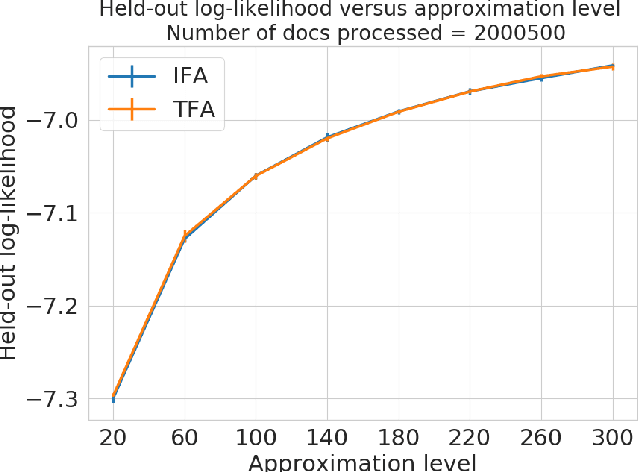
Abstract:Bayesian nonparametrics based on completely random measures (CRMs) offers a flexible modeling approach when the number of clusters or latent components in a dataset is unknown. However, managing the infinite dimensionality of CRMs often leads to slow computation. Practical inference typically relies on either integrating out the infinite-dimensional parameter or using a finite approximation: a truncated finite approximation (TFA) or an independent finite approximation (IFA). The atom weights of TFAs are constructed sequentially, while the atoms of IFAs are independent, which (1) make them well-suited for parallel and distributed computation and (2) facilitates more convenient inference schemes. While IFAs have been developed in certain special cases in the past, there has not yet been a general template for construction or a systematic comparison to TFAs. We show how to construct IFAs for approximating distributions in a large family of CRMs, encompassing all those typically used in practice. We quantify the approximation error between IFAs and the target nonparametric prior, and prove that, in the worst-case, TFAs provide more component-efficient approximations than IFAs. However, in experiments on image denoising and topic modeling tasks with real data, we find that the error of Bayesian approximation methods overwhelms any finite approximation error, and IFAs perform very similarly to TFAs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge