Songting Shi
Visualizing Data using GTSNE
Aug 03, 2021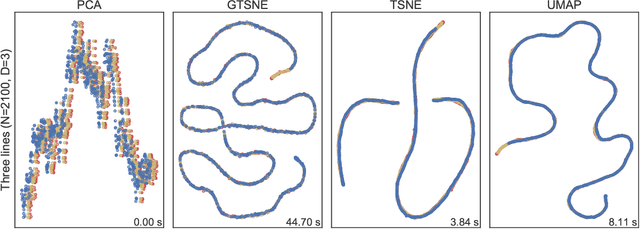

Abstract:We present a new method GTSNE to visualize high-dimensional data points in the two dimensional map. The technique is a variation of t-SNE that produces better visualizations by capturing both the local neighborhood structure and the macro structure in the data. This is particularly important for high-dimensional data that lie on continuous low-dimensional manifolds. We illustrate the performance of GTSNE on a wide variety of datasets and compare it the state of art methods, including t-SNE and UMAP. The visualizations produced by GTSNE are better than those produced by the other techniques on almost all of the datasets on the macro structure preservation.
Visualizing Data Velocity using DSNE
Mar 15, 2021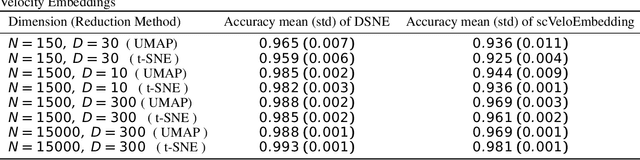
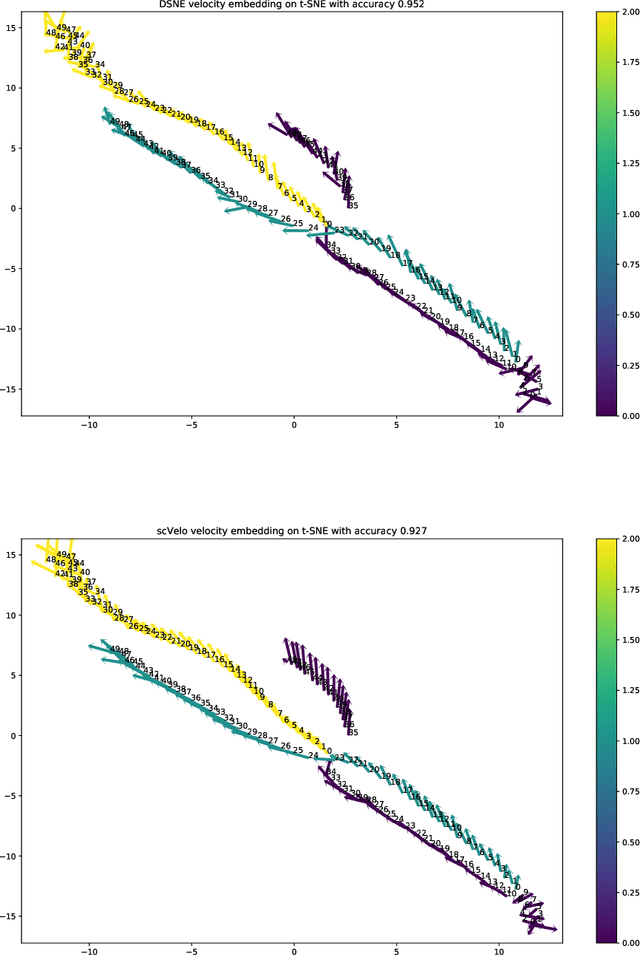
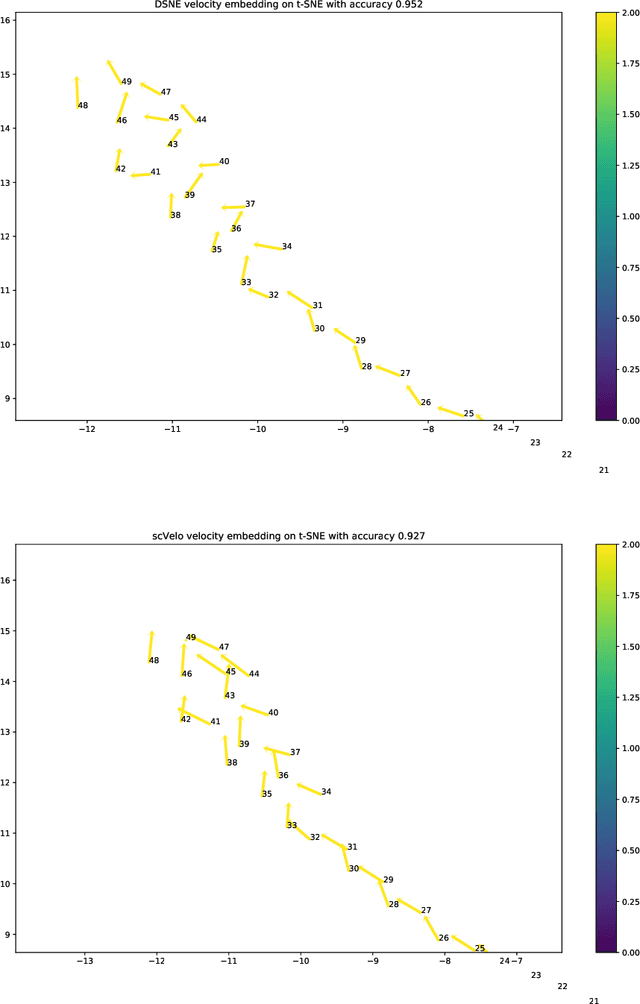
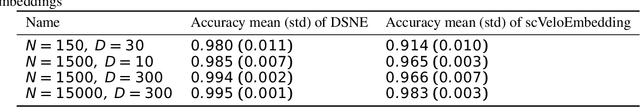
Abstract:We present a new technique called "DSNE" which learns the velocity embeddings of low dimensional map points when given the high-dimensional data points with its velocities. The technique is a variation of Stochastic Neighbor Embedding, which uses the Euclidean distance on the unit sphere between the unit-length velocity of the point and the unit-length direction from the point to its near neighbors to define similarities, and try to match the two kinds of similarities in the high dimension space and low dimension space to find the velocity embeddings on the low dimension space. DSNE can help to visualize how the data points move in the high dimension space by presenting the movements in two or three dimensions space. It is helpful for understanding the mechanism of cell differentiation and embryo development.
A Tutorial on the Mathematical Model of Single Cell Variational Inference
Jan 03, 2021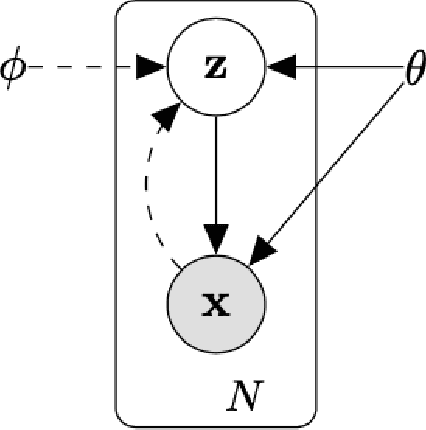
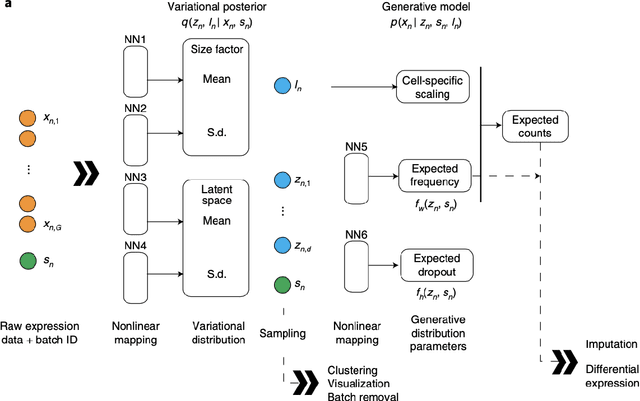
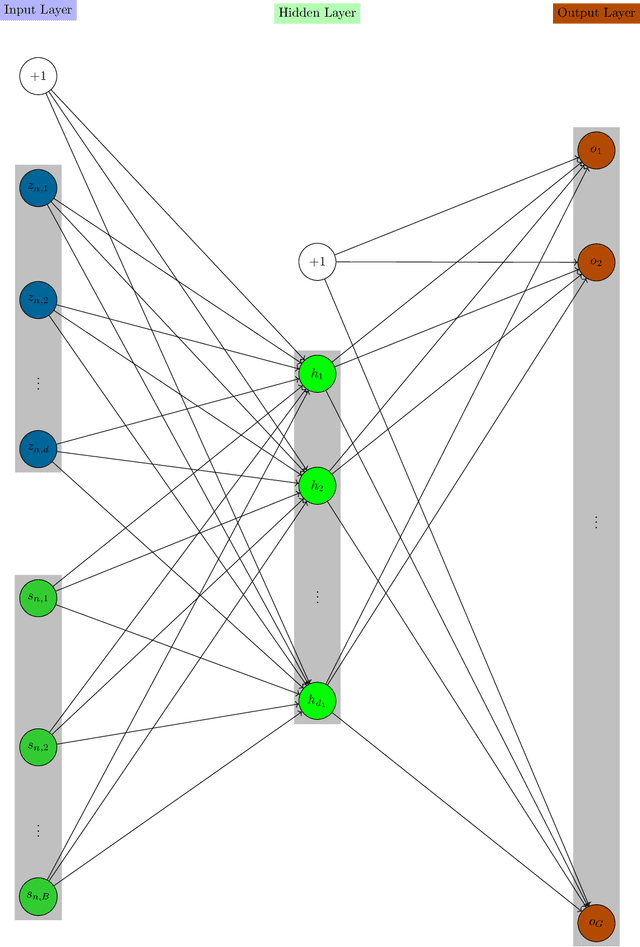
Abstract:As the large amount of sequencing data accumulated in past decades and it is still accumulating, we need to handle the more and more sequencing data. As the fast development of the computing technologies, we now can handle a large amount of data by a reasonable of time using the neural network based model. This tutorial will introduce the the mathematical model of the single cell variational inference (scVI), which use the variational auto-encoder (building on the neural networks) to learn the distribution of the data to gain insights. It was written for beginners in the simple and intuitive way with many deduction details to encourage more researchers into this field.
Learning the Sparse and Low Rank PARAFAC Decomposition via the Elastic Net
May 29, 2017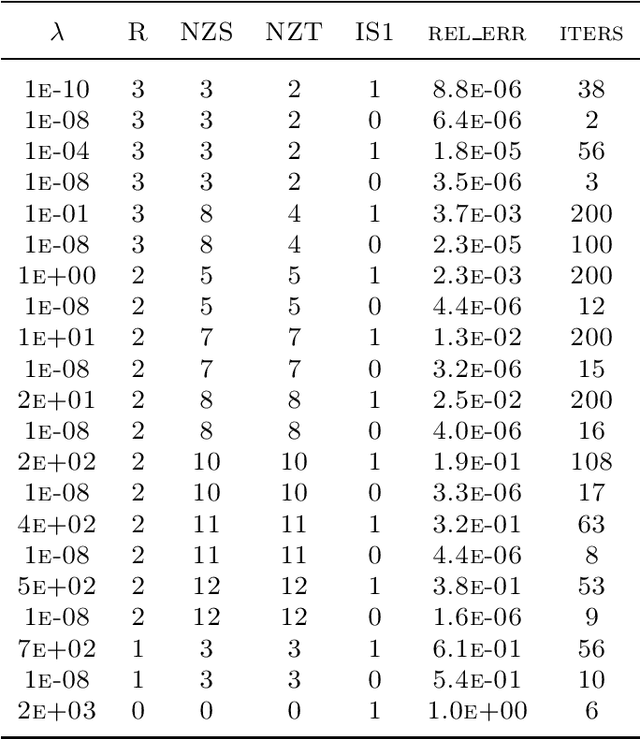
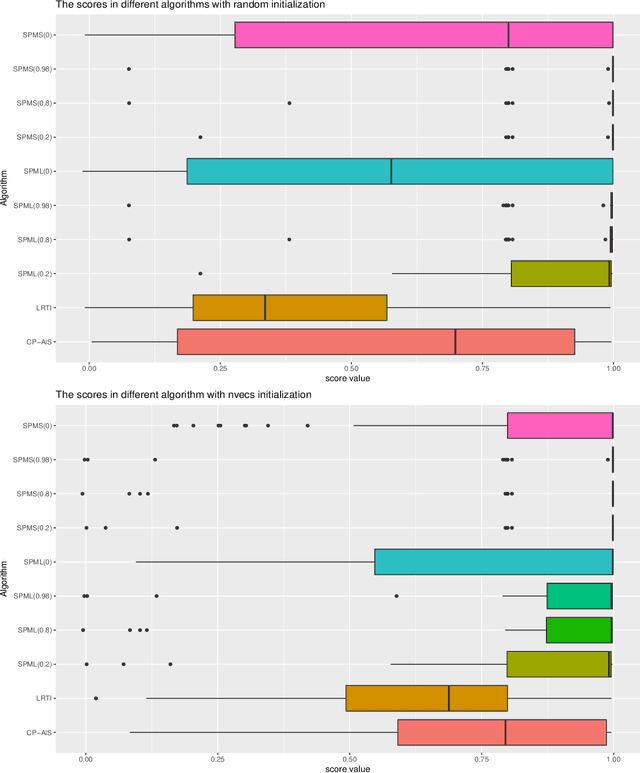
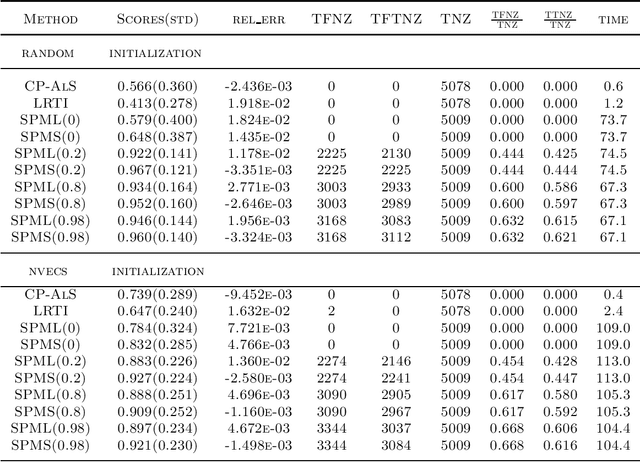
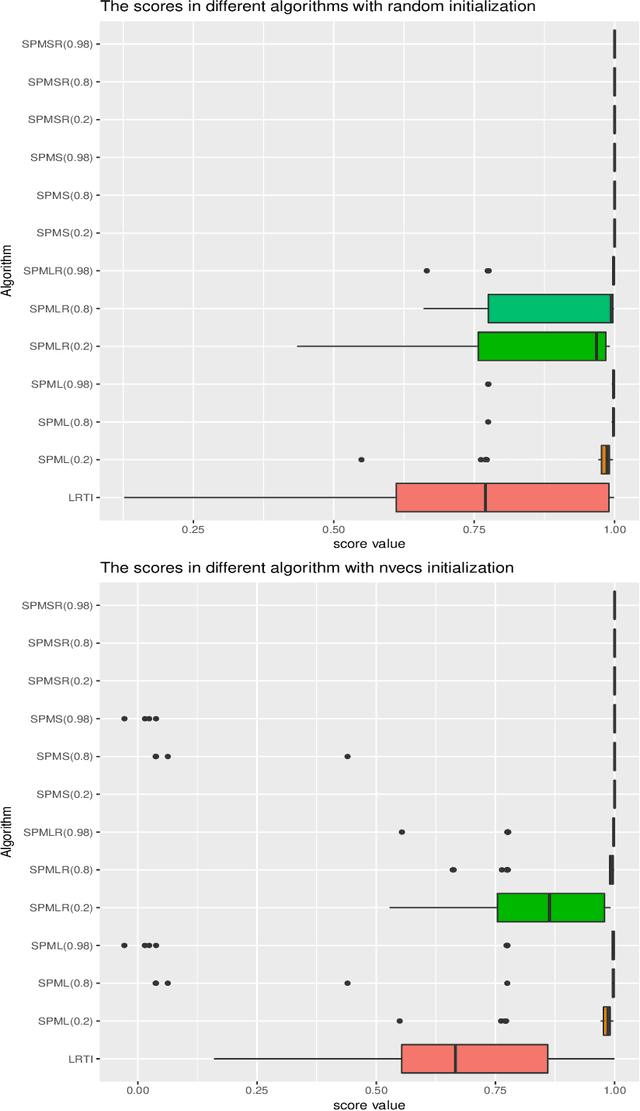
Abstract:In this article, we derive a Bayesian model to learning the sparse and low rank PARAFAC decomposition for the observed tensor with missing values via the elastic net, with property to find the true rank and sparse factor matrix which is robust to the noise. We formulate efficient block coordinate descent algorithm and admax stochastic block coordinate descent algorithm to solve it, which can be used to solve the large scale problem. To choose the appropriate rank and sparsity in PARAFAC decomposition, we will give a solution path by gradually increasing the regularization to increase the sparsity and decrease the rank. When we find the sparse structure of the factor matrix, we can fixed the sparse structure, using a small to regularization to decreasing the recovery error, and one can choose the proper decomposition from the solution path with sufficient sparse factor matrix with low recovery error. We test the power of our algorithm on the simulation data and real data, which show it is powerful.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge