Simon Bartels
A survey and benchmark of high-dimensional Bayesian optimization of discrete sequences
Jun 07, 2024Abstract:Optimizing discrete black-box functions is key in several domains, e.g. protein engineering and drug design. Due to the lack of gradient information and the need for sample efficiency, Bayesian optimization is an ideal candidate for these tasks. Several methods for high-dimensional continuous and categorical Bayesian optimization have been proposed recently. However, our survey of the field reveals highly heterogeneous experimental set-ups across methods and technical barriers for the replicability and application of published algorithms to real-world tasks. To address these issues, we develop a unified framework to test a vast array of high-dimensional Bayesian optimization methods and a collection of standardized black-box functions representing real-world application domains in chemistry and biology. These two components of the benchmark are each supported by flexible, scalable, and easily extendable software libraries (poli and poli-baselines), allowing practitioners to readily incorporate new optimization objectives or discrete optimizers. Project website: https://machinelearninglifescience.github.io/hdbo_benchmark
A Continuous Relaxation for Discrete Bayesian Optimization
Apr 26, 2024Abstract:To optimize efficiently over discrete data and with only few available target observations is a challenge in Bayesian optimization. We propose a continuous relaxation of the objective function and show that inference and optimization can be computationally tractable. We consider in particular the optimization domain where very few observations and strict budgets exist; motivated by optimizing protein sequences for expensive to evaluate bio-chemical properties. The advantages of our approach are two-fold: the problem is treated in the continuous setting, and available prior knowledge over sequences can be incorporated directly. More specifically, we utilize available and learned distributions over the problem domain for a weighting of the Hellinger distance which yields a covariance function. We show that the resulting acquisition function can be optimized with both continuous or discrete optimization algorithms and empirically assess our method on two bio-chemical sequence optimization tasks.
Adaptive Cholesky Gaussian Processes
Feb 23, 2022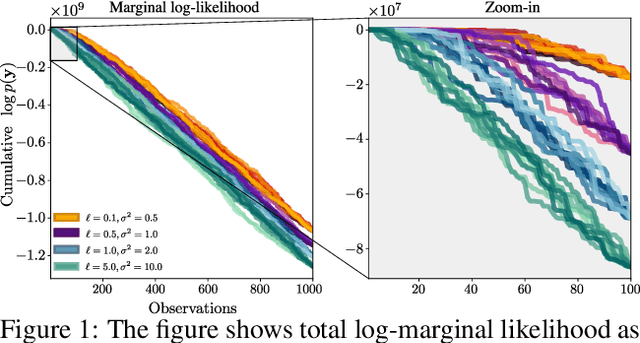
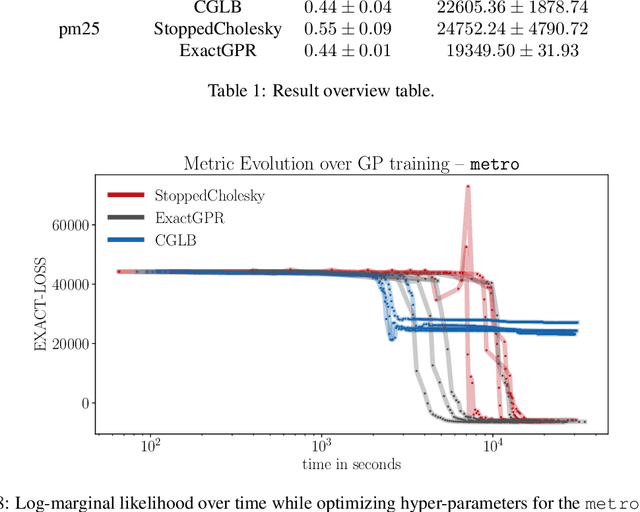
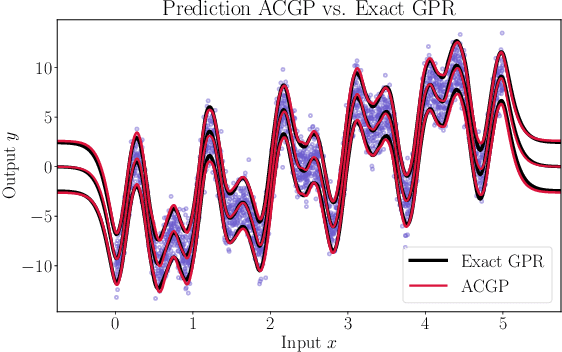
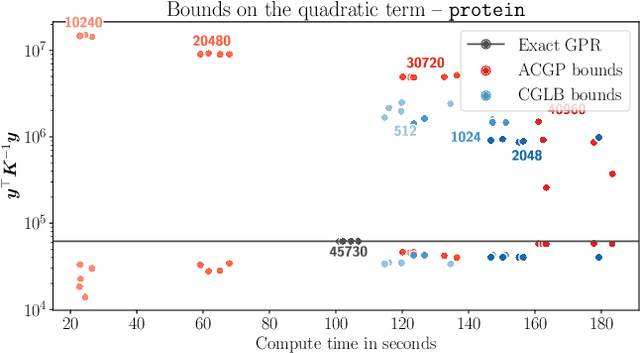
Abstract:We present a method to fit exact Gaussian process models to large datasets by considering only a subset of the data. Our approach is novel in that the size of the subset is selected on the fly during exact inference with little computational overhead. From an empirical observation that the log-marginal likelihood often exhibits a linear trend once a sufficient subset of a dataset has been observed, we conclude that many large datasets contain redundant information that only slightly affects the posterior. Based on this, we provide probabilistic bounds on the full model evidence that can identify such subsets. Remarkably, these bounds are largely composed of terms that appear in intermediate steps of the standard Cholesky decomposition, allowing us to modify the algorithm to adaptively stop the decomposition once enough data have been observed. Empirically, we show that our method can be directly plugged into well-known inference schemes to fit exact Gaussian process models to large datasets.
Conjugate Gradients for Kernel Machines
Nov 14, 2019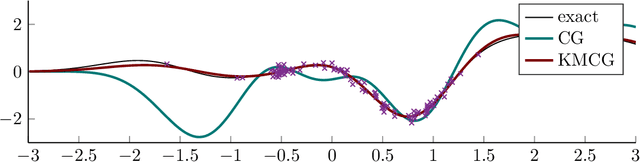
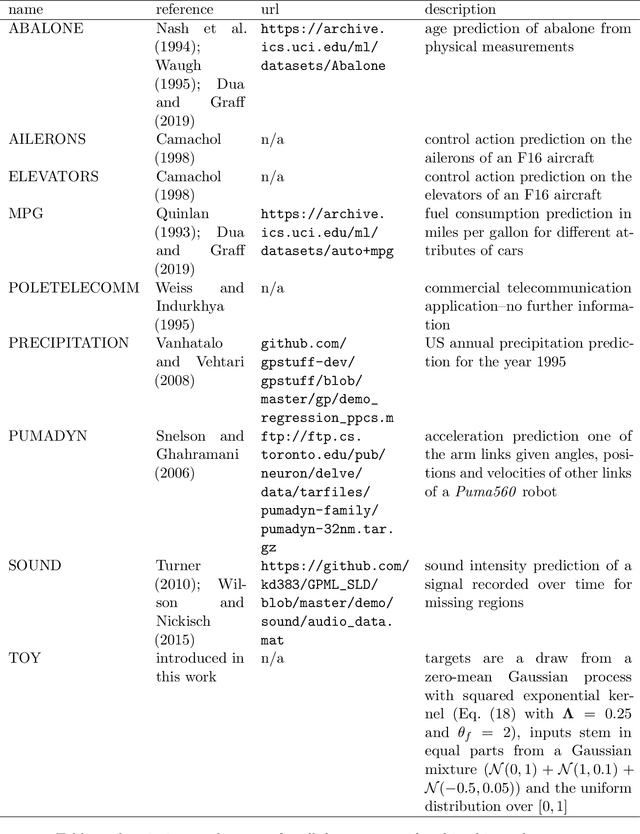
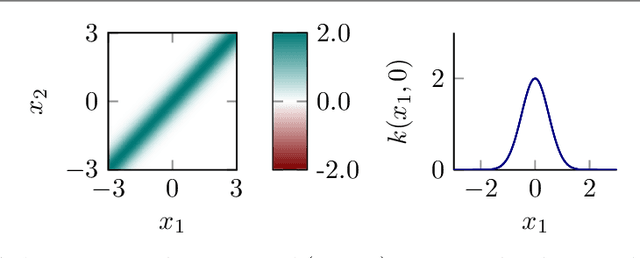
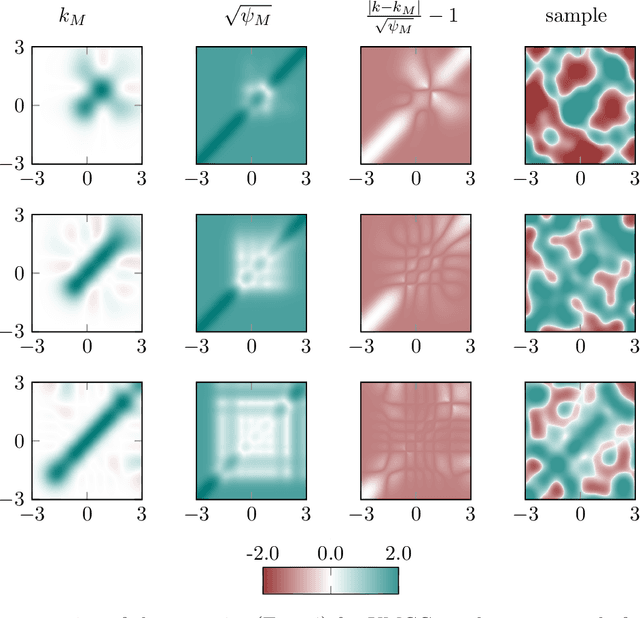
Abstract:Regularized least-squares (kernel-ridge / Gaussian process) regression is a fundamental algorithm of statistics and machine learning. Because generic algorithms for the exact solution have cubic complexity in the number of datapoints, large datasets require to resort to approximations. In this work, the computation of the least-squares prediction is itself treated as a probabilistic inference problem. We propose a structured Gaussian regression model on the kernel function that uses projections of the kernel matrix to obtain a low-rank approximation of the kernel and the matrix. A central result is an enhanced way to use the method of conjugate gradients for the specific setting of least-squares regression as encountered in machine learning. Our method improves the approximation of the kernel ridge regressor / Gaussian process posterior mean over vanilla conjugate gradients and, allows computation of the posterior variance and the log marginal likelihood (evidence) without further overhead.
Fast Bayesian Optimization of Machine Learning Hyperparameters on Large Datasets
Mar 07, 2017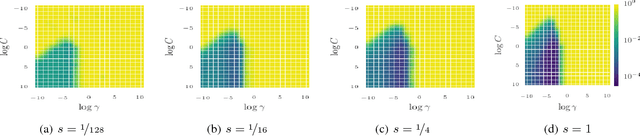
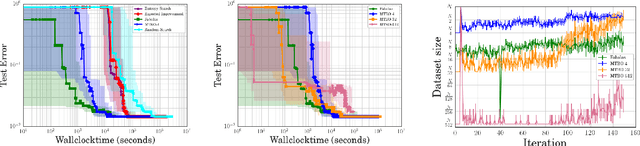
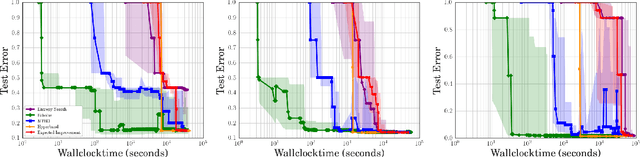
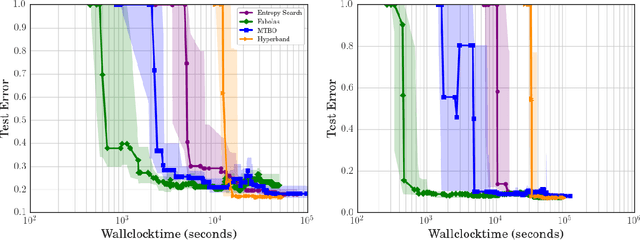
Abstract:Bayesian optimization has become a successful tool for hyperparameter optimization of machine learning algorithms, such as support vector machines or deep neural networks. Despite its success, for large datasets, training and validating a single configuration often takes hours, days, or even weeks, which limits the achievable performance. To accelerate hyperparameter optimization, we propose a generative model for the validation error as a function of training set size, which is learned during the optimization process and allows exploration of preliminary configurations on small subsets, by extrapolating to the full dataset. We construct a Bayesian optimization procedure, dubbed Fabolas, which models loss and training time as a function of dataset size and automatically trades off high information gain about the global optimum against computational cost. Experiments optimizing support vector machines and deep neural networks show that Fabolas often finds high-quality solutions 10 to 100 times faster than other state-of-the-art Bayesian optimization methods or the recently proposed bandit strategy Hyperband.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge