Serge Vasylechko
WristMIR: Coarse-to-Fine Region-Aware Retrieval of Pediatric Wrist Radiographs with Radiology Report-Driven Learning
Feb 08, 2026Abstract:Retrieving wrist radiographs with analogous fracture patterns is challenging because clinically important cues are subtle, highly localized and often obscured by overlapping anatomy or variable imaging views. Progress is further limited by the scarcity of large, well-annotated datasets for case-based medical image retrieval. We introduce WristMIR, a region-aware pediatric wrist radiograph retrieval framework that leverages dense radiology reports and bone-specific localization to learn fine-grained, clinically meaningful image representations without any manual image-level annotations. Using MedGemma-based structured report mining to generate both global and region-level captions, together with pre-processed wrist images and bone-specific crops of the distal radius, distal ulna, and ulnar styloid, WristMIR jointly trains global and local contrastive encoders and performs a two-stage retrieval process: (1) coarse global matching to identify candidate exams, followed by (2) region-conditioned reranking aligned to a predefined anatomical bone region. WristMIR improves retrieval performance over strong vision-language baselines, raising image-to-text Recall@5 from 0.82% to 9.35%. Its embeddings also yield stronger fracture classification (AUROC 0.949, AUPRC 0.953). In region-aware evaluation, the two-stage design markedly improves retrieval-based fracture diagnosis, increasing mean $F_1$ from 0.568 to 0.753, and radiologists rate its retrieved cases as more clinically relevant, with mean scores rising from 3.36 to 4.35. These findings highlight the potential of anatomically guided retrieval to enhance diagnostic reasoning and support clinical decision-making in pediatric musculoskeletal imaging. The source code is publicly available at https://github.com/quin-med-harvard-edu/WristMIR.
An Optimized Binning and Probabilistic Slice Sharing Algorithm for Motion Correction in Abdominal DW-MRI
Sep 01, 2024Abstract:Abdominal diffusion-weighted magnetic resonance imaging (DW-MRI) is a powerful, non-invasive technique for characterizing lesions and facilitating early diagnosis. However, respiratory motion during a scan can degrade image quality. Binning image slices into respiratory phases may reduce motion artifacts, but when the standard binning algorithm is applied to DW-MRI, reconstructed volumes are often incomplete because they lack slices along the superior-inferior axis. Missing slices create black stripes within images, and prolonged scan times are required to generate complete volumes. In this study, we propose a new binning algorithm to minimize missing slices. We acquired free-breathing and shallow-breathing abdominal DW-MRI scans on seven volunteers and used our algorithm to correct for motion in free-breathing scans. First, we drew the optimal rigid bin partitions in the respiratory signal using a dynamic programming approach, assigning each slice to one bin. We then designed a probabilistic approach for selecting some slices to belong in two bins. Our proposed binning algorithm resulted in significantly fewer missing slices than standard binning (p<1.0e-16), yielding an average reduction of 82.98+/-6.07%. Our algorithm also improved lesion conspicuity and reduced motion artifacts in DW-MR images and Apparent Diffusion Coefficient (ADC) maps. ADC maps created from free-breathing images corrected for motion with our algorithm showed lower intra-subject variability compared to uncorrected free-breathing and shallow-breathing maps (p<0.001). Additionally, shallow-breathing ADC maps showed more consistency with corrected free-breathing maps rather than uncorrected free-breathing maps (p<0.01). Our proposed binning algorithm's efficacy in reducing missing slices increases anatomical accuracy and allows for shorter acquisition times compared to standard binning.
Convolution-Free Medical Image Segmentation using Transformers
Feb 26, 2021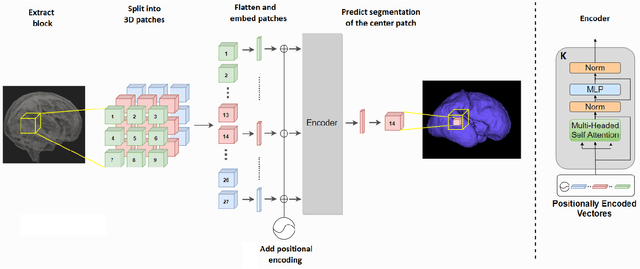
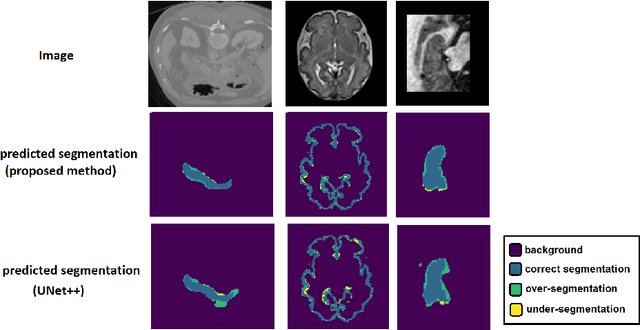
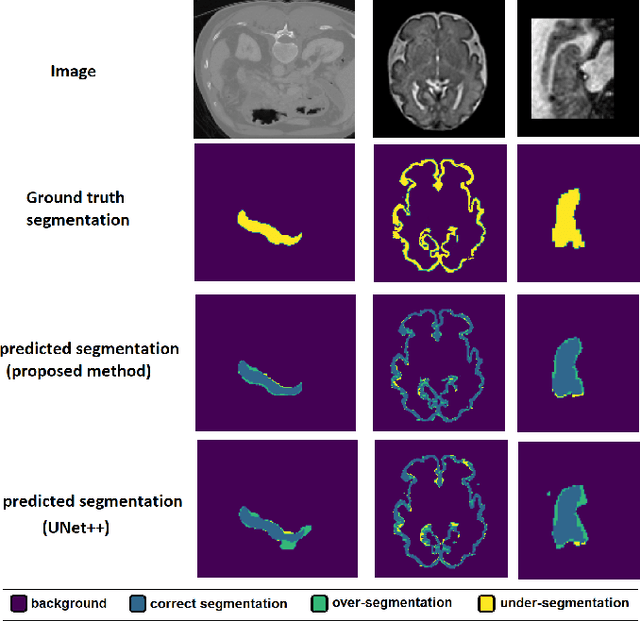
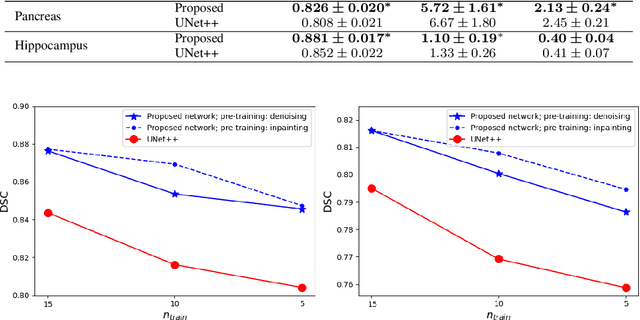
Abstract:Like other applications in computer vision, medical image segmentation has been most successfully addressed using deep learning models that rely on the convolution operation as their main building block. Convolutions enjoy important properties such as sparse interactions, weight sharing, and translation equivariance. These properties give convolutional neural networks (CNNs) a strong and useful inductive bias for vision tasks. In this work we show that a different method, based entirely on self-attention between neighboring image patches and without any convolution operations, can achieve competitive or better results. Given a 3D image block, our network divides it into $n^3$ 3D patches, where $n=3 \text{ or } 5$ and computes a 1D embedding for each patch. The network predicts the segmentation map for the center patch of the block based on the self-attention between these patch embeddings. We show that the proposed model can achieve segmentation accuracies that are better than the state of the art CNNs on three datasets. We also propose methods for pre-training this model on large corpora of unlabeled images. Our experiments show that with pre-training the advantage of our proposed network over CNNs can be significant when labeled training data is small.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge