Scott Raymond
Autonomous Skeletal Landmark Localization towards Agentic C-Arm Control
Apr 20, 2026Abstract:Purpose: Automated C-arm positioning ensures timely treatment in patients requiring emergent interventions. When a conventional Deep Learning (DL) approach for C-arm control fails, clinicians must revert to manual operation, resulting in additional delays. Consequently, an agentic C-arm control framework based on multimodal large language models (MLLMs) is highly desirable, as it can incorporate clinician feedback and use reasoning to make adjustments toward more accurate positioning. Skeletal landmark localization is essential for C-arm control, and we investigate adapting MLLMs for autonomous landmark localization. Methods: We used an annotated synthetic X-ray dataset and a real X-ray dataset. Each X-ray in both datasets is paired with several skeletal landmarks. We fine-tuned two MLLMs and tasked them with retrieving the closest landmarks from each X-ray. Quantitative evaluations of landmark localization were performed and compared against a leading DL approach. We further conducted qualitative experiments demonstrating: (1) how an MLLM can correct an initially incorrect prediction through reasoning, and (2) how the MLLM can sequentially navigate the C-arm toward a target location. Results: On both datasets, fine-tuned MLLMs demonstrate competitive performance across all localization tasks when compared with the DL approach. In the qualitative experiments, the MLLMs provide evidence of reasoning and spatial awareness. Conclusion: This study shows that fine-tuned MLLMs achieve accurate skeletal landmark localization and hold promise for agentic autonomous C-arm control. Our code is available athttps://github.com/marszzibros/C-arm-localization-LLMs.git
Path and Bone-Contour Regularized Unpaired MRI-to-CT Translation
May 06, 2025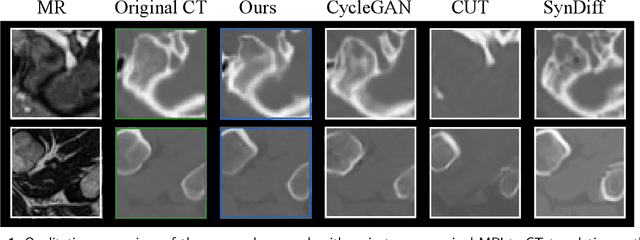
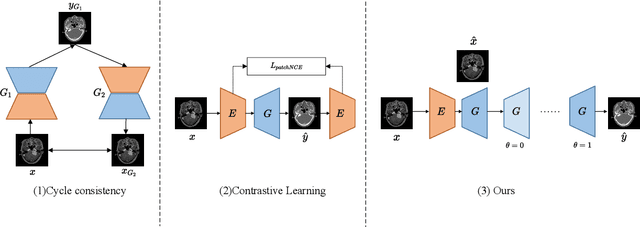
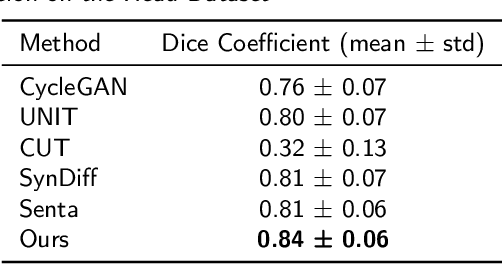
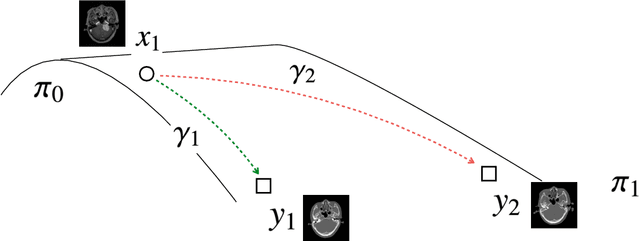
Abstract:Accurate MRI-to-CT translation promises the integration of complementary imaging information without the need for additional imaging sessions. Given the practical challenges associated with acquiring paired MRI and CT scans, the development of robust methods capable of leveraging unpaired datasets is essential for advancing the MRI-to-CT translation. Current unpaired MRI-to-CT translation methods, which predominantly rely on cycle consistency and contrastive learning frameworks, frequently encounter challenges in accurately translating anatomical features that are highly discernible on CT but less distinguishable on MRI, such as bone structures. This limitation renders these approaches less suitable for applications in radiation therapy, where precise bone representation is essential for accurate treatment planning. To address this challenge, we propose a path- and bone-contour regularized approach for unpaired MRI-to-CT translation. In our method, MRI and CT images are projected to a shared latent space, where the MRI-to-CT mapping is modeled as a continuous flow governed by neural ordinary differential equations. The optimal mapping is obtained by minimizing the transition path length of the flow. To enhance the accuracy of translated bone structures, we introduce a trainable neural network to generate bone contours from MRI and implement mechanisms to directly and indirectly encourage the model to focus on bone contours and their adjacent regions. Evaluations conducted on three datasets demonstrate that our method outperforms existing unpaired MRI-to-CT translation approaches, achieving lower overall error rates. Moreover, in a downstream bone segmentation task, our approach exhibits superior performance in preserving the fidelity of bone structures. Our code is available at: https://github.com/kennysyp/PaBoT.
Analysis of the MICCAI Brain Tumor Segmentation -- Metastases (BraTS-METS) 2025 Lighthouse Challenge: Brain Metastasis Segmentation on Pre- and Post-treatment MRI
Apr 16, 2025Abstract:Despite continuous advancements in cancer treatment, brain metastatic disease remains a significant complication of primary cancer and is associated with an unfavorable prognosis. One approach for improving diagnosis, management, and outcomes is to implement algorithms based on artificial intelligence for the automated segmentation of both pre- and post-treatment MRI brain images. Such algorithms rely on volumetric criteria for lesion identification and treatment response assessment, which are still not available in clinical practice. Therefore, it is critical to establish tools for rapid volumetric segmentations methods that can be translated to clinical practice and that are trained on high quality annotated data. The BraTS-METS 2025 Lighthouse Challenge aims to address this critical need by establishing inter-rater and intra-rater variability in dataset annotation by generating high quality annotated datasets from four individual instances of segmentation by neuroradiologists while being recorded on video (two instances doing "from scratch" and two instances after AI pre-segmentation). This high-quality annotated dataset will be used for testing phase in 2025 Lighthouse challenge and will be publicly released at the completion of the challenge. The 2025 Lighthouse challenge will also release the 2023 and 2024 segmented datasets that were annotated using an established pipeline of pre-segmentation, student annotation, two neuroradiologists checking, and one neuroradiologist finalizing the process. It builds upon its previous edition by including post-treatment cases in the dataset. Using these high-quality annotated datasets, the 2025 Lighthouse challenge plans to test benchmark algorithms for automated segmentation of pre-and post-treatment brain metastases (BM), trained on diverse and multi-institutional datasets of MRI images obtained from patients with brain metastases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge