Scott Floyd
Analysis of the MICCAI Brain Tumor Segmentation -- Metastases (BraTS-METS) 2025 Lighthouse Challenge: Brain Metastasis Segmentation on Pre- and Post-treatment MRI
Apr 16, 2025Abstract:Despite continuous advancements in cancer treatment, brain metastatic disease remains a significant complication of primary cancer and is associated with an unfavorable prognosis. One approach for improving diagnosis, management, and outcomes is to implement algorithms based on artificial intelligence for the automated segmentation of both pre- and post-treatment MRI brain images. Such algorithms rely on volumetric criteria for lesion identification and treatment response assessment, which are still not available in clinical practice. Therefore, it is critical to establish tools for rapid volumetric segmentations methods that can be translated to clinical practice and that are trained on high quality annotated data. The BraTS-METS 2025 Lighthouse Challenge aims to address this critical need by establishing inter-rater and intra-rater variability in dataset annotation by generating high quality annotated datasets from four individual instances of segmentation by neuroradiologists while being recorded on video (two instances doing "from scratch" and two instances after AI pre-segmentation). This high-quality annotated dataset will be used for testing phase in 2025 Lighthouse challenge and will be publicly released at the completion of the challenge. The 2025 Lighthouse challenge will also release the 2023 and 2024 segmented datasets that were annotated using an established pipeline of pre-segmentation, student annotation, two neuroradiologists checking, and one neuroradiologist finalizing the process. It builds upon its previous edition by including post-treatment cases in the dataset. Using these high-quality annotated datasets, the 2025 Lighthouse challenge plans to test benchmark algorithms for automated segmentation of pre-and post-treatment brain metastases (BM), trained on diverse and multi-institutional datasets of MRI images obtained from patients with brain metastases.
Brain Tumor Segmentation (BraTS) Challenge 2024: Meningioma Radiotherapy Planning Automated Segmentation
May 28, 2024Abstract:The 2024 Brain Tumor Segmentation Meningioma Radiotherapy (BraTS-MEN-RT) challenge aims to advance automated segmentation algorithms using the largest known multi-institutional dataset of radiotherapy planning brain MRIs with expert-annotated target labels for patients with intact or post-operative meningioma that underwent either conventional external beam radiotherapy or stereotactic radiosurgery. Each case includes a defaced 3D post-contrast T1-weighted radiotherapy planning MRI in its native acquisition space, accompanied by a single-label "target volume" representing the gross tumor volume (GTV) and any at-risk post-operative site. Target volume annotations adhere to established radiotherapy planning protocols, ensuring consistency across cases and institutions. For pre-operative meningiomas, the target volume encompasses the entire GTV and associated nodular dural tail, while for post-operative cases, it includes at-risk resection cavity margins as determined by the treating institution. Case annotations were reviewed and approved by expert neuroradiologists and radiation oncologists. Participating teams will develop, containerize, and evaluate automated segmentation models using this comprehensive dataset. Model performance will be assessed using the lesion-wise Dice Similarity Coefficient and the 95% Hausdorff distance. The top-performing teams will be recognized at the Medical Image Computing and Computer Assisted Intervention Conference in October 2024. BraTS-MEN-RT is expected to significantly advance automated radiotherapy planning by enabling precise tumor segmentation and facilitating tailored treatment, ultimately improving patient outcomes.
Analysis of the BraTS 2023 Intracranial Meningioma Segmentation Challenge
May 16, 2024



Abstract:We describe the design and results from the BraTS 2023 Intracranial Meningioma Segmentation Challenge. The BraTS Meningioma Challenge differed from prior BraTS Glioma challenges in that it focused on meningiomas, which are typically benign extra-axial tumors with diverse radiologic and anatomical presentation and a propensity for multiplicity. Nine participating teams each developed deep-learning automated segmentation models using image data from the largest multi-institutional systematically expert annotated multilabel multi-sequence meningioma MRI dataset to date, which included 1000 training set cases, 141 validation set cases, and 283 hidden test set cases. Each case included T2, T2/FLAIR, T1, and T1Gd brain MRI sequences with associated tumor compartment labels delineating enhancing tumor, non-enhancing tumor, and surrounding non-enhancing T2/FLAIR hyperintensity. Participant automated segmentation models were evaluated and ranked based on a scoring system evaluating lesion-wise metrics including dice similarity coefficient (DSC) and 95% Hausdorff Distance. The top ranked team had a lesion-wise median dice similarity coefficient (DSC) of 0.976, 0.976, and 0.964 for enhancing tumor, tumor core, and whole tumor, respectively and a corresponding average DSC of 0.899, 0.904, and 0.871, respectively. These results serve as state-of-the-art benchmarks for future pre-operative meningioma automated segmentation algorithms. Additionally, we found that 1286 of 1424 cases (90.3%) had at least 1 compartment voxel abutting the edge of the skull-stripped image edge, which requires further investigation into optimal pre-processing face anonymization steps.
A personalized Uncertainty Quantification framework for patient survival models: estimating individual uncertainty of patients with metastatic brain tumors in the absence of ground truth
Nov 28, 2023



Abstract:TodevelopanovelUncertaintyQuantification (UQ) framework to estimate the uncertainty of patient survival models in the absence of ground truth, we developed and evaluated our approach based on a dataset of 1383 patients treated with stereotactic radiosurgery (SRS) for brain metastases between January 2015 and December 2020. Our motivating hypothesis is that a time-to-event prediction of a test patient on inference is more certain given a higher feature-space-similarity to patients in the training set. Therefore, the uncertainty for a particular patient-of-interest is represented by the concordance index between a patient similarity rank and a prediction similarity rank. Model uncertainty was defined as the increased percentage of the max uncertainty-constrained-AUC compared to the model AUC. We evaluated our method on multiple clinically-relevant endpoints, including time to intracranial progression (ICP), progression-free survival (PFS) after SRS, overall survival (OS), and time to ICP and/or death (ICPD), on a variety of both statistical and non-statistical models, including CoxPH, conditional survival forest (CSF), and neural multi-task linear regression (NMTLR). Our results show that all models had the lowest uncertainty on ICP (2.21%) and the highest uncertainty (17.28%) on ICPD. OS models demonstrated high variation in uncertainty performance, where NMTLR had the lowest uncertainty(1.96%)and CSF had the highest uncertainty (14.29%). In conclusion, our method can estimate the uncertainty of individual patient survival modeling results. As expected, our data empirically demonstrate that as model uncertainty measured via our technique increases, the similarity between a feature-space and its predicted outcome decreases.
A Neural Ordinary Differential Equation Model for Visualizing Deep Neural Network Behaviors in Multi-Parametric MRI based Glioma Segmentation
Mar 23, 2022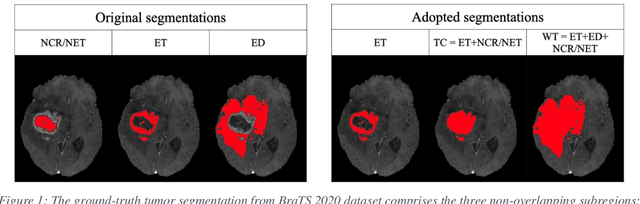
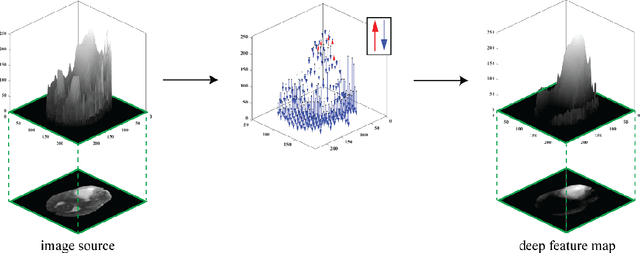
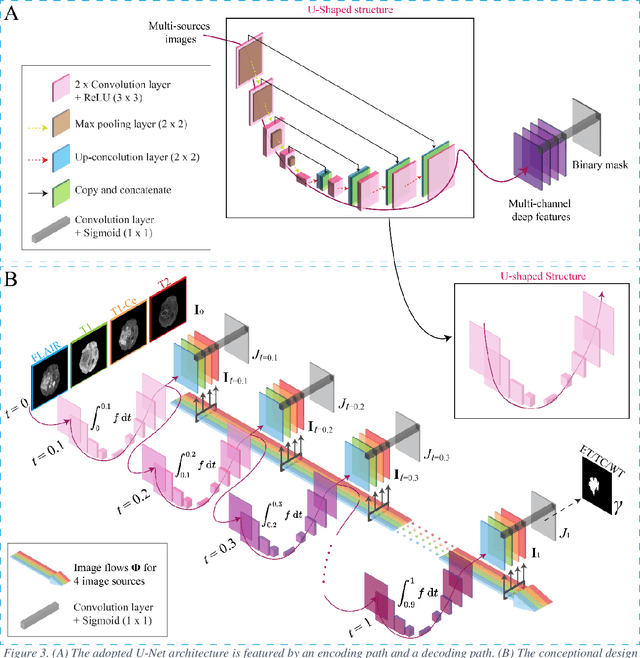
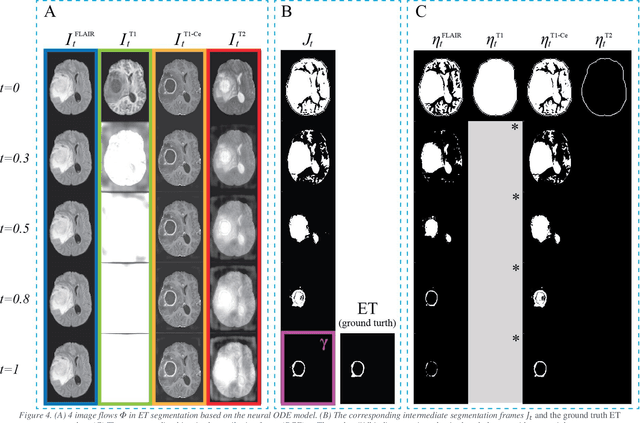
Abstract:Purpose: To develop a neural ordinary differential equation (ODE) model for visualizing deep neural network (DNN) behavior during multi-parametric MRI (mp-MRI) based glioma segmentation as a method to enhance deep learning explainability. Methods: By hypothesizing that deep feature extraction can be modeled as a spatiotemporally continuous process, we designed a novel deep learning model, neural ODE, in which deep feature extraction was governed by an ODE without explicit expression. The dynamics of 1) MR images after interactions with DNN and 2) segmentation formation can be visualized after solving ODE. An accumulative contribution curve (ACC) was designed to quantitatively evaluate the utilization of each MRI by DNN towards the final segmentation results. The proposed neural ODE model was demonstrated using 369 glioma patients with a 4-modality mp-MRI protocol: T1, contrast-enhanced T1 (T1-Ce), T2, and FLAIR. Three neural ODE models were trained to segment enhancing tumor (ET), tumor core (TC), and whole tumor (WT). The key MR modalities with significant utilization by DNN were identified based on ACC analysis. Segmentation results by DNN using only the key MR modalities were compared to the ones using all 4 MR modalities. Results: All neural ODE models successfully illustrated image dynamics as expected. ACC analysis identified T1-Ce as the only key modality in ET and TC segmentations, while both FLAIR and T2 were key modalities in WT segmentation. Compared to the U-Net results using all 4 MR modalities, Dice coefficient of ET (0.784->0.775), TC (0.760->0.758), and WT (0.841->0.837) using the key modalities only had minimal differences without significance. Conclusion: The neural ODE model offers a new tool for optimizing the deep learning model inputs with enhanced explainability. The presented methodology can be generalized to other medical image-related deep learning applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge