Sanand Sasidharan
Negation is Not Semantic: Diagnosing Dense Retrieval Failure Modes for Trade-offs in Contradiction-Aware Biomedical QA
Mar 18, 2026Abstract:Large Language Models (LLMs) have demonstrated strong capabilities in biomedical question answering, yet their tendency to generate plausible but unverified claims poses serious risks in clinical settings. To mitigate these risks, the TREC 2025 BioGen track mandates grounded answers that explicitly surface contradictory evidence (Task A) and the generation of narrative driven, fully attributed responses (Task B). Addressing the absence of target ground truth, we present a proxy-based development framework using the SciFact dataset to systematically optimize retrieval architectures. Our iterative evaluation revealed a "Simplicity Paradox": complex adversarial dense retrieval strategies failed catastrophically at contradiction detection (MRR 0.023) due to Semantic Collapse, where negation signals become indistinguishable in vector space. We further identify a Retrieval Asymmetry: filtering dense embeddings improves contradiction detection but degrades support recall, compromising reliability. We resolve this via a Decoupled Lexical Architecture built on a unified BM25 backbone, balancing semantic support recall (0.810) with precise contradiction surfacing (0.750). This approach achieves the highest Weighted MRR (0.790) on the proxy benchmark while remaining the only viable strategy for scaling to the 30 million document PubMed corpus. For answer generation, we introduce Narrative Aware Reranking and One-Shot In-Context Learning, improving citation coverage from 50% (zero-shot) to 100%. Official TREC results confirm our findings: our system ranks 2nd on Task A contradiction F1 and 3rd out of 50 runs on Task B citation coverage (98.77%), achieving zero citation contradict rate. Our work transforms LLMs from stochastic generators into honest evidence synthesizers, showing that epistemic integrity in biomedical AI requires precision and architectural scalability isolated metric optimization.
Efficient Biomedical Entity Linking: Clinical Text Standardization with Low-Resource Techniques
May 27, 2024Abstract:Clinical text is rich in information, with mentions of treatment, medication and anatomy among many other clinical terms. Multiple terms can refer to the same core concepts which can be referred as a clinical entity. Ontologies like the Unified Medical Language System (UMLS) are developed and maintained to store millions of clinical entities including the definitions, relations and other corresponding information. These ontologies are used for standardization of clinical text by normalizing varying surface forms of a clinical term through Biomedical entity linking. With the introduction of transformer-based language models, there has been significant progress in Biomedical entity linking. In this work, we focus on learning through synonym pairs associated with the entities. As compared to the existing approaches, our approach significantly reduces the training data and resource consumption. Moreover, we propose a suite of context-based and context-less reranking techniques for performing the entity disambiguation. Overall, we achieve similar performance to the state-of-the-art zero-shot and distant supervised entity linking techniques on the Medmentions dataset, the largest annotated dataset on UMLS, without any domain-based training. Finally, we show that retrieval performance alone might not be sufficient as an evaluation metric and introduce an article level quantitative and qualitative analysis to reveal further insights on the performance of entity linking methods.
Double Ramp Loss Based Reject Option Classifier
Dec 08, 2014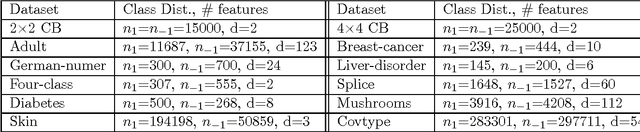
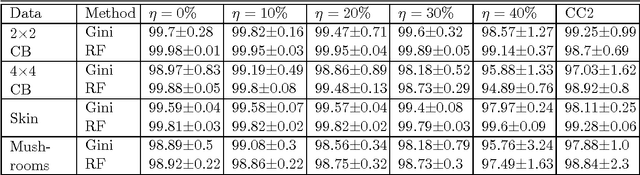
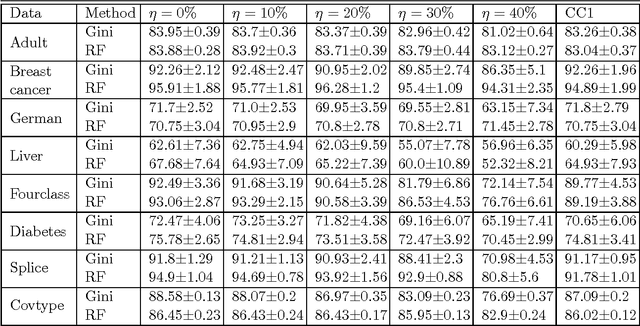
Abstract:We consider the problem of learning reject option classifiers. The goodness of a reject option classifier is quantified using $0-d-1$ loss function wherein a loss $d \in (0,.5)$ is assigned for rejection. In this paper, we propose {\em double ramp loss} function which gives a continuous upper bound for $(0-d-1)$ loss. Our approach is based on minimizing regularized risk under the double ramp loss using {\em difference of convex (DC) programming}. We show the effectiveness of our approach through experiments on synthetic and benchmark datasets. Our approach performs better than the state of the art reject option classification approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge