Rita Fioresi
Flow Matching on Symmetric Spaces
May 05, 2026Abstract:We introduce a general framework for training flow matching models on Riemannian symmetric spaces, a large class of manifolds that includes the sphere, hyperbolic space and Grassmannians. We exploit their algebraic structure to reformulate flow matching on symmetric spaces as flow matching on a subspace of the Lie algebra of their isometry group, thus linearizing the problem and greatly simplifying the handling of geodesics. As an application, we showcase our framework on the real Grassmannians $\operatorname{SO}(n) / \operatorname{SO}(k) \times \operatorname{SO}(n-k)$.
Sinkhorn doubly stochastic attention rank decay analysis
Apr 09, 2026Abstract:The self-attention mechanism is central to the success of Transformer architectures. However, standard row-stochastic attention has been shown to suffer from significant signal degradation across layers. In particular, it can induce rank collapse, resulting in increasingly uniform token representations, as well as entropy collapse, characterized by highly concentrated attention distributions. Recent work has highlighted the benefits of doubly stochastic attention as a form of entropy regularization, promoting a more balanced attention distribution and leading to improved empirical performance. In this paper, we study rank collapse across network depth and show that doubly stochastic attention matrices normalized with Sinkhorn algorithm preserve rank more effectively than standard Softmax row-stochastic ones. As previously shown for Softmax, skip connections are crucial to mitigate rank collapse. We empirically validate this phenomenon on both sentiment analysis and image classification tasks. Moreover, we derive a theoretical bound for the pure self-attention rank decay when using Sinkhorn normalization and find that rank decays to one doubly exponentially with depth, a phenomenon that has already been shown for Softmax.
A Multi-Label Temporal Convolutional Framework for Transcription Factor Binding Characterization
Mar 12, 2026Abstract:Transcription factors (TFs) regulate gene expression through complex and co-operative mechanisms. While many TFs act together, the logic underlying TFs binding and their interactions is not fully understood yet. Most current approaches for TF binding site prediction focus on individual TFs and binary classification tasks, without a full analysis of the possible interactions among various TFs. In this paper we investigate DNA TF binding site recognition as a multi-label classification problem, achieving reliable predictions for multiple TFs on DNA sequences retrieved in public repositories. Our deep learning models are based on Temporal Convolutional Networks (TCNs), which are able to predict multiple TF binding profiles, capturing correlations among TFs andtheir cooperative regulatory mechanisms. Our results suggest that multi-label learning leading to reliable predictive performances can reveal biologically meaningful motifs and co-binding patterns consistent with known TF interactions, while also suggesting novel relationships and cooperation among TFs.
Bioinspired CNNs for border completion in occluded images
Mar 11, 2026Abstract:We exploit the mathematical modeling of the border completion problem in the visual cortex to design convolutional neural network (CNN) filters that enhance robustness to image occlusions. We evaluate our CNN architecture, BorderNet, on three occluded datasets (MNIST, Fashion-MNIST, and EMNIST) under two types of occlusions: stripes and grids. In all cases, BorderNet demonstrates improved performance, with gains varying depending on the severity of the occlusions and the dataset.
Sheaf Neural Networks and biomedical applications
Jan 29, 2026Abstract:The purpose of this paper is to elucidate the theory and mathematical modelling behind the sheaf neural network (SNN) algorithm and then show how SNN can effectively answer to biomedical questions in a concrete case study and outperform the most popular graph neural networks (GNNs) as graph convolutional networks (GCNs), graph attention networks (GAT) and GraphSage.
Enhancing CNNs robustness to occlusions with bioinspired filters for border completion
Apr 24, 2025



Abstract:We exploit the mathematical modeling of the visual cortex mechanism for border completion to define custom filters for CNNs. We see a consistent improvement in performance, particularly in accuracy, when our modified LeNet 5 is tested with occluded MNIST images.
Cartan moving frames and the data manifolds
Sep 18, 2024Abstract:The purpose of this paper is to employ the language of Cartan moving frames to study the geometry of the data manifolds and its Riemannian structure, via the data information metric and its curvature at data points. Using this framework and through experiments, explanations on the response of a neural network are given by pointing out the output classes that are easily reachable from a given input. This emphasizes how the proposed mathematical relationship between the output of the network and the geometry of its inputs can be exploited as an explainable artificial intelligence tool.
Model-centric Data Manifold: the Data Through the Eyes of the Model
Apr 26, 2021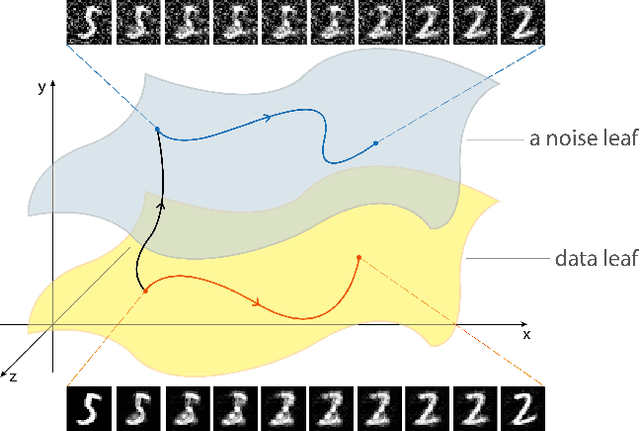
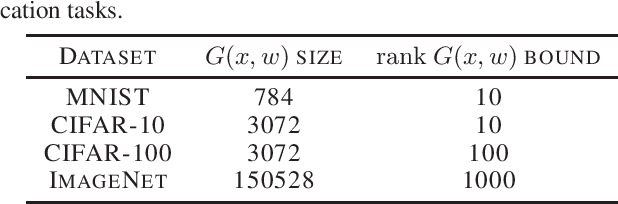
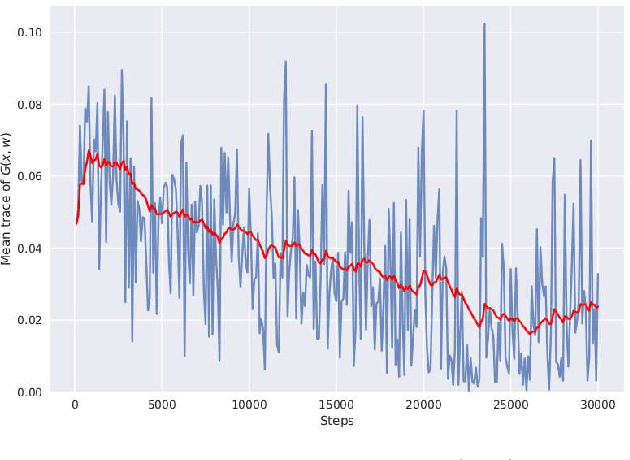

Abstract:We discover that deep ReLU neural network classifiers can see a low-dimensional Riemannian manifold structure on data. Such structure comes via the local data matrix, a variation of the Fisher information matrix, where the role of the model parameters is taken by the data variables. We obtain a foliation of the data domain and we show that the dataset on which the model is trained lies on a leaf, the data leaf, whose dimension is bounded by the number of classification labels. We validate our results with some experiments with the MNIST dataset: paths on the data leaf connect valid images, while other leaves cover noisy images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge