Przemysław Biecek
Warsaw University of Technology
Trojan horse hunt in deep forecasting models: Insights from the European Space Agency competition
Mar 20, 2026Abstract:Forecasting plays a crucial role in modern safety-critical applications, such as space operations. However, the increasing use of deep forecasting models introduces a new security risk of trojan horse attacks, carried out by hiding a backdoor in the training data or directly in the model weights. Once implanted, the backdoor is activated by a specific trigger pattern at test time, causing the model to produce manipulated predictions. We focus on this issue in our \textit{Trojan Horse Hunt} data science competition, where more than 200 teams faced the task of identifying triggers hidden in deep forecasting models for spacecraft telemetry. We describe the novel task formulation, benchmark set, evaluation protocol, and best solutions from the competition. We further summarize key insights and research directions for effective identification of triggers in time series forecasting models. All materials are publicly available on the official competition webpage https://www.kaggle.com/competitions/trojan-horse-hunt-in-space.
Analyzing the Temporal Factors for Anxiety and Depression Symptoms with the Rashomon Perspective
Jan 18, 2026Abstract:This paper introduces a new modeling perspective in the public mental health domain to provide a robust interpretation of the relations between anxiety and depression, and the demographic and temporal factors. This perspective particularly leverages the Rashomon Effect, where multiple models exhibit similar predictive performance but rely on diverse internal structures. Instead of considering these multiple models, choosing a single best model risks masking alternative narratives embedded in the data. To address this, we employed this perspective in the interpretation of a large-scale psychological dataset, specifically focusing on the Patient Health Questionnaire-4. We use a random forest model combined with partial dependence profiles to rigorously assess the robustness and stability of predictive relationships across the resulting Rashomon set, which consists of multiple models that exhibit similar predictive performance. Our findings confirm that demographic variables \texttt{age}, \texttt{sex}, and \texttt{education} lead to consistent structural shifts in anxiety and depression risk. Crucially, we identify significant temporal effects: risk probability demonstrates clear diurnal and circaseptan fluctuations, peaking during early morning hours. This work demonstrates the necessity of moving beyond the best model to analyze the entire Rashomon set. Our results highlight that the observed variability, particularly due to circadian and circaseptan rhythms, must be meticulously considered for robust interpretation in psychological screening. We advocate for a multiplicity-aware approach to enhance the stability and generalizability of ML-based conclusions in mental health research.
Fake or Real: The Impostor Hunt in Texts for Space Operations
Jul 17, 2025Abstract:The "Fake or Real" competition hosted on Kaggle (\href{https://www.kaggle.com/competitions/fake-or-real-the-impostor-hunt}{https://www.kaggle.com/competitions/fake-or-real-the-impostor-hunt}) is the second part of a series of follow-up competitions and hackathons related to the "Assurance for Space Domain AI Applications" project funded by the European Space Agency (\href{https://assurance-ai.space-codev.org/}{https://assurance-ai.space-codev.org/}). The competition idea is based on two real-life AI security threats identified within the project -- data poisoning and overreliance in Large Language Models. The task is to distinguish between the proper output from LLM and the output generated under malicious modification of the LLM. As this problem was not extensively researched, participants are required to develop new techniques to address this issue or adjust already existing ones to this problem's statement.
MASCOTS: Model-Agnostic Symbolic COunterfactual explanations for Time Series
Mar 28, 2025



Abstract:Counterfactual explanations provide an intuitive way to understand model decisions by identifying minimal changes required to alter an outcome. However, applying counterfactual methods to time series models remains challenging due to temporal dependencies, high dimensionality, and the lack of an intuitive human-interpretable representation. We introduce MASCOTS, a method that leverages the Bag-of-Receptive-Fields representation alongside symbolic transformations inspired by Symbolic Aggregate Approximation. By operating in a symbolic feature space, it enhances interpretability while preserving fidelity to the original data and model. Unlike existing approaches that either depend on model structure or autoencoder-based sampling, MASCOTS directly generates meaningful and diverse counterfactual observations in a model-agnostic manner, operating on both univariate and multivariate data. We evaluate MASCOTS on univariate and multivariate benchmark datasets, demonstrating comparable validity, proximity, and plausibility to state-of-the-art methods, while significantly improving interpretability and sparsity. Its symbolic nature allows for explanations that can be expressed visually, in natural language, or through semantic representations, making counterfactual reasoning more accessible and actionable.
The Dark Patterns of Personalized Persuasion in Large Language Models: Exposing Persuasive Linguistic Features for Big Five Personality Traits in LLMs Responses
Nov 12, 2024



Abstract:This study explores how the Large Language Models (LLMs) adjust linguistic features to create personalized persuasive outputs. While research showed that LLMs personalize outputs, a gap remains in understanding the linguistic features of their persuasive capabilities. We identified 13 linguistic features crucial for influencing personalities across different levels of the Big Five model of personality. We analyzed how prompts with personality trait information influenced the output of 19 LLMs across five model families. The findings show that models use more anxiety-related words for neuroticism, increase achievement-related words for conscientiousness, and employ fewer cognitive processes words for openness to experience. Some model families excel at adapting language for openness to experience, others for conscientiousness, while only one model adapts language for neuroticism. Our findings show how LLMs tailor responses based on personality cues in prompts, indicating their potential to create persuasive content affecting the mind and well-being of the recipients.
Does context matter in digital pathology?
May 23, 2024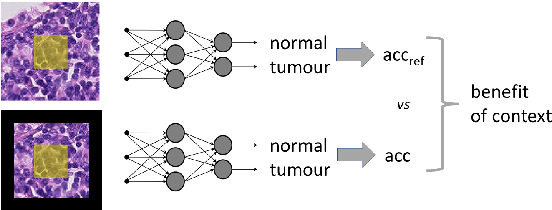
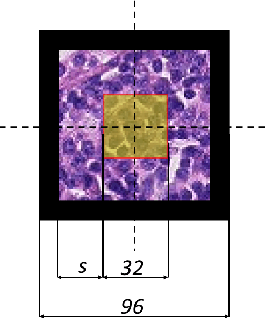
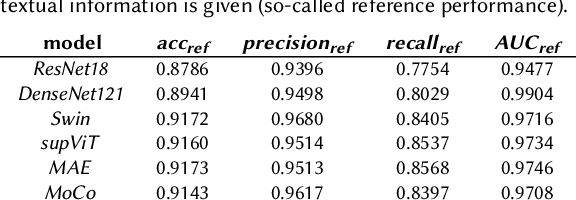
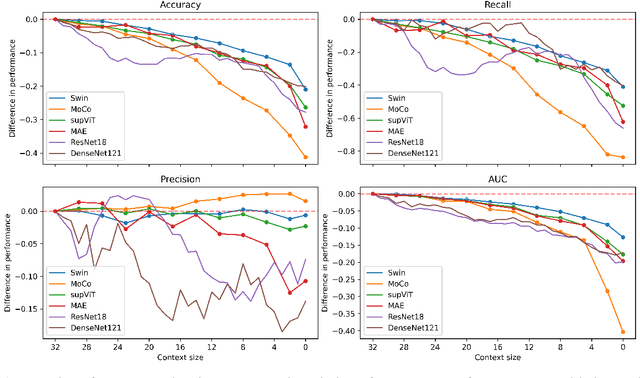
Abstract:The development of Artificial Intelligence for healthcare is of great importance. Models can sometimes achieve even superior performance to human experts, however, they can reason based on spurious features. This is not acceptable to the experts as it is expected that the models catch the valid patterns in the data following domain expertise. In the work, we analyse whether Deep Learning (DL) models for vision follow the histopathologists' practice so that when diagnosing a part of a lesion, they take into account also the surrounding tissues which serve as context. It turns out that the performance of DL models significantly decreases when the amount of contextual information is limited, therefore contextual information is valuable at prediction time. Moreover, we show that the models sometimes behave in an unstable way as for some images, they change the predictions many times depending on the size of the context. It may suggest that partial contextual information can be misleading.
Position paper: Do not explain without context
Apr 28, 2024Abstract:Does the stethoscope in the picture make the adjacent person a doctor or a patient? This, of course, depends on the contextual relationship of the two objects. If it is obvious, why don not explanation methods for vision models use contextual information? In this paper, we (1) review the most popular methods of explaining computer vision models by pointing out that they do not take into account context information, (2) provide examples of real-world use cases where spatial context plays a significant role, (3) propose new research directions that may lead to better use of context information in explaining computer vision models, (4) argue that a change in approach to explanations is needed from 'where' to 'how'.
Global Counterfactual Directions
Apr 18, 2024Abstract:Despite increasing progress in development of methods for generating visual counterfactual explanations, especially with the recent rise of Denoising Diffusion Probabilistic Models, previous works consider them as an entirely local technique. In this work, we take the first step at globalizing them. Specifically, we discover that the latent space of Diffusion Autoencoders encodes the inference process of a given classifier in the form of global directions. We propose a novel proxy-based approach that discovers two types of these directions with the use of only single image in an entirely black-box manner. Precisely, g-directions allow for flipping the decision of a given classifier on an entire dataset of images, while h-directions further increase the diversity of explanations. We refer to them in general as Global Counterfactual Directions (GCDs). Moreover, we show that GCDs can be naturally combined with Latent Integrated Gradients resulting in a new black-box attribution method, while simultaneously enhancing the understanding of counterfactual explanations. We validate our approach on existing benchmarks and show that it generalizes to real-world use-cases.
CNN-based explanation ensembling for dataset, representation and explanations evaluation
Apr 16, 2024



Abstract:Explainable Artificial Intelligence has gained significant attention due to the widespread use of complex deep learning models in high-stake domains such as medicine, finance, and autonomous cars. However, different explanations often present different aspects of the model's behavior. In this research manuscript, we explore the potential of ensembling explanations generated by deep classification models using convolutional model. Through experimentation and analysis, we aim to investigate the implications of combining explanations to uncover a more coherent and reliable patterns of the model's behavior, leading to the possibility of evaluating the representation learned by the model. With our method, we can uncover problems of under-representation of images in a certain class. Moreover, we discuss other side benefits like features' reduction by replacing the original image with its explanations resulting in the removal of some sensitive information. Through the use of carefully selected evaluation metrics from the Quantus library, we demonstrated the method's superior performance in terms of Localisation and Faithfulness, compared to individual explanations.
A comparative analysis of deep learning models for lung segmentation on X-ray images
Apr 09, 2024


Abstract:Robust and highly accurate lung segmentation in X-rays is crucial in medical imaging. This study evaluates deep learning solutions for this task, ranking existing methods and analyzing their performance under diverse image modifications. Out of 61 analyzed papers, only nine offered implementation or pre-trained models, enabling assessment of three prominent methods: Lung VAE, TransResUNet, and CE-Net. The analysis revealed that CE-Net performs best, demonstrating the highest values in dice similarity coefficient and intersection over union metric.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge