P. Liatsis
Department of Electrical Engineering and Computer Science, Khalifa University
k-MS: A novel clustering algorithm based on morphological reconstruction
Aug 30, 2022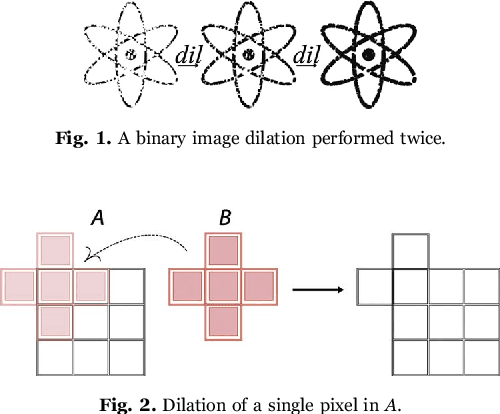
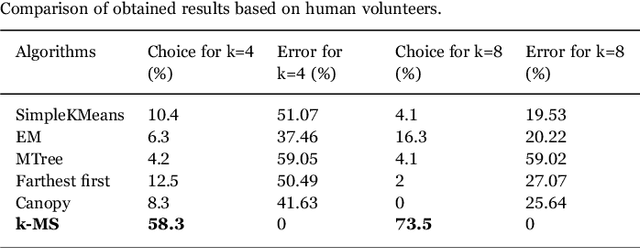
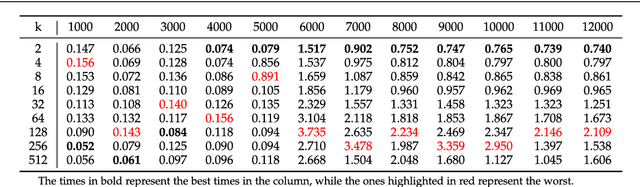

Abstract:This work proposes a clusterization algorithm called k-Morphological Sets (k-MS), based on morphological reconstruction and heuristics. k-MS is faster than the CPU-parallel k-Means in worst case scenarios and produces enhanced visualizations of the dataset as well as very distinct clusterizations. It is also faster than similar clusterization methods that are sensitive to density and shapes such as Mitosis and TRICLUST. In addition, k-MS is deterministic and has an intrinsic sense of maximal clusters that can be created for a given input sample and input parameters, differing from k-Means and other clusterization algorithms. In other words, given a constant k, a structuring element and a dataset, k-MS produces k or less clusters without using random/ pseudo-random functions. Finally, the proposed algorithm also provides a straightforward means for removing noise from images or datasets in general.
Automated recognition of the pericardium contour on processed CT images using genetic algorithms
Aug 30, 2022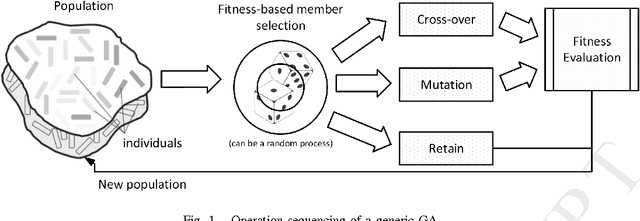

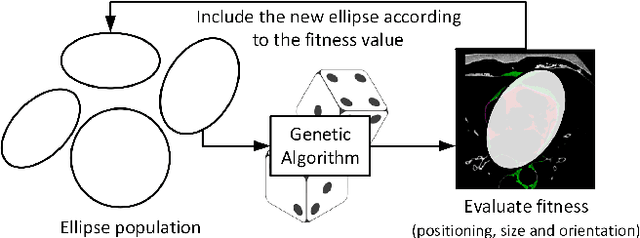

Abstract:This work proposes the use of Genetic Algorithms (GA) in tracing and recognizing the pericardium contour of the human heart using Computed Tomography (CT) images. We assume that each slice of the pericardium can be modelled by an ellipse, the parameters of which need to be optimally determined. An optimal ellipse would be one that closely follows the pericardium contour and, consequently, separates appropriately the epicardial and mediastinal fats of the human heart. Tracing and automatically identifying the pericardium contour aids in medical diagnosis. Usually, this process is done manually or not done at all due to the effort required. Besides, detecting the pericardium may improve previously proposed automated methodologies that separate the two types of fat associated to the human heart. Quantification of these fats provides important health risk marker information, as they are associated with the development of certain cardiovascular pathologies. Finally, we conclude that GA offers satisfiable solutions in a feasible amount of processing time.
Machine learning in the prediction of cardiac epicardial and mediastinal fat volumes
Aug 30, 2022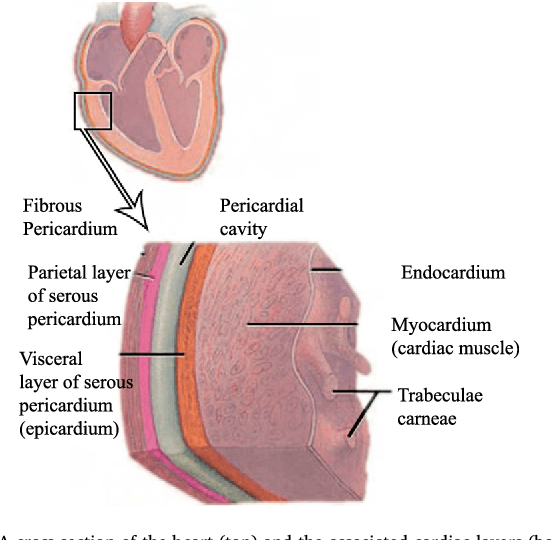
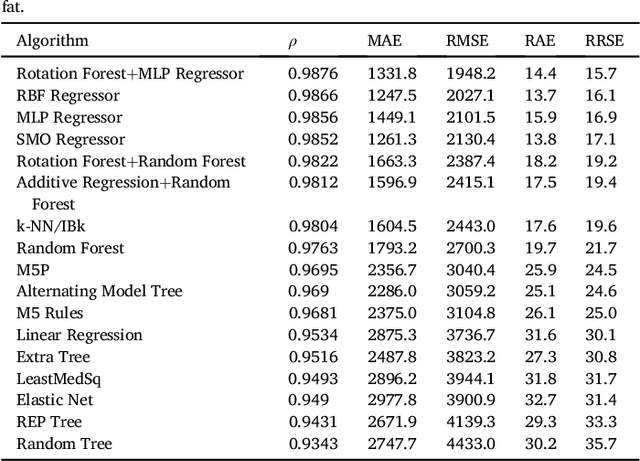
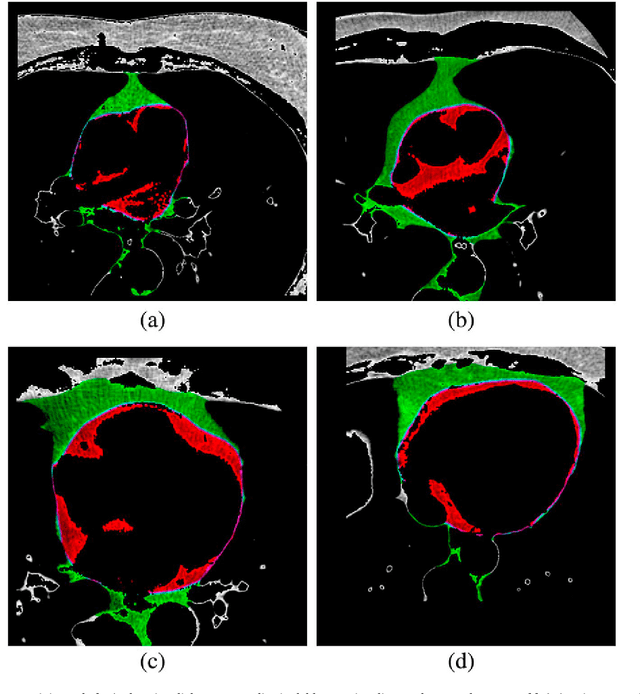
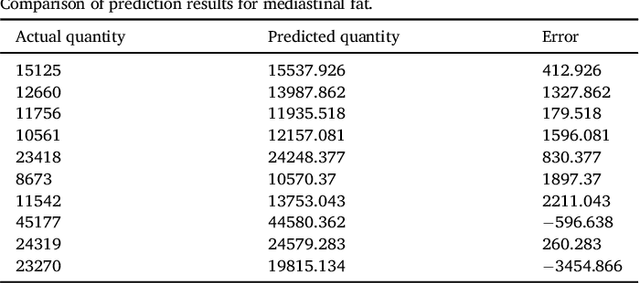
Abstract:We propose a methodology to predict the cardiac epicardial and mediastinal fat volumes in computed tomography images using regression algorithms. The obtained results indicate that it is feasible to predict these fats with a high degree of correlation, thus alleviating the requirement for manual or automatic segmentation of both fat volumes. Instead, segmenting just one of them suffices, while the volume of the other may be predicted fairly precisely. The correlation coefficient obtained by the Rotation Forest algorithm using MLP Regressor for predicting the mediastinal fat based on the epicardial fat was 0.9876, with a relative absolute error of 14.4% and a root relative squared error of 15.7%. The best correlation coefficient obtained in the prediction of the epicardial fat based on the mediastinal was 0.9683 with a relative absolute error of 19.6% and a relative squared error of 24.9%. Moreover, we analysed the feasibility of using linear regressors, which provide an intuitive interpretation of the underlying approximations. In this case, the obtained correlation coefficient was 0.9534 for predicting the mediastinal fat based on the epicardial, with a relative absolute error of 31.6% and a root relative squared error of 30.1%. On the prediction of the epicardial fat based on the mediastinal fat, the correlation coefficient was 0.8531, with a relative absolute error of 50.43% and a root relative squared error of 52.06%. In summary, it is possible to speed up general medical analyses and some segmentation and quantification methods that are currently employed in the state-of-the-art by using this prediction approach, which consequently reduces costs and therefore enables preventive treatments that may lead to a reduction of health problems.
Multi-Label Retinal Disease Classification using Transformers
Jul 07, 2022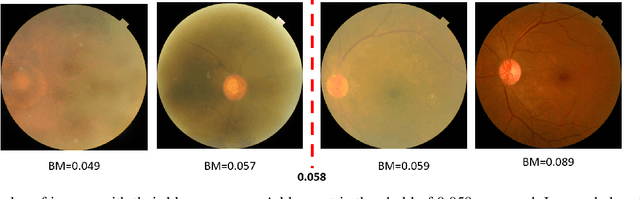
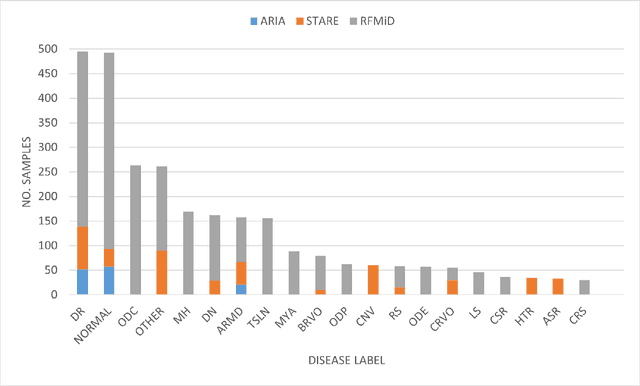
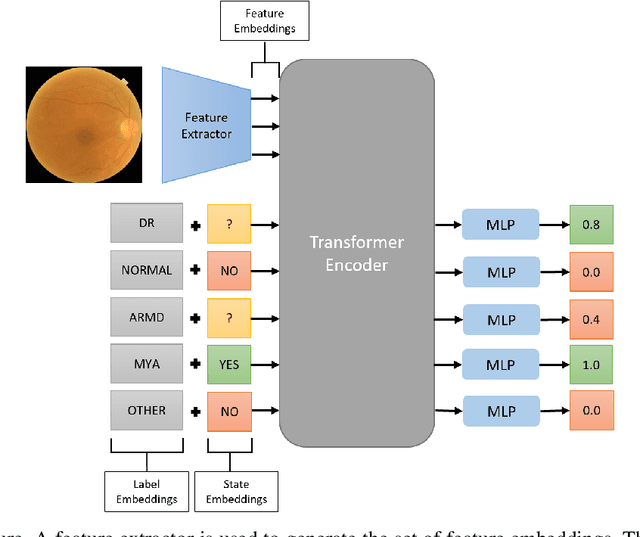
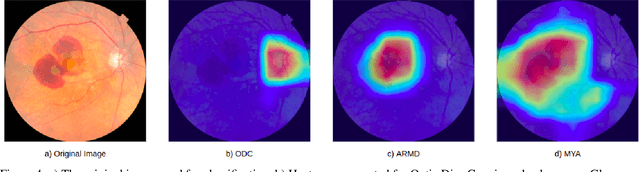
Abstract:Early detection of retinal diseases is one of the most important means of preventing partial or permanent blindness in patients. In this research, a novel multi-label classification system is proposed for the detection of multiple retinal diseases, using fundus images collected from a variety of sources. First, a new multi-label retinal disease dataset, the MuReD dataset, is constructed, using a number of publicly available datasets for fundus disease classification. Next, a sequence of post-processing steps is applied to ensure the quality of the image data and the range of diseases, present in the dataset. For the first time in fundus multi-label disease classification, a transformer-based model optimized through extensive experimentation is used for image analysis and decision making. Numerous experiments are performed to optimize the configuration of the proposed system. It is shown that the approach performs better than state-of-the-art works on the same task by 7.9% and 8.1% in terms of AUC score for disease detection and disease classification, respectively. The obtained results further support the potential applications of transformer-based architectures in the medical imaging field.
Morphological classifiers
Dec 21, 2021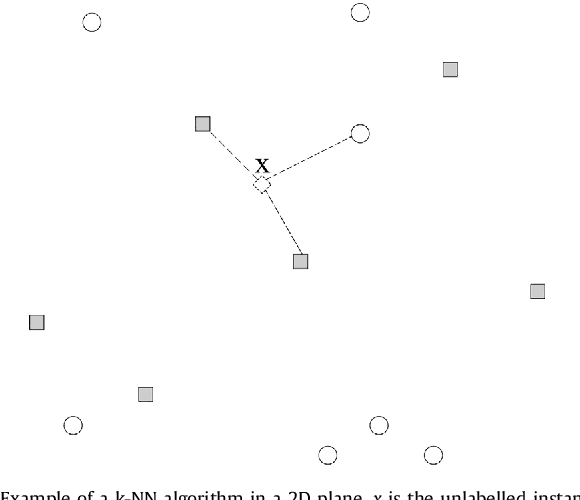

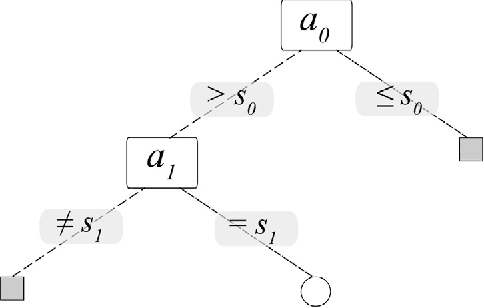
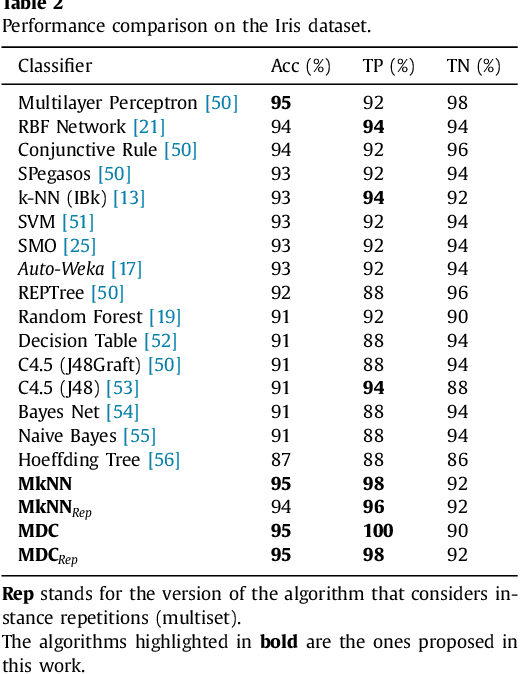
Abstract:This work proposes a new type of classifier called Morphological Classifier (MC). MCs aggregate concepts from mathematical morphology and supervised learning. The outcomes of this aggregation are classifiers that may preserve shape characteristics of classes, subject to the choice of a stopping criterion and structuring element. MCs are fundamentally based on set theory, and their classification model can be a mathematical set itself. Two types of morphological classifiers are proposed in the current work, namely, Morphological k-NN (MkNN) and Morphological Dilation Classifier (MDC), which demonstrate the feasibility of the approach. This work provides evidence regarding the advantages of MCs, e.g., very fast classification times as well as competitive accuracy rates. The performance of MkNN and MDC was tested using p -dimensional datasets. MCs tied or outperformed 14 well established classifiers in 5 out of 8 datasets. In all occasions, the obtained accuracies were higher than the average accuracy obtained with all classifiers. Moreover, the proposed implementations utilize the power of the Graphics Processing Units (GPUs) to speed up processing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge