Oskar Taubert
AutoPQ: Automating Quantile estimation from Point forecasts in the context of sustainability
Nov 30, 2024



Abstract:Optimizing smart grid operations relies on critical decision-making informed by uncertainty quantification, making probabilistic forecasting a vital tool. Designing such forecasting models involves three key challenges: accurate and unbiased uncertainty quantification, workload reduction for data scientists during the design process, and limitation of the environmental impact of model training. In order to address these challenges, we introduce AutoPQ, a novel method designed to automate and optimize probabilistic forecasting for smart grid applications. AutoPQ enhances forecast uncertainty quantification by generating quantile forecasts from an existing point forecast by using a conditional Invertible Neural Network (cINN). AutoPQ also automates the selection of the underlying point forecasting method and the optimization of hyperparameters, ensuring that the best model and configuration is chosen for each application. For flexible adaptation to various performance needs and available computing power, AutoPQ comes with a default and an advanced configuration, making it suitable for a wide range of smart grid applications. Additionally, AutoPQ provides transparency regarding the electricity consumption required for performance improvements. We show that AutoPQ outperforms state-of-the-art probabilistic forecasting methods while effectively limiting computational effort and hence environmental impact. Additionally and in the context of sustainability, we quantify the electricity consumption required for performance improvements.
ReCycle: Fast and Efficient Long Time Series Forecasting with Residual Cyclic Transformers
May 06, 2024Abstract:Transformers have recently gained prominence in long time series forecasting by elevating accuracies in a variety of use cases. Regrettably, in the race for better predictive performance the overhead of model architectures has grown onerous, leading to models with computational demand infeasible for most practical applications. To bridge the gap between high method complexity and realistic computational resources, we introduce the Residual Cyclic Transformer, ReCycle. ReCycle utilizes primary cycle compression to address the computational complexity of the attention mechanism in long time series. By learning residuals from refined smoothing average techniques, ReCycle surpasses state-of-the-art accuracy in a variety of application use cases. The reliable and explainable fallback behavior ensured by simple, yet robust, smoothing average techniques additionally lowers the barrier for user acceptance. At the same time, our approach reduces the run time and energy consumption by more than an order of magnitude, making both training and inference feasible on low-performance, low-power and edge computing devices. Code is available at https://github.com/Helmholtz-AI-Energy/ReCycle
Massively Parallel Genetic Optimization through Asynchronous Propagation of Populations
Jan 20, 2023Abstract:We present Propulate, an evolutionary optimization algorithm and software package for global optimization and in particular hyperparameter search. For efficient use of HPC resources, Propulate omits the synchronization after each generation as done in conventional genetic algorithms. Instead, it steers the search with the complete population present at time of breeding new individuals. We provide an MPI-based implementation of our algorithm, which features variants of selection, mutation, crossover, and migration and is easy to extend with custom functionality. We compare Propulate to the established optimization tool Optuna. We find that Propulate is up to three orders of magnitude faster without sacrificing solution accuracy, demonstrating the efficiency and efficacy of our lazy synchronization approach. Code and documentation are available at https://github.com/Helmholtz-AI-Energy/propulate
Learning Tree Structures from Leaves For Particle Decay Reconstruction
Sep 01, 2022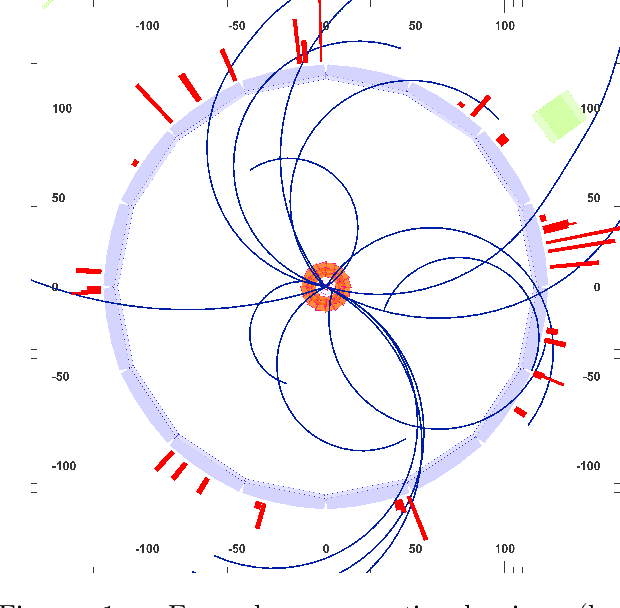
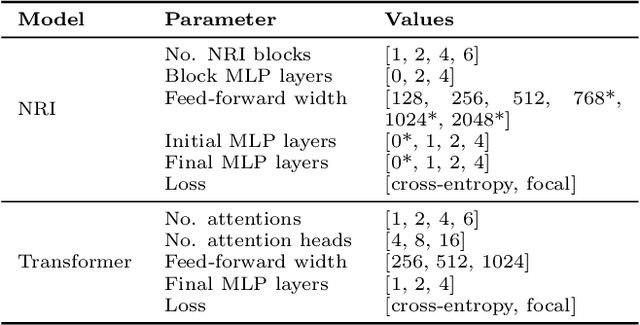
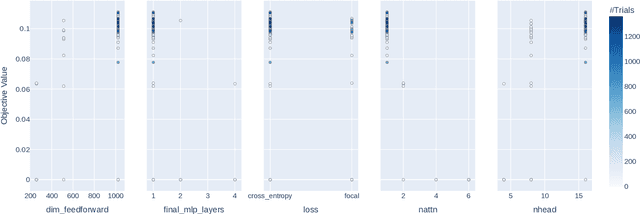
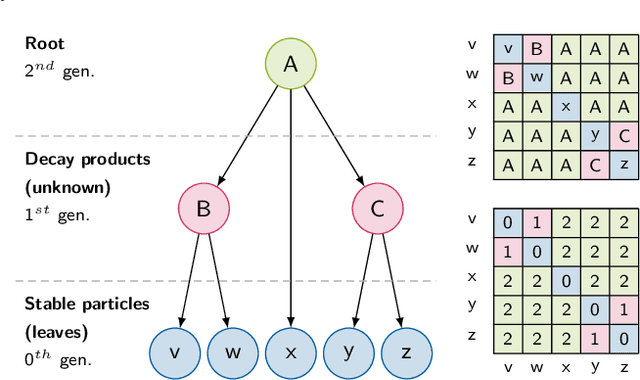
Abstract:In this work, we present a neural approach to reconstructing rooted tree graphs describing hierarchical interactions, using a novel representation we term the Lowest Common Ancestor Generations (LCAG) matrix. This compact formulation is equivalent to the adjacency matrix, but enables learning a tree's structure from its leaves alone without the prior assumptions required if using the adjacency matrix directly. Employing the LCAG therefore enables the first end-to-end trainable solution which learns the hierarchical structure of varying tree sizes directly, using only the terminal tree leaves to do so. In the case of high-energy particle physics, a particle decay forms a hierarchical tree structure of which only the final products can be observed experimentally, and the large combinatorial space of possible trees makes an analytic solution intractable. We demonstrate the use of the LCAG as a target in the task of predicting simulated particle physics decay structures using both a Transformer encoder and a Neural Relational Inference encoder Graph Neural Network. With this approach, we are able to correctly predict the LCAG purely from leaf features for a maximum tree-depth of $8$ in $92.5\%$ of cases for trees up to $6$ leaves (including) and $59.7\%$ for trees up to $10$ in our simulated dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge