Okito Yamashita
Correntropy-Based Improper Likelihood Model for Robust Electrophysiological Source Imaging
Aug 27, 2024



Abstract:Bayesian learning provides a unified skeleton to solve the electrophysiological source imaging task. From this perspective, existing source imaging algorithms utilize the Gaussian assumption for the observation noise to build the likelihood function for Bayesian inference. However, the electromagnetic measurements of brain activity are usually affected by miscellaneous artifacts, leading to a potentially non-Gaussian distribution for the observation noise. Hence the conventional Gaussian likelihood model is a suboptimal choice for the real-world source imaging task. In this study, we aim to solve this problem by proposing a new likelihood model which is robust with respect to non-Gaussian noises. Motivated by the robust maximum correntropy criterion, we propose a new improper distribution model concerning the noise assumption. This new noise distribution is leveraged to structure a robust likelihood function and integrated with hierarchical prior distributions to estimate source activities by variational inference. In particular, the score matching is adopted to determine the hyperparameters for the improper likelihood model. A comprehensive performance evaluation is performed to compare the proposed noise assumption to the conventional Gaussian model. Simulation results show that, the proposed method can realize more precise source reconstruction by designing known ground-truth. The real-world dataset also demonstrates the superiority of our new method with the visual perception task. This study provides a new backbone for Bayesian source imaging, which would facilitate its application using real-world noisy brain signal.
Fast, accurate, and interpretable decoding of electrocorticographic signals using dynamic mode decomposition
Oct 31, 2023Abstract:Dynamic mode (DM) decomposition decomposes spatiotemporal signals into basic oscillatory components (DMs). DMs can improve the accuracy of neural decoding when used with the nonlinear Grassmann kernel, compared to conventional power features. However, such kernel-based machine learning algorithms have three limitations: large computational time preventing real-time application, incompatibility with non-kernel algorithms, and low interpretability. Here, we propose a mapping function corresponding to the Grassmann kernel that explicitly transforms DMs into spatial DM (sDM) features, which can be used in any machine learning algorithm. Using electrocorticographic signals recorded during various movement and visual perception tasks, the sDM features were shown to improve the decoding accuracy and computational time compared to conventional methods. Furthermore, the components of the sDM features informative for decoding showed similar characteristics to the high-$\gamma$ power of the signals, but with higher trial-to-trial reproducibility. The proposed sDM features enable fast, accurate, and interpretable neural decoding.
Adaptive sparseness for correntropy-based robust regression via automatic relevance determination
Jan 31, 2023



Abstract:Sparseness and robustness are two important properties for many machine learning scenarios. In the present study, regarding the maximum correntropy criterion (MCC) based robust regression algorithm, we investigate to integrate the MCC method with the automatic relevance determination (ARD) technique in a Bayesian framework, so that MCC-based robust regression could be implemented with adaptive sparseness. To be specific, we use an inherent noise assumption from the MCC to derive an explicit likelihood function, and realize the maximum a posteriori (MAP) estimation with the ARD prior by variational Bayesian inference. Compared to the existing robust and sparse L1-regularized MCC regression, the proposed MCC-ARD regression can eradicate the troublesome tuning for the regularization hyper-parameter which controls the regularization strength. Further, MCC-ARD achieves superior prediction performance and feature selection capability than L1-regularized MCC, as demonstrated by a noisy and high-dimensional simulation study.
Multiple-view clustering for correlation matrices based on Wishart mixture model
Oct 20, 2020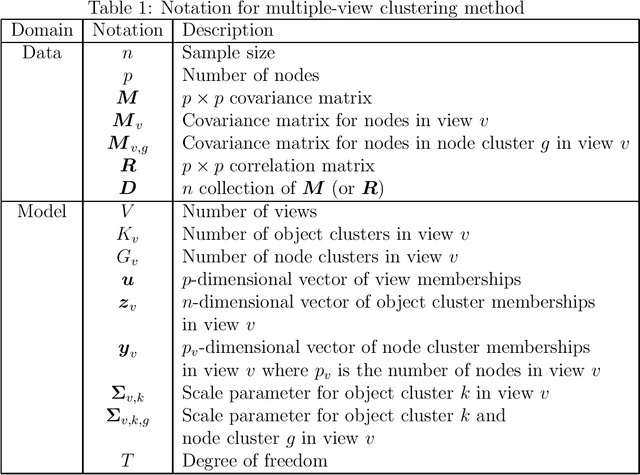
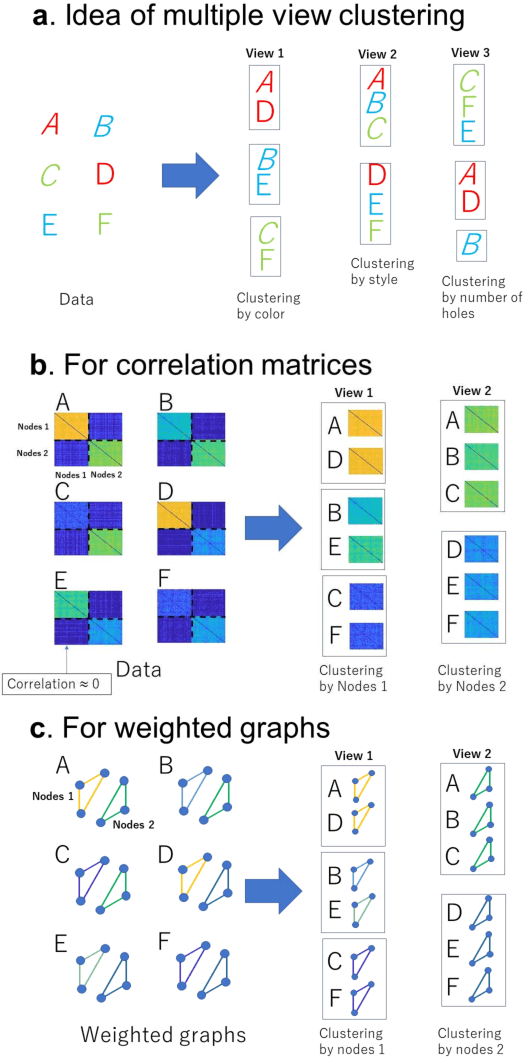
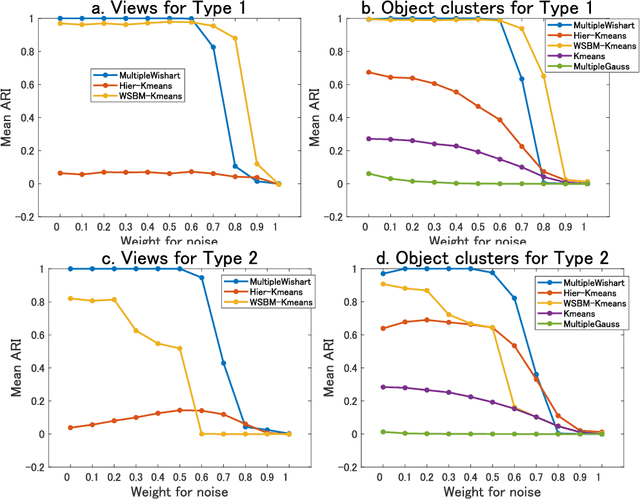
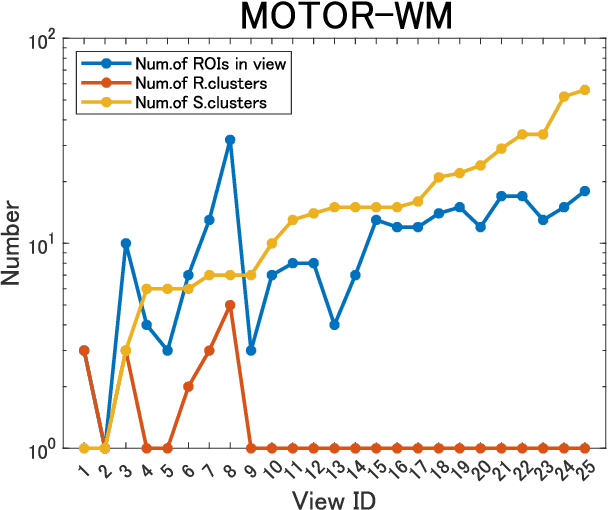
Abstract:A multiple-view clustering method is a powerful analytical tool for high-dimensional data, such as functional magnetic resonance imaging (fMRI). It can identify clustering patterns of subjects depending on their functional connectivity in specific brain areas. However, when one applies an existing method to fMRI data, there is a need to simplify the data structure, independently dealing with elements in a functional connectivity matrix, that is, a correlation matrix. In general, elements in a correlation matrix are closely associated. Hence, such a simplification may distort the clustering results. To overcome this problem, we propose a novel multiple-view clustering method based on the Wishart mixture model, which preserves the correlation matrix structure. The uniqueness of this method is that the multiple-view clustering of subjects is based on particular networks of nodes (or regions of interest (ROIs) in fMRI), optimized in a data-driven manner. Hence, it can identify multiple underlying pairs of associations between a subject cluster solution and a ROI network. The key assumption of the method is independence among networks, which is effectively addressed by whitening correlation matrices. We applied the proposed method to synthetic and fMRI data, demonstrating the usefulness and power of the proposed method.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge