Neil R. Smalheiser
Robust Biomedical Publication Type and Study Design Classification with Knowledge-Guided Perturbations
May 12, 2026Abstract:Accurately and consistently indexing biomedical literature by publication type and study design is essential for supporting evidence synthesis and knowledge discovery. Prior work on automated publication type and study design indexing has primarily focused on expanding label coverage, enriching feature representations, and improving in-domain accuracy, with evaluation typically conducted on data drawn from the same distribution as training. Although pretrained biomedical language models achieve strong performance under these settings, models optimized for in-domain accuracy may rely on superficial lexical or dataset-specific cues, resulting in reduced robustness under distributional shift. In this study, we introduce an evaluation framework based on controlled semantic perturbations to assess the robustness of a publication type classifier and investigate robustness-oriented training strategies that combine entity masking and domain-adversarial training to mitigate reliance on spurious topical correlations. Our results show that the commonly observed trade-off between robustness and in-domain accuracy can be mitigated when robustness objectives are designed to selectively suppress non-task-defining features while preserving salient methodological signals. We find that these improvements arise from two complementary mechanisms: (1) increased reliance on explicit methodological cues when such cues are present in the input, and (2) reduced reliance on spurious domain-specific topical features. These findings highlight the importance of feature-level robustness analysis for publication type and study design classification and suggest that refining masking and adversarial objectives to more selectively suppress topical information may further improve robustness. Data, code, and models are available at: https://github.com/ScienceNLP-Lab/MultiTagger-v2/tree/main/ICHI
Unsupervised Low-Dimensional Vector Representations for Words, Phrases and Text that are Transparent, Scalable, and produce Similarity Metrics that are Complementary to Neural Embeddings
Jan 09, 2018


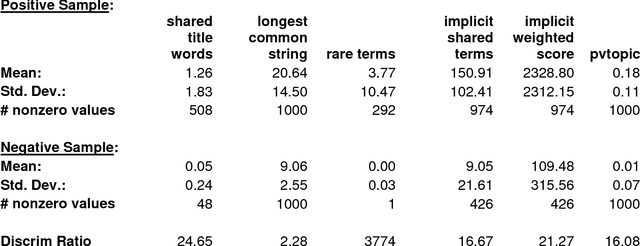
Abstract:Neural embeddings are a popular set of methods for representing words, phrases or text as a low dimensional vector (typically 50-500 dimensions). However, it is difficult to interpret these dimensions in a meaningful manner, and creating neural embeddings requires extensive training and tuning of multiple parameters and hyperparameters. We present here a simple unsupervised method for representing words, phrases or text as a low dimensional vector, in which the meaning and relative importance of dimensions is transparent to inspection. We have created a near-comprehensive vector representation of words, and selected bigrams, trigrams and abbreviations, using the set of titles and abstracts in PubMed as a corpus. This vector is used to create several novel implicit word-word and text-text similarity metrics. The implicit word-word similarity metrics correlate well with human judgement of word pair similarity and relatedness, and outperform or equal all other reported methods on a variety of biomedical benchmarks, including several implementations of neural embeddings trained on PubMed corpora. Our implicit word-word metrics capture different aspects of word-word relatedness than word2vec-based metrics and are only partially correlated (rho = ~0.5-0.8 depending on task and corpus). The vector representations of words, bigrams, trigrams, abbreviations, and PubMed title+abstracts are all publicly available from http://arrowsmith.psych.uic.edu for release under CC-BY-NC license. Several public web query interfaces are also available at the same site, including one which allows the user to specify a given word and view its most closely related terms according to direct co-occurrence as well as different implicit similarity metrics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge