Mohammad Golbabaee
MRI2Qmap: multi-parametric quantitative mapping with MRI-driven denoising priors
Mar 11, 2026Abstract:Magnetic Resonance Fingerprinting (MRF) and other highly accelerated transient-state parameter mapping techniques enable simultaneous quantification of multiple tissue properties, but often suffer from aliasing artifacts due to compressed sampling. Incorporating spatial image priors can mitigate these artifacts, and deep learning has shown strong potential when large training datasets are available. However, extending this paradigm to MRF-type sequences remains challenging due to the scarcity of quantitative imaging data for training. Can this limitation be overcome by leveraging sources of training data from clinically-routine weighted MRI images? To this end, we introduce MRI2Qmap, a plug-and-play quantitative reconstruction framework that integrates the physical acquisition model with priors learned from deep denoising autoencoders pretrained on large multimodal weighted-MRI datasets. MRI2Qmap demonstrates that spatial-domain structural priors learned from independently acquired datasets of routine weighted-MRI images can be effectively used for quantitative MRI reconstruction. The proposed method is validated on highly accelerated 3D whole-brain MRF data from both in-vivo and simulated acquisitions, achieving competitive or superior performance relative to existing baselines without requiring ground-truth quantitative imaging data for training. By decoupling quantitative reconstruction from the need for ground-truth MRF training data, this framework points toward a scalable paradigm for quantitative MRI that can capitalize on the large and growing repositories of routine clinical MRI.
Grouped Sequential Optimization Strategy -- the Application of Hyperparameter Importance Assessment in Deep Learning
Mar 07, 2025Abstract:Hyperparameter optimization (HPO) is a critical component of machine learning pipelines, significantly affecting model robustness, stability, and generalization. However, HPO is often a time-consuming and computationally intensive task. Traditional HPO methods, such as grid search and random search, often suffer from inefficiency. Bayesian optimization, while more efficient, still struggles with high-dimensional search spaces. In this paper, we contribute to the field by exploring how insights gained from hyperparameter importance assessment (HIA) can be leveraged to accelerate HPO, reducing both time and computational resources. Building on prior work that quantified hyperparameter importance by evaluating 10 hyperparameters on CNNs using 10 common image classification datasets, we implement a novel HPO strategy called 'Sequential Grouping.' That prior work assessed the importance weights of the investigated hyperparameters based on their influence on model performance, providing valuable insights that we leverage to optimize our HPO process. Our experiments, validated across six additional image classification datasets, demonstrate that incorporating hyperparameter importance assessment (HIA) can significantly accelerate HPO without compromising model performance, reducing optimization time by an average of 31.9\% compared to the conventional simultaneous strategy.
Improved Patch Denoising Diffusion Probabilistic Models for Magnetic Resonance Fingerprinting
Oct 29, 2024



Abstract:Magnetic Resonance Fingerprinting (MRF) is a time-efficient approach to quantitative MRI, enabling the mapping of multiple tissue properties from a single, accelerated scan. However, achieving accurate reconstructions remains challenging, particularly in highly accelerated and undersampled acquisitions, which are crucial for reducing scan times. While deep learning techniques have advanced image reconstruction, the recent introduction of diffusion models offers new possibilities for imaging tasks, though their application in the medical field is still emerging. Notably, diffusion models have not yet been explored for the MRF problem. In this work, we propose for the first time a conditional diffusion probabilistic model for MRF image reconstruction. Qualitative and quantitative comparisons on in-vivo brain scan data demonstrate that the proposed approach can outperform established deep learning and compressed sensing algorithms for MRF reconstruction. Extensive ablation studies also explore strategies to improve computational efficiency of our approach.
Efficient Hyperparameter Importance Assessment for CNNs
Oct 11, 2024



Abstract:Hyperparameter selection is an essential aspect of the machine learning pipeline, profoundly impacting models' robustness, stability, and generalization capabilities. Given the complex hyperparameter spaces associated with Neural Networks and the constraints of computational resources and time, optimizing all hyperparameters becomes impractical. In this context, leveraging hyperparameter importance assessment (HIA) can provide valuable guidance by narrowing down the search space. This enables machine learning practitioners to focus their optimization efforts on the hyperparameters with the most significant impact on model performance while conserving time and resources. This paper aims to quantify the importance weights of some hyperparameters in Convolutional Neural Networks (CNNs) with an algorithm called N-RReliefF, laying the groundwork for applying HIA methodologies in the Deep Learning field. We conduct an extensive study by training over ten thousand CNN models across ten popular image classification datasets, thereby acquiring a comprehensive dataset containing hyperparameter configuration instances and their corresponding performance metrics. It is demonstrated that among the investigated hyperparameters, the top five important hyperparameters of the CNN model are the number of convolutional layers, learning rate, dropout rate, optimizer and epoch.
StoDIP: Efficient 3D MRF image reconstruction with deep image priors and stochastic iterations
Aug 05, 2024


Abstract:Magnetic Resonance Fingerprinting (MRF) is a time-efficient approach to quantitative MRI for multiparametric tissue mapping. The reconstruction of quantitative maps requires tailored algorithms for removing aliasing artefacts from the compressed sampled MRF acquisitions. Within approaches found in the literature, many focus solely on two-dimensional (2D) image reconstruction, neglecting the extension to volumetric (3D) scans despite their higher relevance and clinical value. A reason for this is that transitioning to 3D imaging without appropriate mitigations presents significant challenges, including increased computational cost and storage requirements, and the need for large amount of ground-truth (artefact-free) data for training. To address these issues, we introduce StoDIP, a new algorithm that extends the ground-truth-free Deep Image Prior (DIP) reconstruction to 3D MRF imaging. StoDIP employs memory-efficient stochastic updates across the multicoil MRF data, a carefully selected neural network architecture, as well as faster nonuniform FFT (NUFFT) transformations. This enables a faster convergence compared against a conventional DIP implementation without these features. Tested on a dataset of whole-brain scans from healthy volunteers, StoDIP demonstrated superior performance over the ground-truth-free reconstruction baselines, both quantitatively and qualitatively.
Deep Image Priors for Magnetic Resonance Fingerprinting with pretrained Bloch-consistent denoising autoencoders
Jul 29, 2024



Abstract:The estimation of multi-parametric quantitative maps from Magnetic Resonance Fingerprinting (MRF) compressed sampled acquisitions, albeit successful, remains a challenge due to the high underspampling rate and artifacts naturally occuring during image reconstruction. Whilst state-of-the-art DL methods can successfully address the task, to fully exploit their capabilities they often require training on a paired dataset, in an area where ground truth is seldom available. In this work, we propose a method that combines a deep image prior (DIP) module that, without ground truth and in conjunction with a Bloch consistency enforcing autoencoder, can tackle the problem, resulting in a method faster and of equivalent or better accuracy than DIP-MRF.
Performance of data-driven inner speech decoding with same-task EEG-fMRI data fusion and bimodal models
Jun 19, 2023



Abstract:Decoding inner speech from the brain signal via hybridisation of fMRI and EEG data is explored to investigate the performance benefits over unimodal models. Two different bimodal fusion approaches are examined: concatenation of probability vectors output from unimodal fMRI and EEG machine learning models, and data fusion with feature engineering. Same task inner speech data are recorded from four participants, and different processing strategies are compared and contrasted to previously-employed hybridisation methods. Data across participants are discovered to encode different underlying structures, which results in varying decoding performances between subject-dependent fusion models. Decoding performance is demonstrated as improved when pursuing bimodal fMRI-EEG fusion strategies, if the data show underlying structure.
Nonlinear Equivariant Imaging: Learning Multi-Parametric Tissue Mapping without Ground Truth for Compressive Quantitative MRI
Nov 23, 2022Abstract:Current state-of-the-art reconstruction for quantitative tissue maps from fast, compressive, Magnetic Resonance Fingerprinting (MRF), use supervised deep learning, with the drawback of requiring high-fidelity ground truth tissue map training data which is limited. This paper proposes NonLinear Equivariant Imaging (NLEI), a self-supervised learning approach to eliminate the need for ground truth for deep MRF image reconstruction. NLEI extends the recent Equivariant Imaging framework to nonlinear inverse problems such as MRF. Only fast, compressed-sampled MRF scans are used for training. NLEI learns tissue mapping using spatiotemporal priors: spatial priors are obtained from the invariance of MRF data to a group of geometric image transformations, while temporal priors are obtained from a nonlinear Bloch response model approximated by a pre-trained neural network. Tested retrospectively on two acquisition settings, we observe that NLEI (self-supervised learning) closely approaches the performance of supervised learning, despite not using ground truth during training.
A Plug-and-Play Approach to Multiparametric Quantitative MRI: Image Reconstruction using Pre-Trained Deep Denoisers
Feb 10, 2022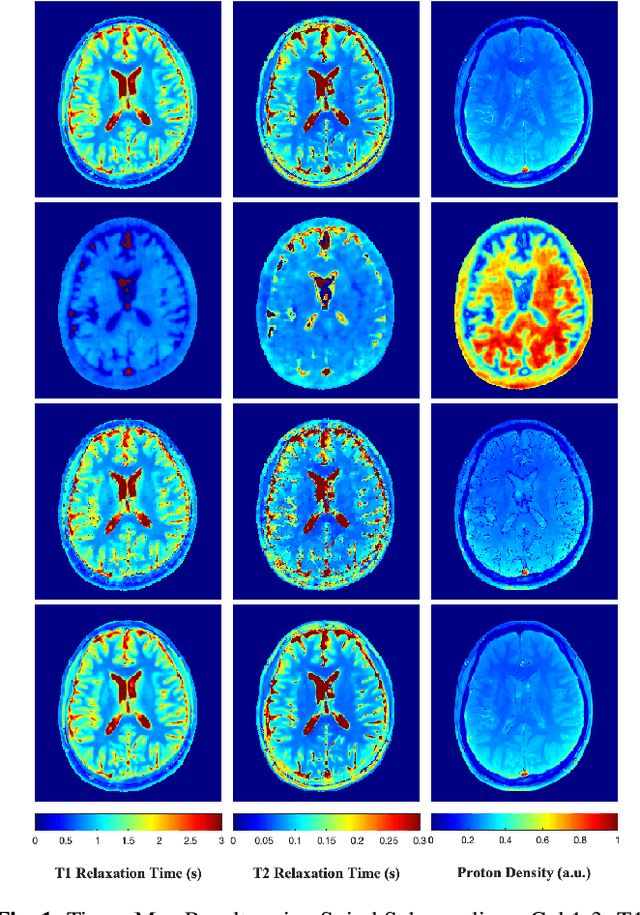
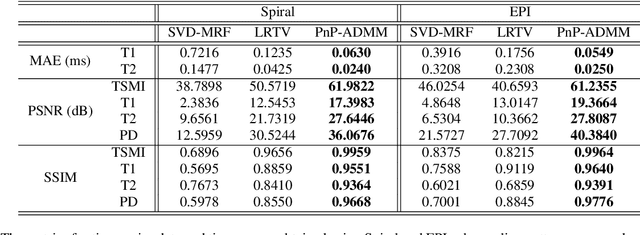
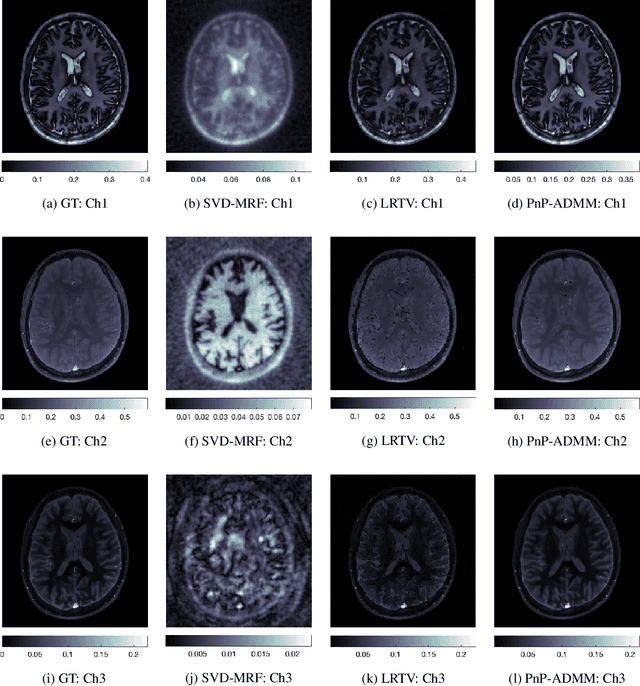
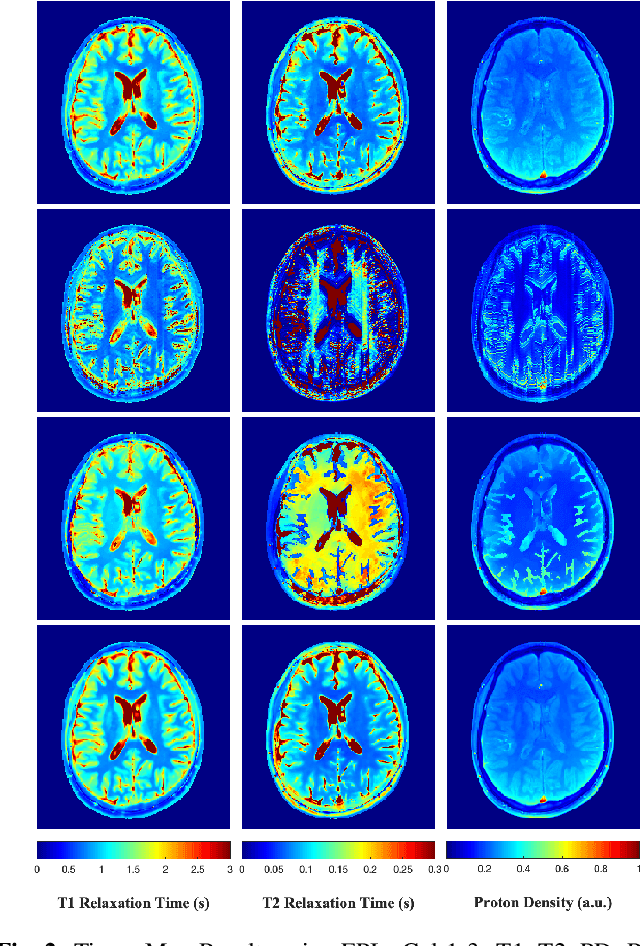
Abstract:Current spatiotemporal deep learning approaches to Magnetic Resonance Fingerprinting (MRF) build artefact-removal models customised to a particular k-space subsampling pattern which is used for fast (compressed) acquisition. This may not be useful when the acquisition process is unknown during training of the deep learning model and/or changes during testing time. This paper proposes an iterative deep learning plug-and-play reconstruction approach to MRF which is adaptive to the forward acquisition process. Spatiotemporal image priors are learned by an image denoiser i.e. a Convolutional Neural Network (CNN), trained to remove generic white gaussian noise (not a particular subsampling artefact) from data. This CNN denoiser is then used as a data-driven shrinkage operator within the iterative reconstruction algorithm. This algorithm with the same denoiser model is then tested on two simulated acquisition processes with distinct subsampling patterns. The results show consistent de-aliasing performance against both acquisition schemes and accurate mapping of tissues' quantitative bio-properties. Software available: https://github.com/ketanfatania/QMRI-PnP-Recon-POC
Deep Unrolling for Magnetic Resonance Fingerprinting
Jan 25, 2022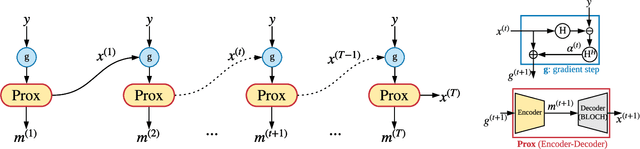
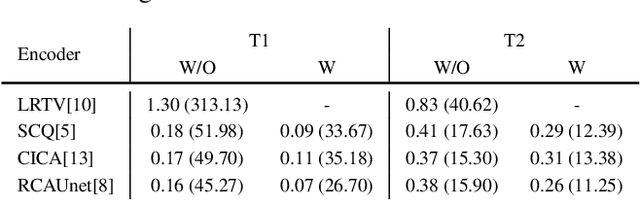
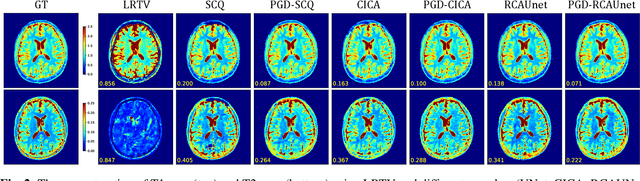
Abstract:Magnetic Resonance Fingerprinting (MRF) has emerged as a promising quantitative MR imaging approach. Deep learning methods have been proposed for MRF and demonstrated improved performance over classical compressed sensing algorithms. However many of these end-to-end models are physics-free, while consistency of the predictions with respect to the physical forward model is crucial for reliably solving inverse problems. To address this, recently [1] proposed a proximal gradient descent framework that directly incorporates the forward acquisition and Bloch dynamic models within an unrolled learning mechanism. However, [1] only evaluated the unrolled model on synthetic data using Cartesian sampling trajectories. In this paper, as a complementary to [1], we investigate other choices of encoders to build the proximal neural network, and evaluate the deep unrolling algorithm on real accelerated MRF scans with non-Cartesian k-space sampling trajectories.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge