Minhaj Nur Alam
Anatomy-Aware Unsupervised Detection and Localization of Retinal Abnormalities in Optical Coherence Tomography
Apr 24, 2026Abstract:Reliable automated analysis of Optical Coherence Tomography (OCT) imaging is crucial for diagnosing retinal disorders but faces a critical barrier: the need for expensive, labor-intensive expert annotations. Supervised deep learning models struggle to generalize across diverse pathologies, imaging devices, and patient populations due to their restricted vocabulary of annotated abnormalities. We propose an unsupervised anomaly detection framework that learns the normative distribution of healthy retinal anatomy without lesion annotations, directly addressing annotation efficiency challenges in clinical deployment. Our approach leverages a discrete latent model trained on normal B-scans to capture OCT-specific structural patterns. To enhance clinical robustness, we incorporate retinal layer-aware supervision and structured triplet learning to separate healthy from pathological representations, improving model reliability across varied imaging conditions. During inference, anomalies are detected and localized via reconstruction discrepancies, enabling both image and pixel-level identification without requiring disease-specific labels. On the Kermany dataset (AUROC: 0.799), our method substantially outperforms VAE, VQVAE, VQGAN, and f-AnoGAN baselines. Critically, cross-dataset evaluation on Srinivasan achieves AUROC 0.884 with superior generalization, demonstrating robust domain adaptation. On the external RETOUCH benchmark, unsupervised anomaly segmentation achieves competitive Dice (0.200) and mIoU (0.117) scores, validating reproducibility across institutions.
FedSIR: Spectral Client Identification and Relabeling for Federated Learning with Noisy Labels
Apr 22, 2026Abstract:Federated learning (FL) enables collaborative model training without sharing raw data; however, the presence of noisy labels across distributed clients can severely degrade the learning performance. In this paper, we propose FedSIR, a multi-stage framework for robust FL under noisy labels. Different from existing approaches that mainly rely on designing noise-tolerant loss functions or exploiting loss dynamics during training, our method leverages the spectral structure of client feature representations to identify and mitigate label noise. Our framework consists of three key components. First, we identify clean and noisy clients by analyzing the spectral consistency of class-wise feature subspaces with minimal communication overhead. Second, clean clients provide spectral references that enable noisy clients to relabel potentially corrupted samples using both dominant class directions and residual subspaces. Third, we employ a noise-aware training strategy that integrates logit-adjusted loss, knowledge distillation, and distance-aware aggregation to further stabilize federated optimization. Extensive experiments on standard FL benchmarks demonstrate that FedSIR consistently outperforms state-of-the-art methods for FL with noisy labels. The code is available at https://github.com/sinagh72/FedSIR.
Multi-OCT-SelfNet: Integrating Self-Supervised Learning with Multi-Source Data Fusion for Enhanced Multi-Class Retinal Disease Classification
Sep 17, 2024



Abstract:In the medical domain, acquiring large datasets poses significant challenges due to privacy concerns. Nonetheless, the development of a robust deep-learning model for retinal disease diagnosis necessitates a substantial dataset for training. The capacity to generalize effectively on smaller datasets remains a persistent challenge. The scarcity of data presents a significant barrier to the practical implementation of scalable medical AI solutions. To address this issue, we've combined a wide range of data sources to improve performance and generalization to new data by giving it a deeper understanding of the data representation from multi-modal datasets and developed a self-supervised framework based on large language models (LLMs), SwinV2 to gain a deeper understanding of multi-modal dataset representations, enhancing the model's ability to extrapolate to new data for the detection of eye diseases using optical coherence tomography (OCT) images. We adopt a two-phase training methodology, self-supervised pre-training, and fine-tuning on a downstream supervised classifier. An ablation study conducted across three datasets employing various encoder backbones, without data fusion, with low data availability setting, and without self-supervised pre-training scenarios, highlights the robustness of our method. Our findings demonstrate consistent performance across these diverse conditions, showcasing superior generalization capabilities compared to the baseline model, ResNet-50.
Quantitative Characterization of Retinal Features in Translated OCTA
Apr 24, 2024



Abstract:Purpose: This study explores the feasibility of using generative machine learning (ML) to translate Optical Coherence Tomography (OCT) images into Optical Coherence Tomography Angiography (OCTA) images, potentially bypassing the need for specialized OCTA hardware. Methods: The method involved implementing a generative adversarial network framework that includes a 2D vascular segmentation model and a 2D OCTA image translation model. The study utilizes a public dataset of 500 patients, divided into subsets based on resolution and disease status, to validate the quality of TR-OCTA images. The validation employs several quality and quantitative metrics to compare the translated images with ground truth OCTAs (GT-OCTA). We then quantitatively characterize vascular features generated in TR-OCTAs with GT-OCTAs to assess the feasibility of using TR-OCTA for objective disease diagnosis. Result: TR-OCTAs showed high image quality in both 3 and 6 mm datasets (high-resolution, moderate structural similarity and contrast quality compared to GT-OCTAs). There were slight discrepancies in vascular metrics, especially in diseased patients. Blood vessel features like tortuosity and vessel perimeter index showed a better trend compared to density features which are affected by local vascular distortions. Conclusion: This study presents a promising solution to the limitations of OCTA adoption in clinical practice by using vascular features from TR-OCTA for disease detection. Translation relevance: This study has the potential to significantly enhance the diagnostic process for retinal diseases by making detailed vascular imaging more widely available and reducing dependency on costly OCTA equipment.
OCT-SelfNet: A Self-Supervised Framework with Multi-Modal Datasets for Generalized and Robust Retinal Disease Detection
Jan 22, 2024



Abstract:Despite the revolutionary impact of AI and the development of locally trained algorithms, achieving widespread generalized learning from multi-modal data in medical AI remains a significant challenge. This gap hinders the practical deployment of scalable medical AI solutions. Addressing this challenge, our research contributes a self-supervised robust machine learning framework, OCT-SelfNet, for detecting eye diseases using optical coherence tomography (OCT) images. In this work, various data sets from various institutions are combined enabling a more comprehensive range of representation. Our method addresses the issue using a two-phase training approach that combines self-supervised pretraining and supervised fine-tuning with a mask autoencoder based on the SwinV2 backbone by providing a solution for real-world clinical deployment. Extensive experiments on three datasets with different encoder backbones, low data settings, unseen data settings, and the effect of augmentation show that our method outperforms the baseline model, Resnet-50 by consistently attaining AUC-ROC performance surpassing 77% across all tests, whereas the baseline model exceeds 54%. Moreover, in terms of the AUC-PR metric, our proposed method exceeded 42%, showcasing a substantial increase of at least 10% in performance compared to the baseline, which exceeded only 33%. This contributes to our understanding of our approach's potential and emphasizes its usefulness in clinical settings.
Contrastive learning-based pretraining improves representation and transferability of diabetic retinopathy classification models
Aug 24, 2022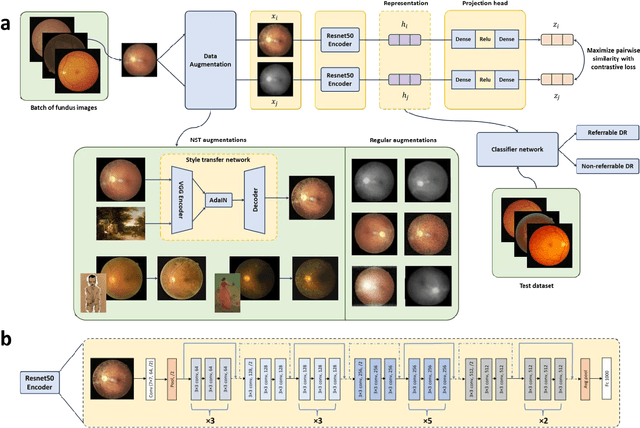
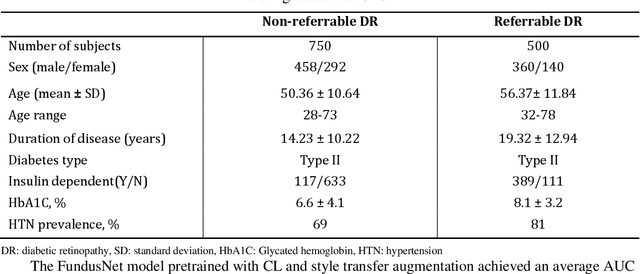

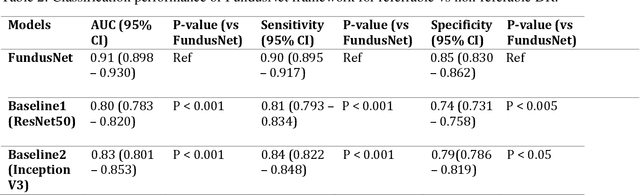
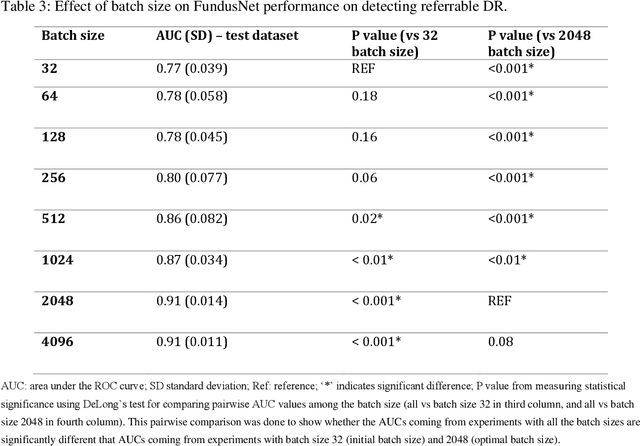
Abstract:Self supervised contrastive learning based pretraining allows development of robust and generalized deep learning models with small, labeled datasets, reducing the burden of label generation. This paper aims to evaluate the effect of CL based pretraining on the performance of referrable vs non referrable diabetic retinopathy (DR) classification. We have developed a CL based framework with neural style transfer (NST) augmentation to produce models with better representations and initializations for the detection of DR in color fundus images. We compare our CL pretrained model performance with two state of the art baseline models pretrained with Imagenet weights. We further investigate the model performance with reduced labeled training data (down to 10 percent) to test the robustness of the model when trained with small, labeled datasets. The model is trained and validated on the EyePACS dataset and tested independently on clinical data from the University of Illinois, Chicago (UIC). Compared to baseline models, our CL pretrained FundusNet model had higher AUC (CI) values (0.91 (0.898 to 0.930) vs 0.80 (0.783 to 0.820) and 0.83 (0.801 to 0.853) on UIC data). At 10 percent labeled training data, the FundusNet AUC was 0.81 (0.78 to 0.84) vs 0.58 (0.56 to 0.64) and 0.63 (0.60 to 0.66) in baseline models, when tested on the UIC dataset. CL based pretraining with NST significantly improves DL classification performance, helps the model generalize well (transferable from EyePACS to UIC data), and allows training with small, annotated datasets, therefore reducing ground truth annotation burden of the clinicians.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge