Matthias Samwald
Benchmark datasets driving artificial intelligence development fail to capture the needs of medical professionals
Jan 18, 2022
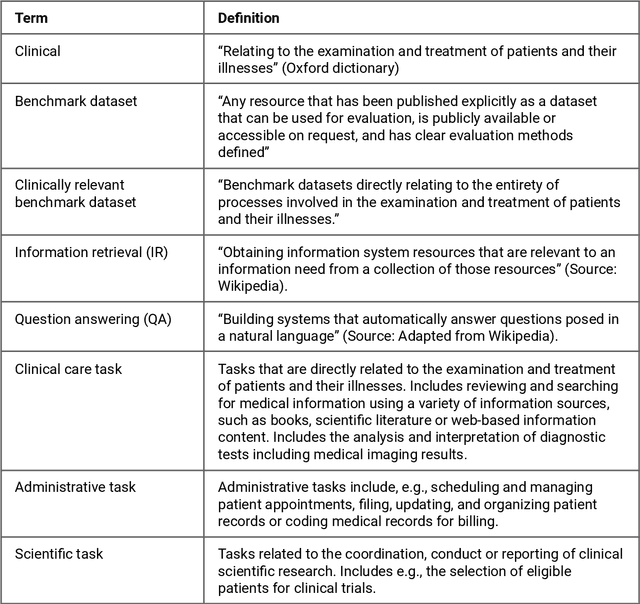
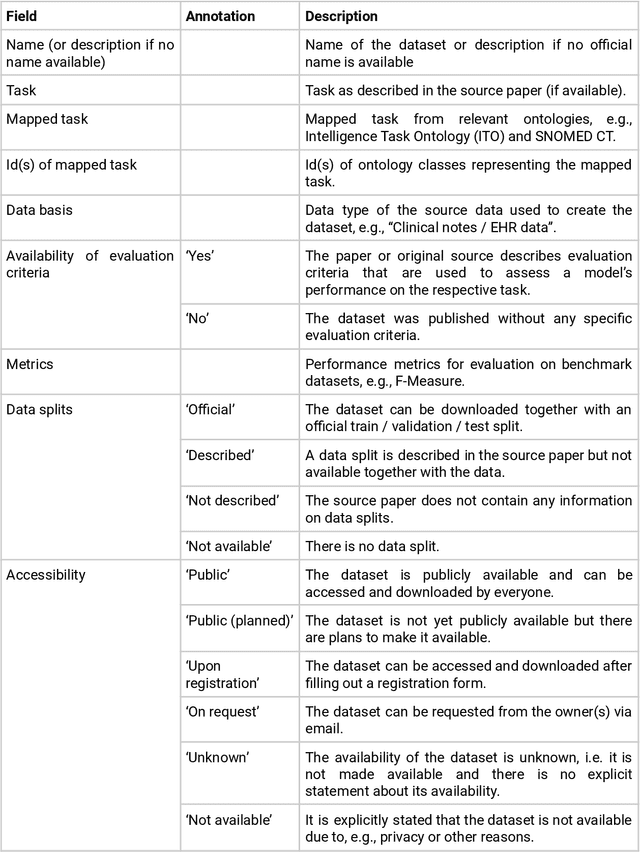
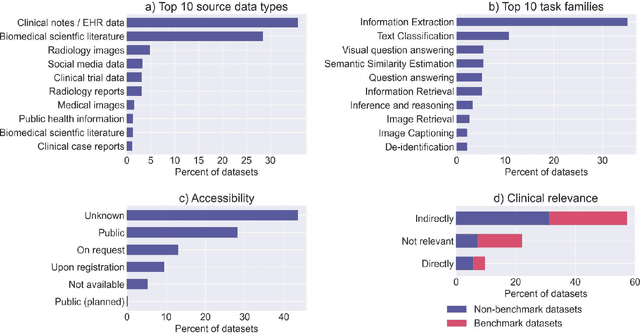
Abstract:Publicly accessible benchmarks that allow for assessing and comparing model performances are important drivers of progress in artificial intelligence (AI). While recent advances in AI capabilities hold the potential to transform medical practice by assisting and augmenting the cognitive processes of healthcare professionals, the coverage of clinically relevant tasks by AI benchmarks is largely unclear. Furthermore, there is a lack of systematized meta-information that allows clinical AI researchers to quickly determine accessibility, scope, content and other characteristics of datasets and benchmark datasets relevant to the clinical domain. To address these issues, we curated and released a comprehensive catalogue of datasets and benchmarks pertaining to the broad domain of clinical and biomedical natural language processing (NLP), based on a systematic review of literature and online resources. A total of 450 NLP datasets were manually systematized and annotated with rich metadata, such as targeted tasks, clinical applicability, data types, performance metrics, accessibility and licensing information, and availability of data splits. We then compared tasks covered by AI benchmark datasets with relevant tasks that medical practitioners reported as highly desirable targets for automation in a previous empirical study. Our analysis indicates that AI benchmarks of direct clinical relevance are scarce and fail to cover most work activities that clinicians want to see addressed. In particular, tasks associated with routine documentation and patient data administration workflows are not represented despite significant associated workloads. Thus, currently available AI benchmarks are improperly aligned with desired targets for AI automation in clinical settings, and novel benchmarks should be created to fill these gaps.
Improving the robustness and accuracy of biomedical language models through adversarial training
Nov 16, 2021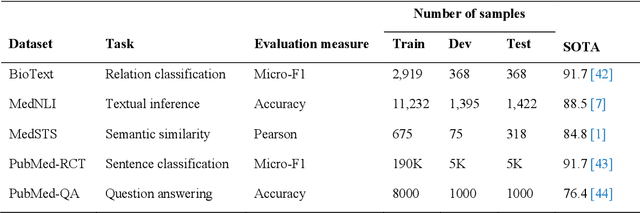
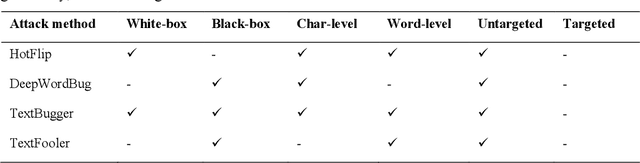
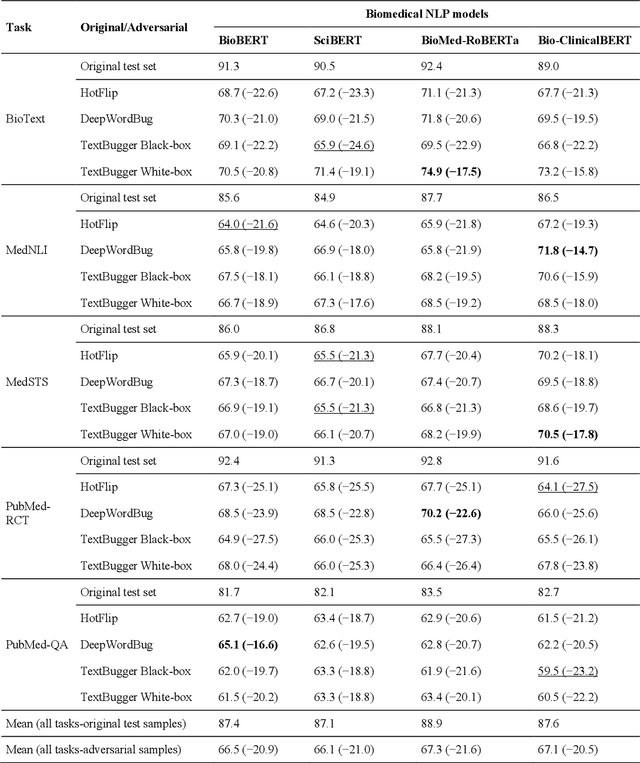
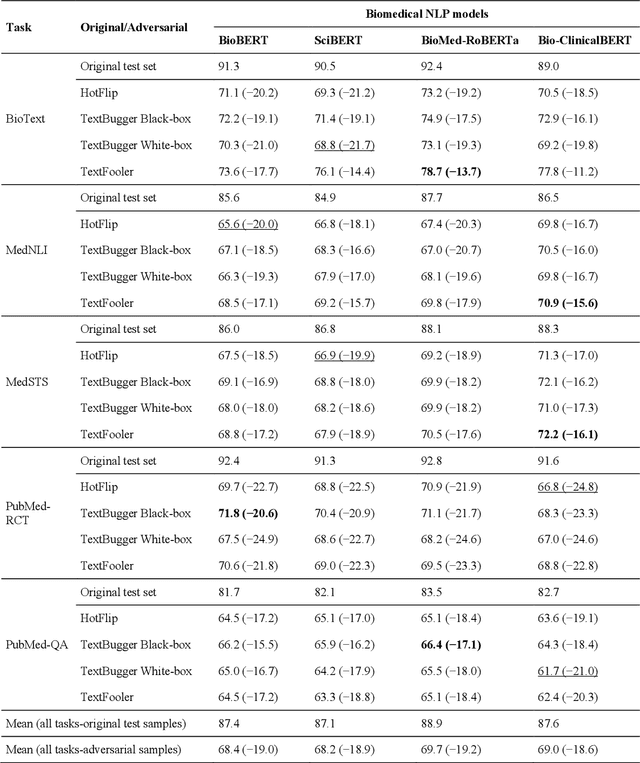
Abstract:Deep transformer neural network models have improved the predictive accuracy of intelligent text processing systems in the biomedical domain. They have obtained state-of-the-art performance scores on a wide variety of biomedical and clinical Natural Language Processing (NLP) benchmarks. However, the robustness and reliability of these models has been less explored so far. Neural NLP models can be easily fooled by adversarial samples, i.e. minor changes to input that preserve the meaning and understandability of the text but force the NLP system to make erroneous decisions. This raises serious concerns about the security and trust-worthiness of biomedical NLP systems, especially when they are intended to be deployed in real-world use cases. We investigated the robustness of several transformer neural language models, i.e. BioBERT, SciBERT, BioMed-RoBERTa, and Bio-ClinicalBERT, on a wide range of biomedical and clinical text processing tasks. We implemented various adversarial attack methods to test the NLP systems in different attack scenarios. Experimental results showed that the biomedical NLP models are sensitive to adversarial samples; their performance dropped in average by 21 and 18.9 absolute percent on character-level and word-level adversarial noise, respectively. Conducting extensive adversarial training experiments, we fine-tuned the NLP models on a mixture of clean samples and adversarial inputs. Results showed that adversarial training is an effective defense mechanism against adversarial noise; the models robustness improved in average by 11.3 absolute percent. In addition, the models performance on clean data increased in average by 2.4 absolute present, demonstrating that adversarial training can boost generalization abilities of biomedical NLP systems.
A curated, ontology-based, large-scale knowledge graph of artificial intelligence tasks and benchmarks
Oct 06, 2021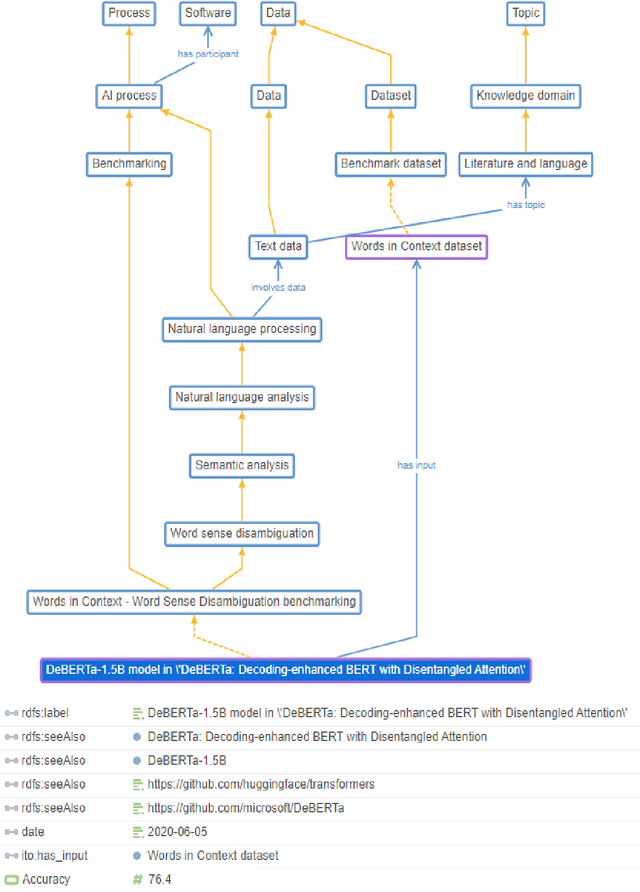
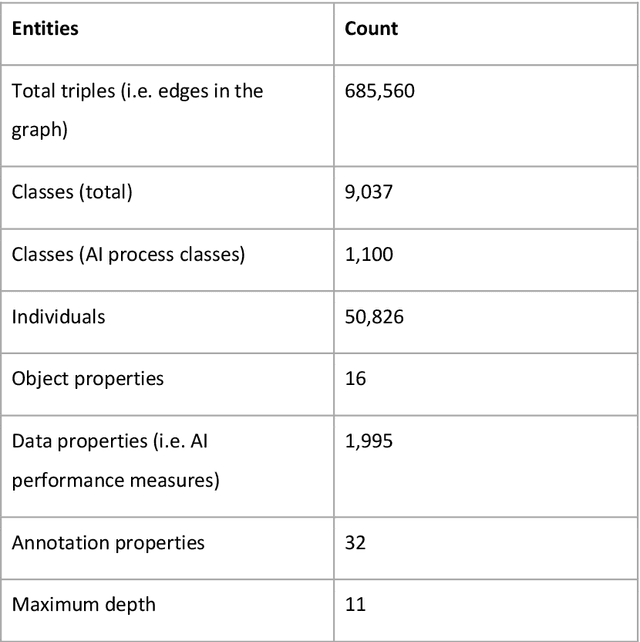
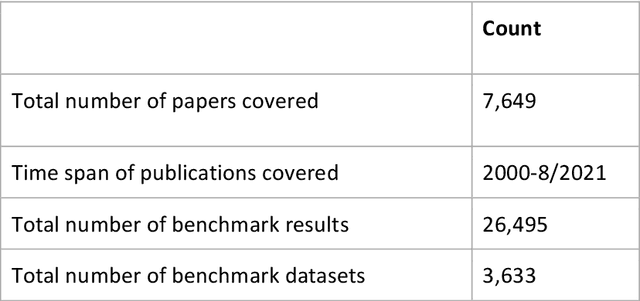
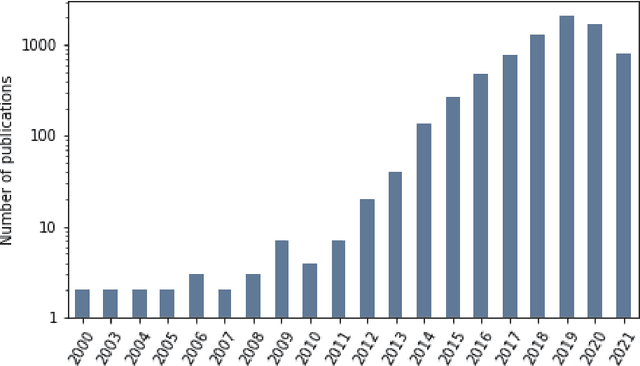
Abstract:Research in artificial intelligence (AI) is addressing a growing number of tasks through a rapidly growing number of models and methodologies. This makes it difficult to keep track of where novel AI methods are successfully -- or still unsuccessfully -- applied, how progress is measured, how different advances might synergize with each other, and how future research should be prioritized. To help address these issues, we created the Intelligence Task Ontology and Knowledge Graph (ITO), a comprehensive, richly structured and manually curated resource on artificial intelligence tasks, benchmark results and performance metrics. The current version of ITO contain 685,560 edges, 1,100 classes representing AI processes and 1,995 properties representing performance metrics. The goal of ITO is to enable precise and network-based analyses of the global landscape of AI tasks and capabilities. ITO is based on technologies that allow for easy integration and enrichment with external data, automated inference and continuous, collaborative expert curation of underlying ontological models. We make the ITO dataset and a collection of Jupyter notebooks utilising ITO openly available.
SAFRAN: An interpretable, rule-based link prediction method outperforming embedding models
Sep 16, 2021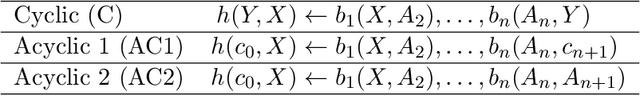
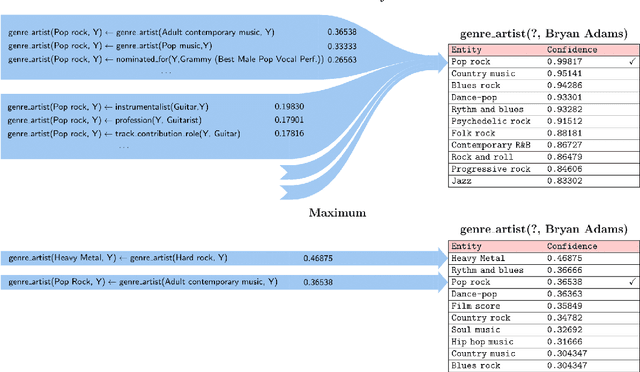

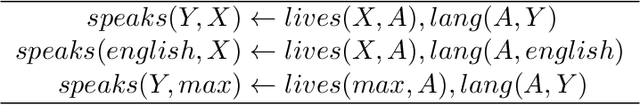
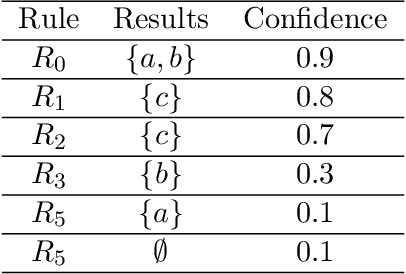
Abstract:Neural embedding-based machine learning models have shown promise for predicting novel links in knowledge graphs. Unfortunately, their practical utility is diminished by their lack of interpretability. Recently, the fully interpretable, rule-based algorithm AnyBURL yielded highly competitive results on many general-purpose link prediction benchmarks. However, current approaches for aggregating predictions made by multiple rules are affected by redundancies. We improve upon AnyBURL by introducing the SAFRAN rule application framework, which uses a novel aggregation approach called Non-redundant Noisy-OR that detects and clusters redundant rules prior to aggregation. SAFRAN yields new state-of-the-art results for fully interpretable link prediction on the established general-purpose benchmarks FB15K-237, WN18RR and YAGO3-10. Furthermore, it exceeds the results of multiple established embedding-based algorithms on FB15K-237 and WN18RR and narrows the gap between rule-based and embedding-based algorithms on YAGO3-10.
GPT-3 Models are Poor Few-Shot Learners in the Biomedical Domain
Sep 06, 2021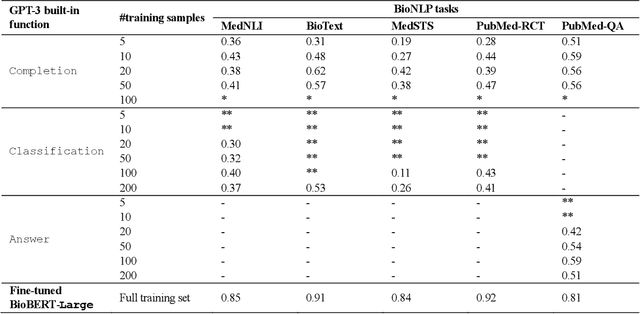
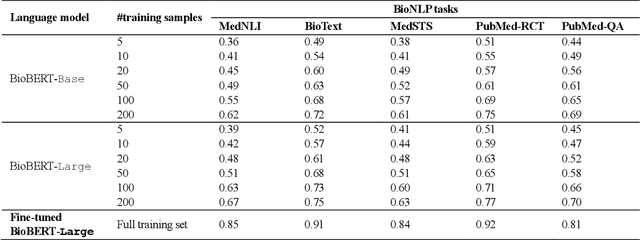
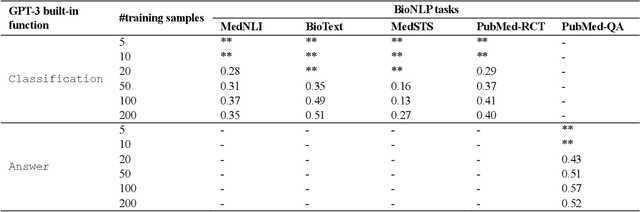
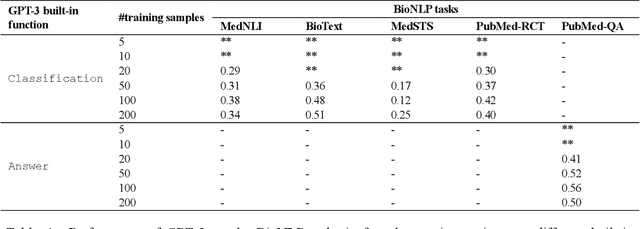
Abstract:Deep neural language models have set new breakthroughs in many tasks of Natural Language Processing (NLP). Recent work has shown that deep transformer language models (pretrained on large amounts of texts) can achieve high levels of task-specific few-shot performance comparable to state-of-the-art models. However, the ability of these large language models in few-shot transfer learning has not yet been explored in the biomedical domain. We investigated the performance of two powerful transformer language models, i.e. GPT-3 and BioBERT, in few-shot settings on various biomedical NLP tasks. The experimental results showed that, to a great extent, both the models underperform a language model fine-tuned on the full training data. Although GPT-3 had already achieved near state-of-the-art results in few-shot knowledge transfer on open-domain NLP tasks, it could not perform as effectively as BioBERT, which is orders of magnitude smaller than GPT-3. Regarding that BioBERT was already pretrained on large biomedical text corpora, our study suggests that language models may largely benefit from in-domain pretraining in task-specific few-shot learning. However, in-domain pretraining seems not to be sufficient; novel pretraining and few-shot learning strategies are required in the biomedical NLP domain.
Deep learning models are not robust against noise in clinical text
Aug 27, 2021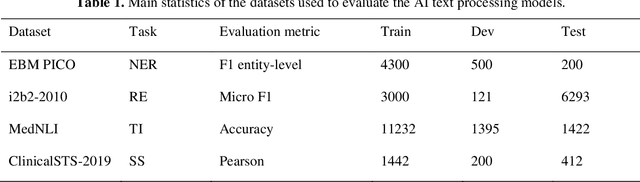
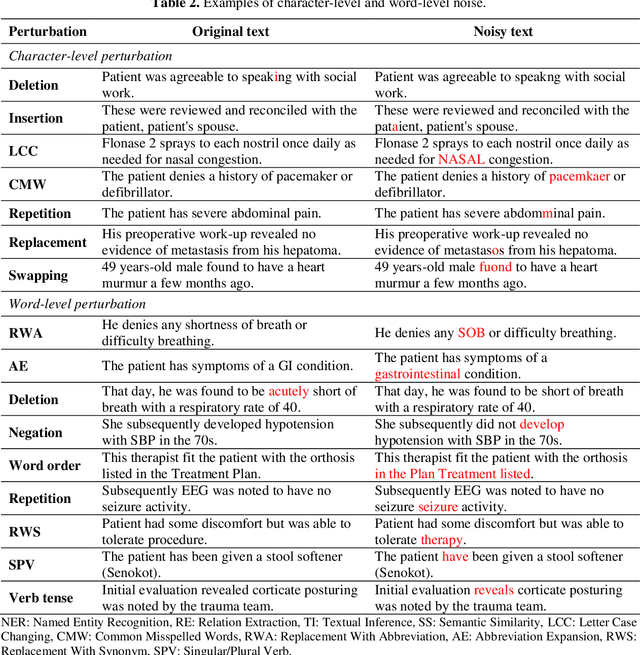
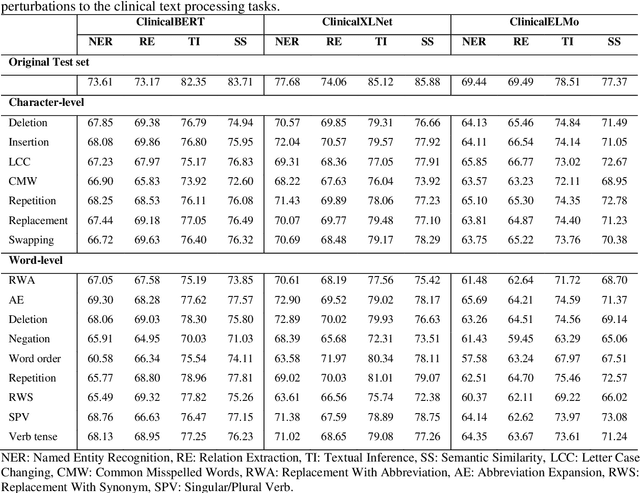
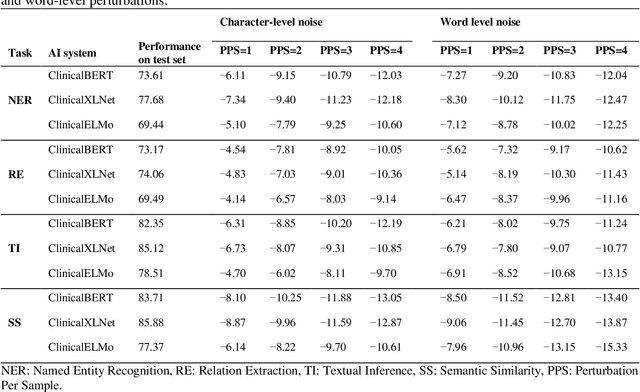
Abstract:Artificial Intelligence (AI) systems are attracting increasing interest in the medical domain due to their ability to learn complicated tasks that require human intelligence and expert knowledge. AI systems that utilize high-performance Natural Language Processing (NLP) models have achieved state-of-the-art results on a wide variety of clinical text processing benchmarks. They have even outperformed human accuracy on some tasks. However, performance evaluation of such AI systems have been limited to accuracy measures on curated and clean benchmark datasets that may not properly reflect how robustly these systems can operate in real-world situations. In order to address this challenge, we introduce and implement a wide variety of perturbation methods that simulate different types of noise and variability in clinical text data. While noisy samples produced by these perturbation methods can often be understood by humans, they may cause AI systems to make erroneous decisions. Conducting extensive experiments on several clinical text processing tasks, we evaluated the robustness of high-performance NLP models against various types of character-level and word-level noise. The results revealed that the NLP models performance degrades when the input contains small amounts of noise. This study is a significant step towards exposing vulnerabilities of AI models utilized in clinical text processing systems. The proposed perturbation methods can be used in performance evaluation tests to assess how robustly clinical NLP models can operate on noisy data, in real-world settings.
Evaluating the Robustness of Neural Language Models to Input Perturbations
Aug 27, 2021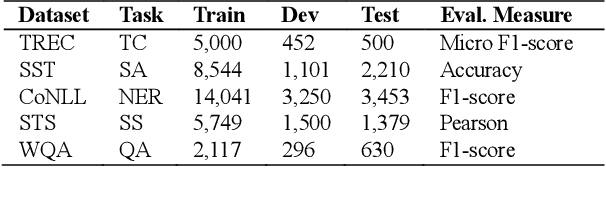
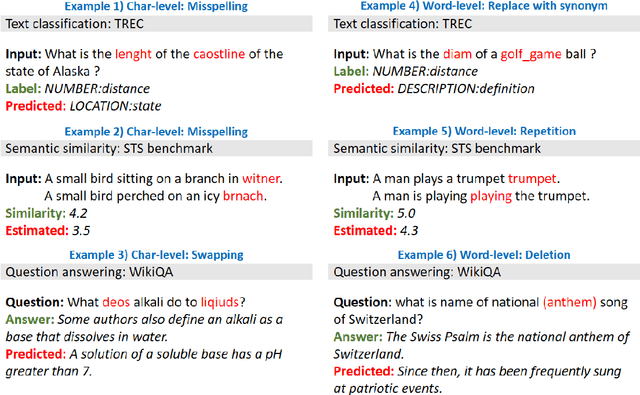
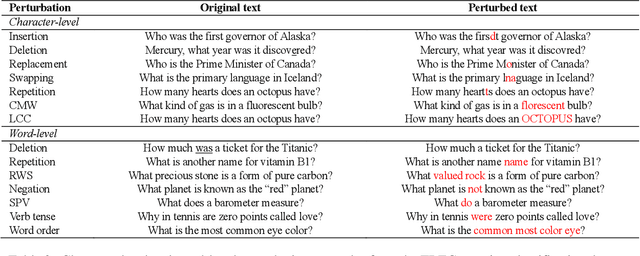
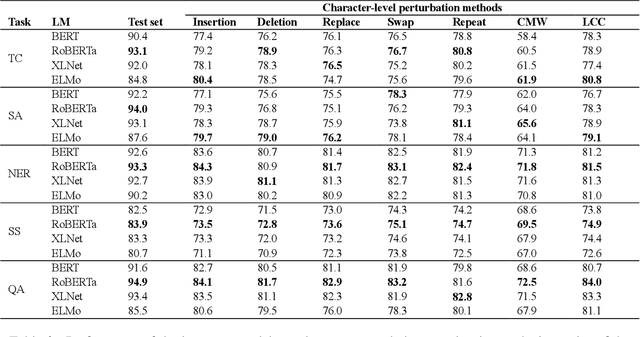
Abstract:High-performance neural language models have obtained state-of-the-art results on a wide range of Natural Language Processing (NLP) tasks. However, results for common benchmark datasets often do not reflect model reliability and robustness when applied to noisy, real-world data. In this study, we design and implement various types of character-level and word-level perturbation methods to simulate realistic scenarios in which input texts may be slightly noisy or different from the data distribution on which NLP systems were trained. Conducting comprehensive experiments on different NLP tasks, we investigate the ability of high-performance language models such as BERT, XLNet, RoBERTa, and ELMo in handling different types of input perturbations. The results suggest that language models are sensitive to input perturbations and their performance can decrease even when small changes are introduced. We highlight that models need to be further improved and that current benchmarks are not reflecting model robustness well. We argue that evaluations on perturbed inputs should routinely complement widely-used benchmarks in order to yield a more realistic understanding of NLP systems robustness.
Explaining Black-box Models for Biomedical Text Classification
Dec 20, 2020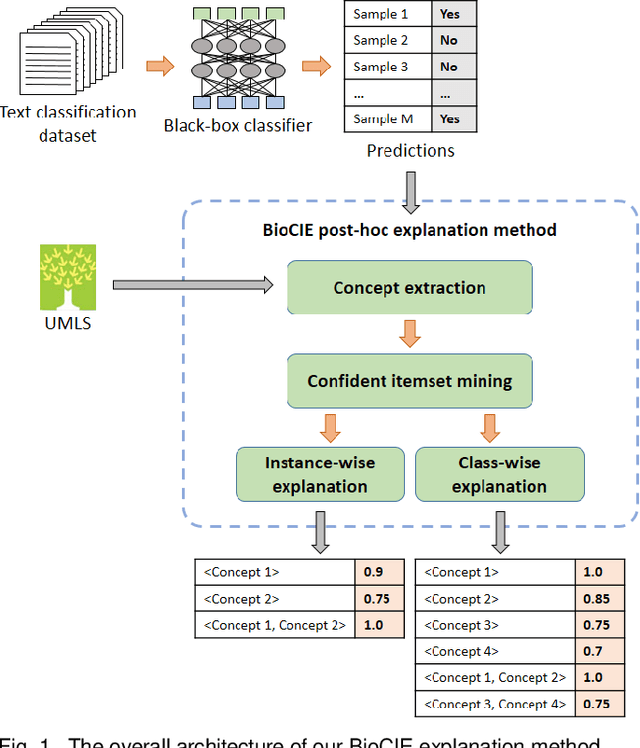

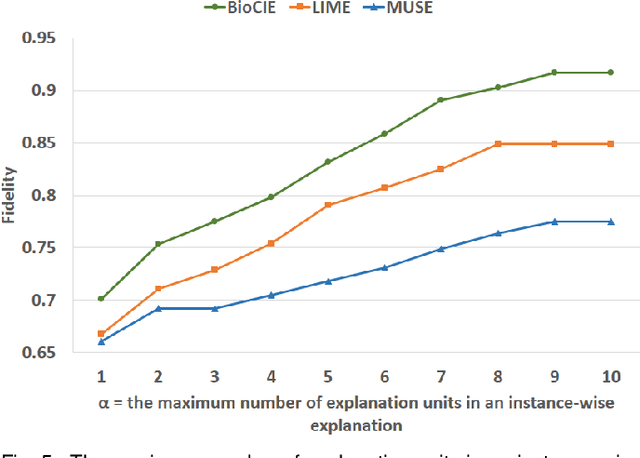
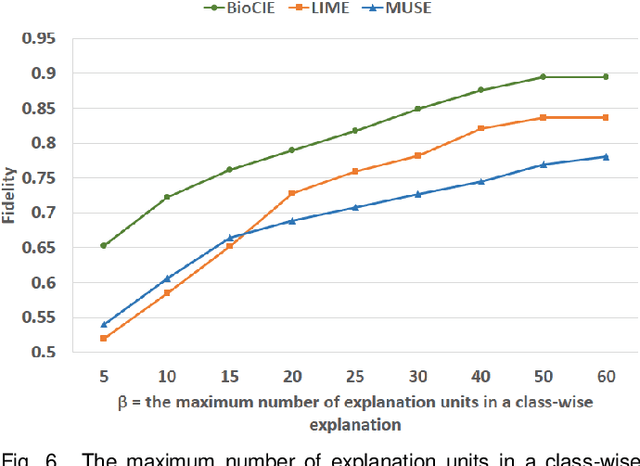
Abstract:In this paper, we propose a novel method named Biomedical Confident Itemsets Explanation (BioCIE), aiming at post-hoc explanation of black-box machine learning models for biomedical text classification. Using sources of domain knowledge and a confident itemset mining method, BioCIE discretizes the decision space of a black-box into smaller subspaces and extracts semantic relationships between the input text and class labels in different subspaces. Confident itemsets discover how biomedical concepts are related to class labels in the black-box's decision space. BioCIE uses the itemsets to approximate the black-box's behavior for individual predictions. Optimizing fidelity, interpretability, and coverage measures, BioCIE produces class-wise explanations that represent decision boundaries of the black-box. Results of evaluations on various biomedical text classification tasks and black-box models demonstrated that BioCIE can outperform perturbation-based and decision set methods in terms of producing concise, accurate, and interpretable explanations. BioCIE improved the fidelity of instance-wise and class-wise explanations by 11.6% and 7.5%, respectively. It also improved the interpretability of explanations by 8%. BioCIE can be effectively used to explain how a black-box biomedical text classification model semantically relates input texts to class labels. The source code and supplementary material are available at https://github.com/mmoradi-iut/BioCIE.
Scalable and interpretable rule-based link prediction for large heterogeneous knowledge graphs
Dec 10, 2020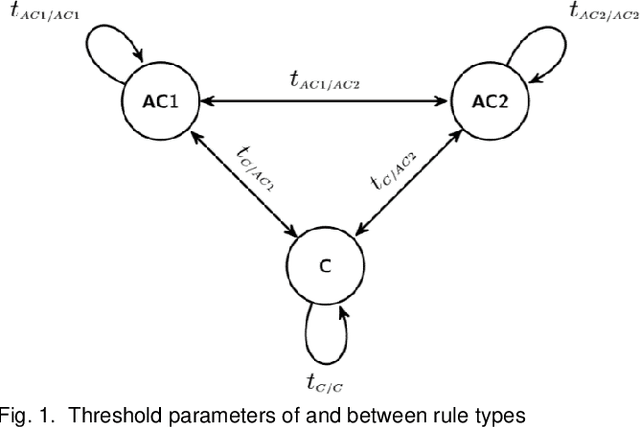
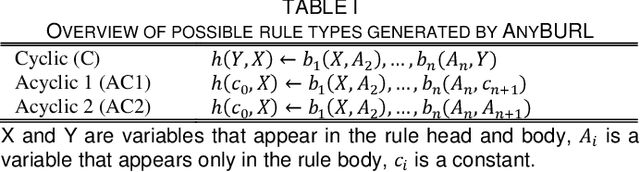


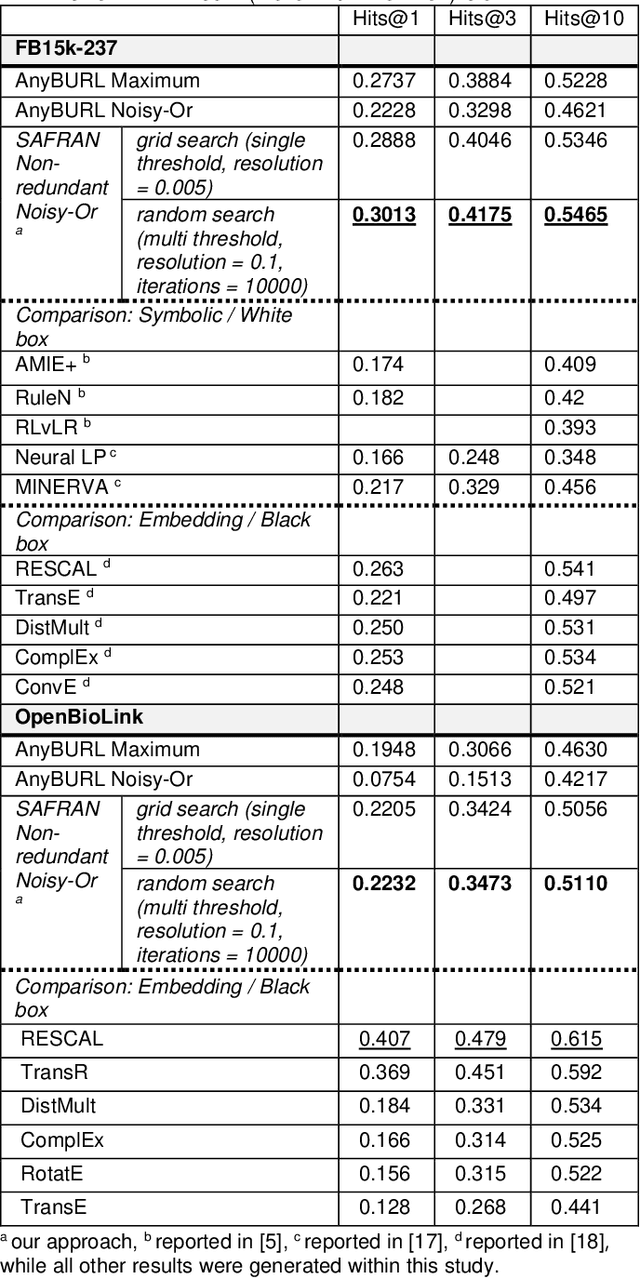
Abstract:Neural embedding-based machine learning models have shown promise for predicting novel links in biomedical knowledge graphs. Unfortunately, their practical utility is diminished by their lack of interpretability. Recently, the fully interpretable, rule-based algorithm AnyBURL yielded highly competitive results on many general-purpose link prediction benchmarks. However, its applicability to large-scale prediction tasks on complex biomedical knowledge bases is limited by long inference times and difficulties with aggregating predictions made by multiple rules. We improve upon AnyBURL by introducing the SAFRAN rule application framework which aggregates rules through a scalable clustering algorithm. SAFRAN yields new state-of-the-art results for fully interpretable link prediction on the established general-purpose benchmark FB15K-237 and the large-scale biomedical benchmark OpenBioLink. Furthermore, it exceeds the results of multiple established embedding-based algorithms on FB15K-237 and narrows the gap between rule-based and embedding-based algorithms on OpenBioLink. We also show that SAFRAN increases inference speeds by up to two orders of magnitude.
Explaining black-box text classifiers for disease-treatment information extraction
Oct 21, 2020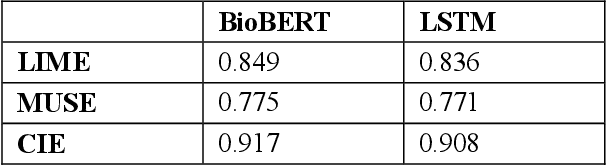
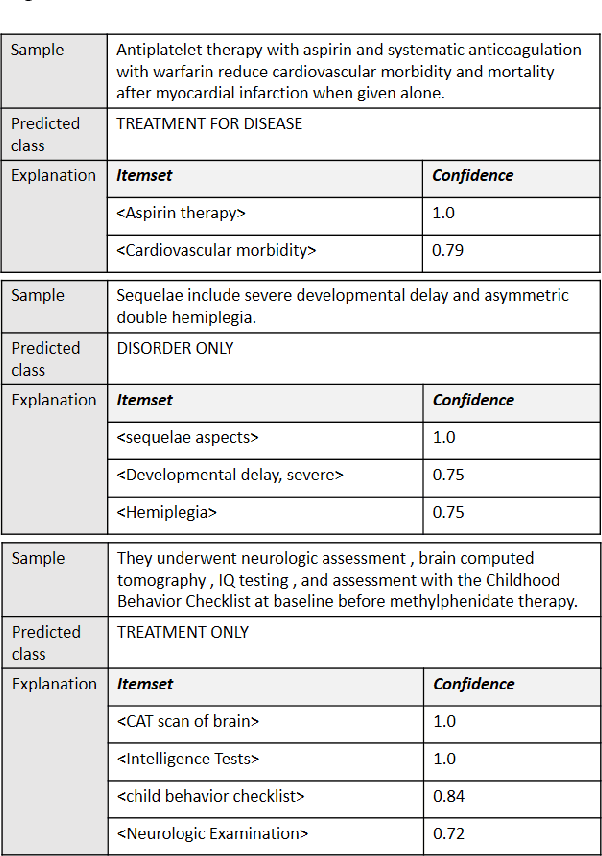
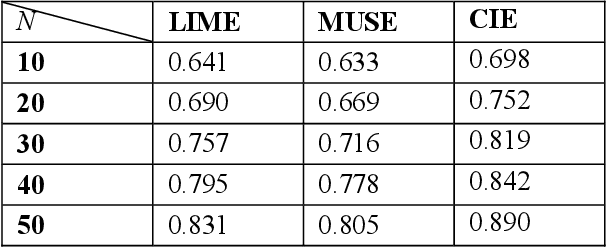
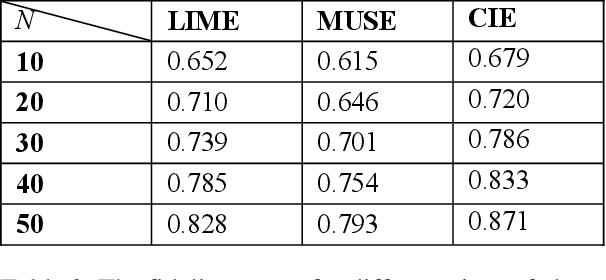
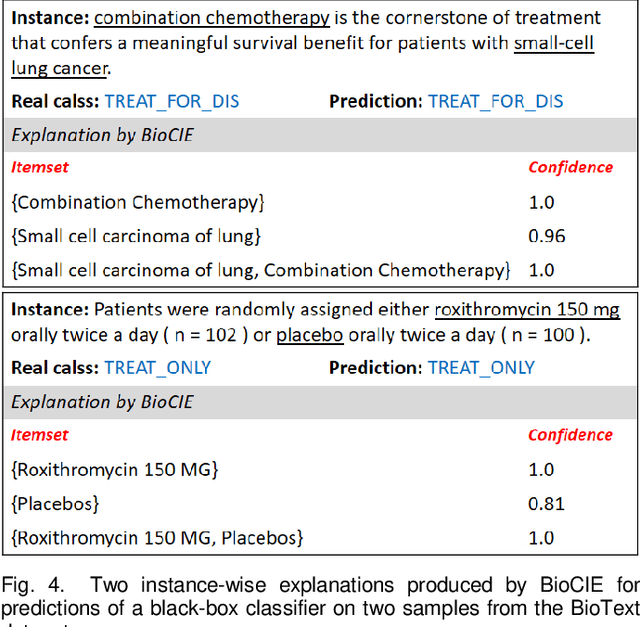
Abstract:Deep neural networks and other intricate Artificial Intelligence (AI) models have reached high levels of accuracy on many biomedical natural language processing tasks. However, their applicability in real-world use cases may be limited due to their vague inner working and decision logic. A post-hoc explanation method can approximate the behavior of a black-box AI model by extracting relationships between feature values and outcomes. In this paper, we introduce a post-hoc explanation method that utilizes confident itemsets to approximate the behavior of black-box classifiers for medical information extraction. Incorporating medical concepts and semantics into the explanation process, our explanator finds semantic relations between inputs and outputs in different parts of the decision space of a black-box classifier. The experimental results show that our explanation method can outperform perturbation and decision set based explanators in terms of fidelity and interpretability of explanations produced for predictions on a disease-treatment information extraction task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge