Matthew Scicluna
scShapeBench: Discovering geometry from high dimensional scRNAseq data
May 12, 2026Abstract:High-dimensional point cloud data arise across many scientific domains, especially single-cell biology. The shapes or topologies of these datasets determine the types of information that can be extracted. For example, clustered data supports cell-type identification, trajectory structures support transition analysis, and archetypal structures capture continua of cellular behaviors. Existing analysis pipelines often assume a specific shape. The standard Seurat pipeline combines UMAP visualization with Louvain clustering and therefore assumes clustered data, while tools such as Monocle and SPADE assume tree-like structures, and flow-based models such as MIOFlow and Conditional Flow Matching target trajectories. Choosing which pipeline to apply is therefore often left to bioinformaticians who visually inspect datasets before selecting an analysis strategy. With the rise of agentic AI scientists, automating shape detection is increasingly important for selecting downstream analysis pipelines. To address this problem, we introduce scShapeBench, a benchmark dataset for shape detection containing both synthetic and expert-annotated single-cell datasets. Synthetic datasets are sampled from ground-truth skeleton graphs with controlled variance. Real single-cell datasets are curated from diverse sources and annotated by experts into four categories: clusters, single trajectory, multi-branching, and archetypal. We additionally introduce scReebTower, a baseline method that uses diffusion geometry to extract Reeb graphs and connect visualization with pipeline selection. We provide topology-aware evaluation metrics and compare scReebTower against PAGA and Mapper on synthetic and real data. Our results indicate that scReebTower outperforms existing baselines. Overall, our contributions span benchmarks, evaluation metrics, and a baseline for automated shape detection in single-cell data.
A Modern Take on the Bias-Variance Tradeoff in Neural Networks
Oct 19, 2018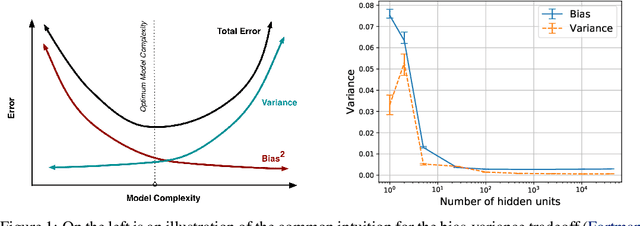
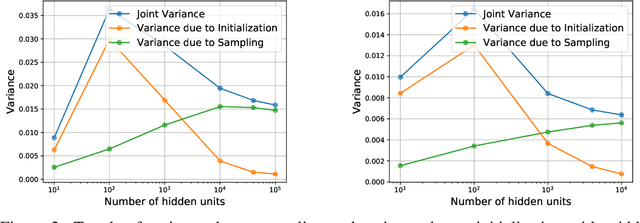
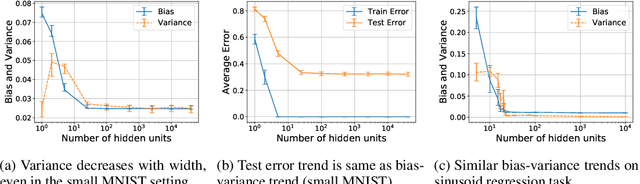
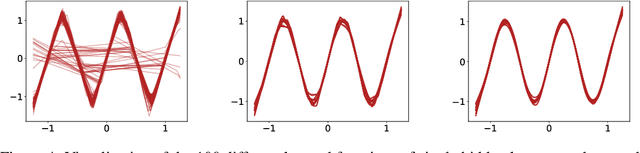
Abstract:We revisit the bias-variance tradeoff for neural networks in light of modern empirical findings. The traditional bias-variance tradeoff in machine learning suggests that as model complexity grows, variance increases. Classical bounds in statistical learning theory point to the number of parameters in a model as a measure of model complexity, which means the tradeoff would indicate that variance increases with the size of neural networks. However, we empirically find that variance due to training set sampling is roughly \textit{constant} (with both width and depth) in practice. Variance caused by the non-convexity of the loss landscape is different. We find that it decreases with width and increases with depth, in our setting. We provide theoretical analysis, in a simplified setting inspired by linear models, that is consistent with our empirical findings for width. We view bias-variance as a useful lens to study generalization through and encourage further theoretical explanation from this perspective.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge