Mario Krenn
On scientific understanding with artificial intelligence
Apr 04, 2022
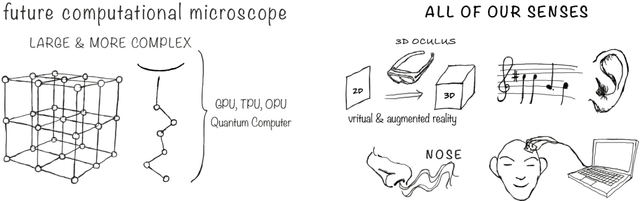
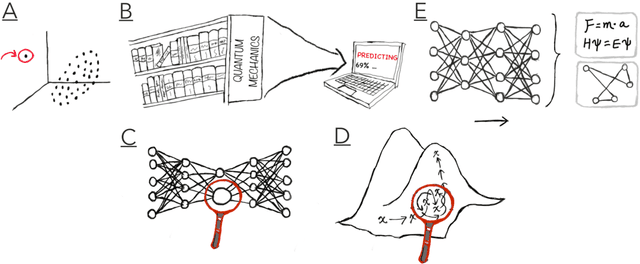
Abstract:Imagine an oracle that correctly predicts the outcome of every particle physics experiment, the products of every chemical reaction, or the function of every protein. Such an oracle would revolutionize science and technology as we know them. However, as scientists, we would not be satisfied with the oracle itself. We want more. We want to comprehend how the oracle conceived these predictions. This feat, denoted as scientific understanding, has frequently been recognized as the essential aim of science. Now, the ever-growing power of computers and artificial intelligence poses one ultimate question: How can advanced artificial systems contribute to scientific understanding or achieve it autonomously? We are convinced that this is not a mere technical question but lies at the core of science. Therefore, here we set out to answer where we are and where we can go from here. We first seek advice from the philosophy of science to understand scientific understanding. Then we review the current state of the art, both from literature and by collecting dozens of anecdotes from scientists about how they acquired new conceptual understanding with the help of computers. Those combined insights help us to define three dimensions of android-assisted scientific understanding: The android as a I) computational microscope, II) resource of inspiration and the ultimate, not yet existent III) agent of understanding. For each dimension, we explain new avenues to push beyond the status quo and unleash the full power of artificial intelligence's contribution to the central aim of science. We hope our perspective inspires and focuses research towards androids that get new scientific understanding and ultimately bring us closer to true artificial scientists.
SELFIES and the future of molecular string representations
Mar 31, 2022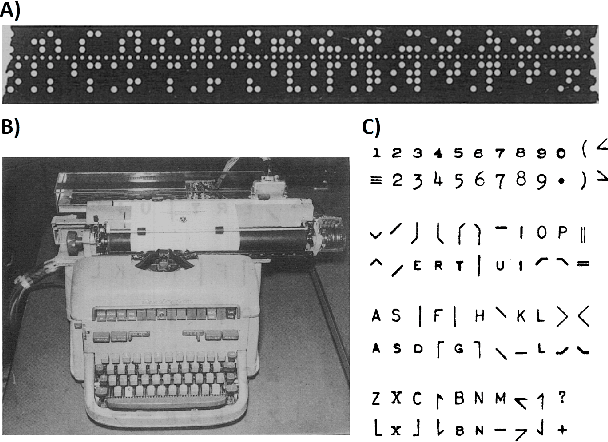
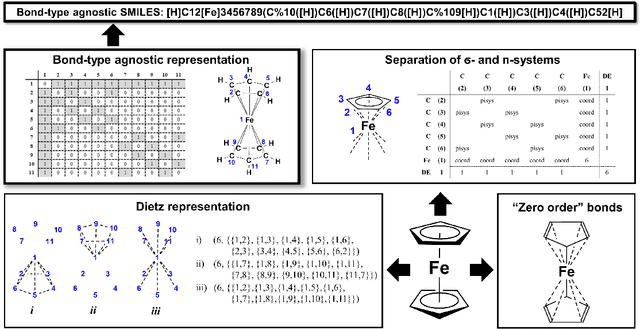

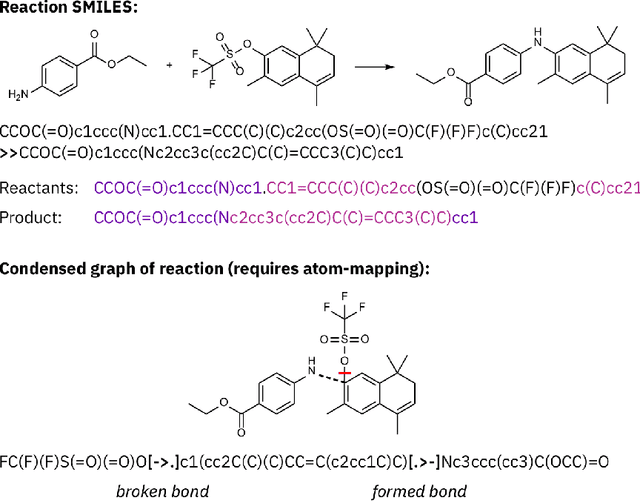
Abstract:Artificial intelligence (AI) and machine learning (ML) are expanding in popularity for broad applications to challenging tasks in chemistry and materials science. Examples include the prediction of properties, the discovery of new reaction pathways, or the design of new molecules. The machine needs to read and write fluently in a chemical language for each of these tasks. Strings are a common tool to represent molecular graphs, and the most popular molecular string representation, SMILES, has powered cheminformatics since the late 1980s. However, in the context of AI and ML in chemistry, SMILES has several shortcomings -- most pertinently, most combinations of symbols lead to invalid results with no valid chemical interpretation. To overcome this issue, a new language for molecules was introduced in 2020 that guarantees 100\% robustness: SELFIES (SELF-referencIng Embedded Strings). SELFIES has since simplified and enabled numerous new applications in chemistry. In this manuscript, we look to the future and discuss molecular string representations, along with their respective opportunities and challenges. We propose 16 concrete Future Projects for robust molecular representations. These involve the extension toward new chemical domains, exciting questions at the interface of AI and robust languages and interpretability for both humans and machines. We hope that these proposals will inspire several follow-up works exploiting the full potential of molecular string representations for the future of AI in chemistry and materials science.
Design of quantum optical experiments with logic artificial intelligence
Sep 27, 2021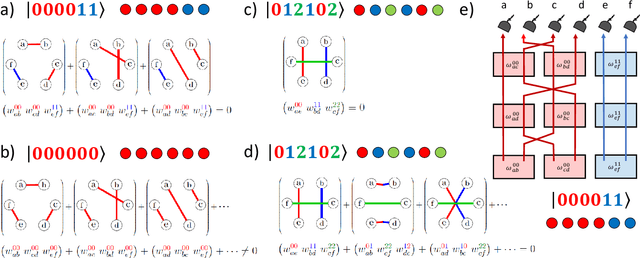
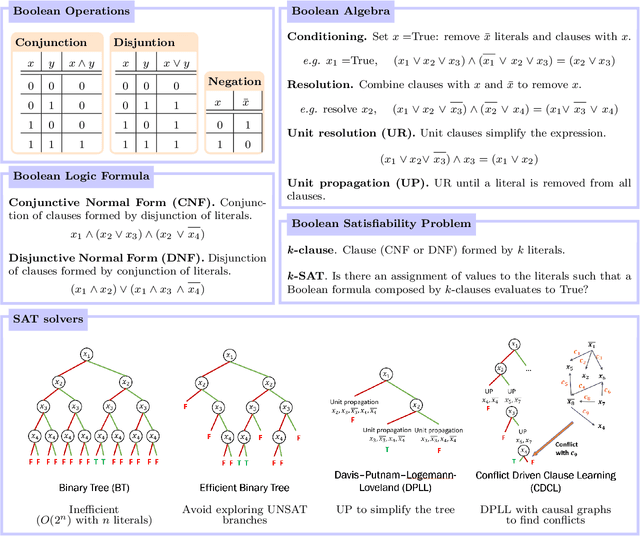
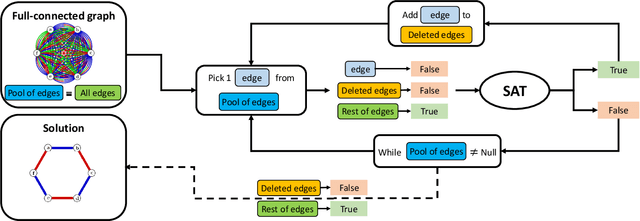
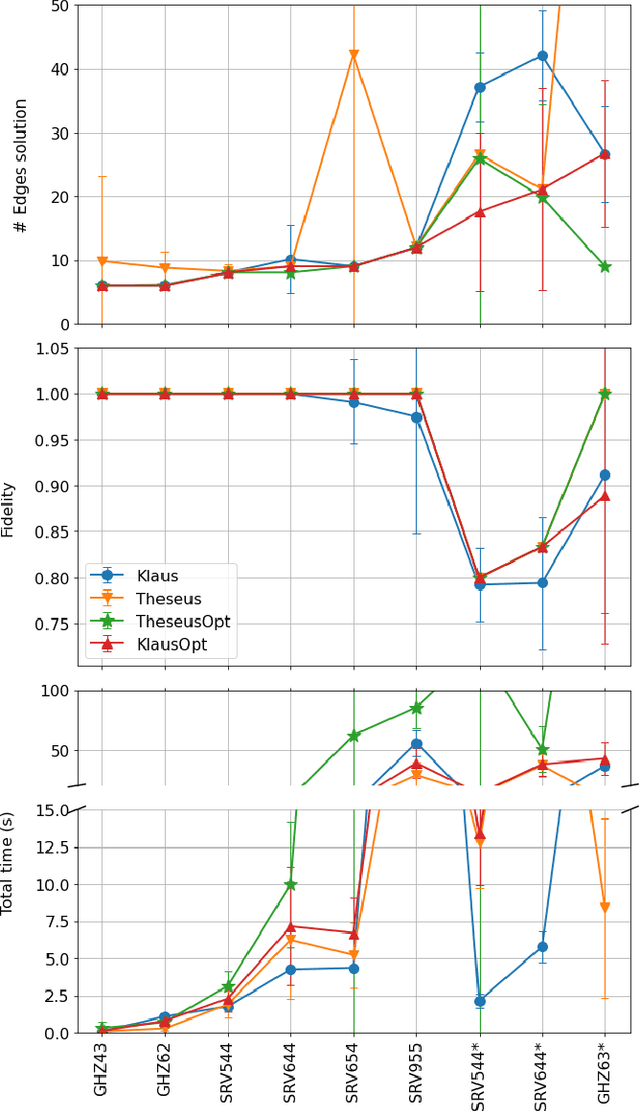
Abstract:Logic artificial intelligence (AI) is a subfield of AI where variables can take two defined arguments, True or False, and are arranged in clauses that follow the rules of formal logic. Several problems that span from physical systems to mathematical conjectures can be encoded into these clauses and be solved by checking their satisfiability (SAT). Recently, SAT solvers have become a sophisticated and powerful computational tool capable, among other things, of solving long-standing mathematical conjectures. In this work, we propose the use of logic AI for the design of optical quantum experiments. We show how to map into a SAT problem the experimental preparation of an arbitrary quantum state and propose a logic-based algorithm, called Klaus, to find an interpretable representation of the photonic setup that generates it. We compare the performance of Klaus with the state-of-the-art algorithm for this purpose based on continuous optimization. We also combine both logic and numeric strategies to find that the use of logic AI improves significantly the resolution of this problem, paving the path to develop more formal-based approaches in the context of quantum physics experiments.
Learning Interpretable Representations of Entanglement in Quantum Optics Experiments using Deep Generative Models
Sep 06, 2021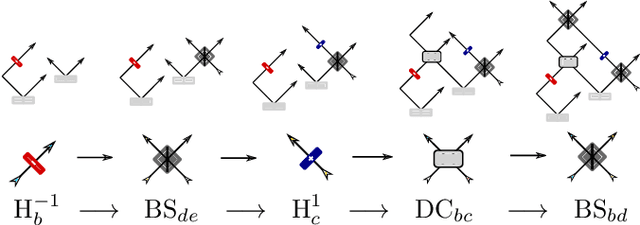
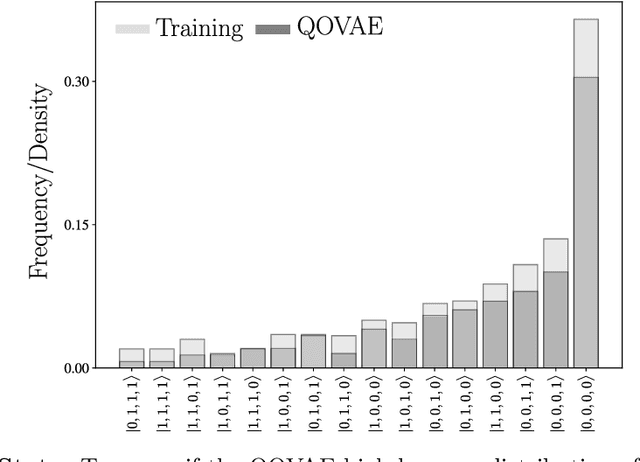
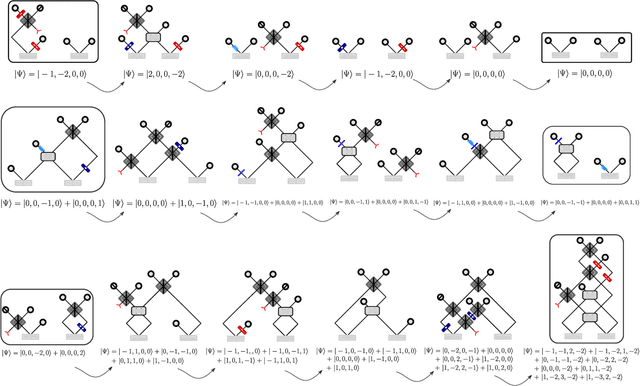
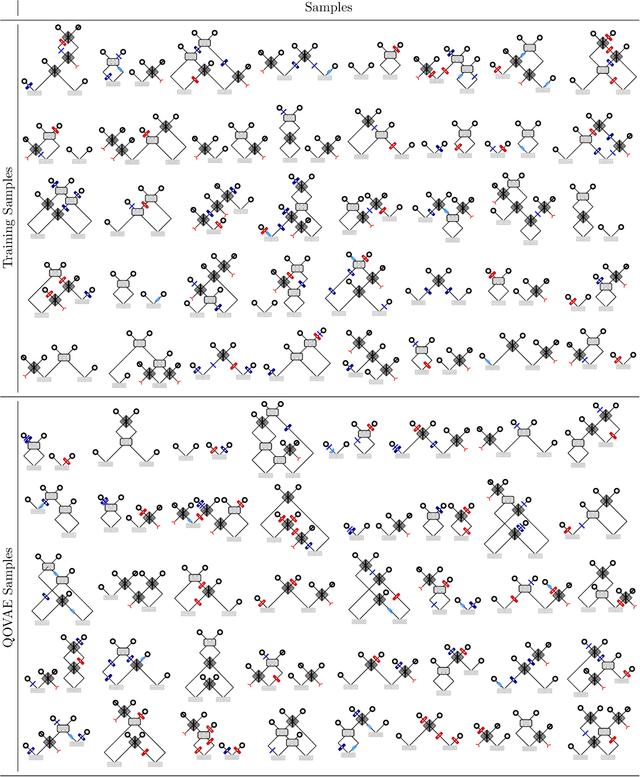
Abstract:Quantum physics experiments produce interesting phenomena such as interference or entanglement, which is a core property of numerous future quantum technologies. The complex relationship between a quantum experiment's structure and its entanglement properties is essential to fundamental research in quantum optics but is difficult to intuitively understand. We present the first deep generative model of quantum optics experiments where a variational autoencoder (QOVAE) is trained on a dataset of experimental setups. In a series of computational experiments, we investigate the learned representation of the QOVAE and its internal understanding of the quantum optics world. We demonstrate that the QOVAE learns an intrepretable representation of quantum optics experiments and the relationship between experiment structure and entanglement. We show the QOVAE is able to generate novel experiments for highly entangled quantum states with specific distributions that match its training data. Importantly, we are able to fully interpret how the QOVAE structures its latent space, finding curious patterns that we can entirely explain in terms of quantum physics. The results demonstrate how we can successfully use and understand the internal representations of deep generative models in a complex scientific domain. The QOVAE and the insights from our investigations can be immediately applied to other physical systems throughout fundamental scientific research.
Curiosity in exploring chemical space: Intrinsic rewards for deep molecular reinforcement learning
Dec 17, 2020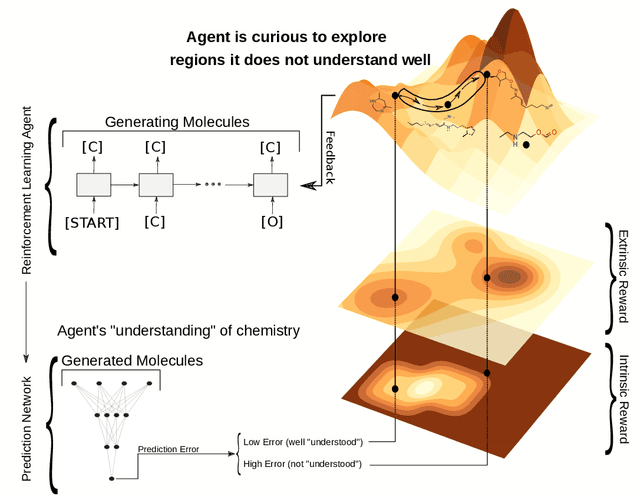
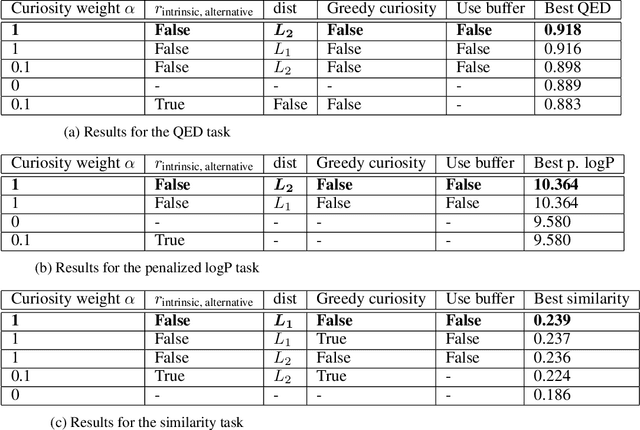
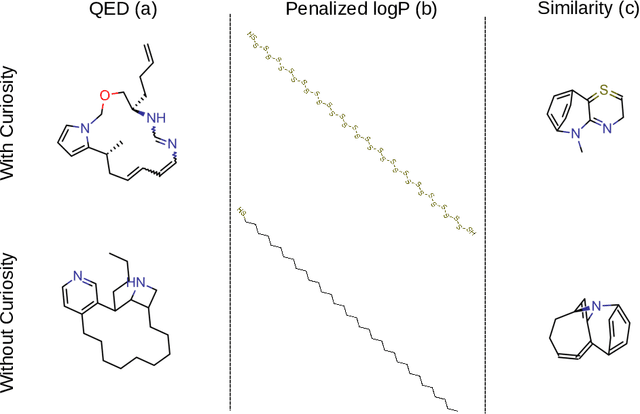
Abstract:Computer-aided design of molecules has the potential to disrupt the field of drug and material discovery. Machine learning, and deep learning, in particular, have been topics where the field has been developing at a rapid pace. Reinforcement learning is a particularly promising approach since it allows for molecular design without prior knowledge. However, the search space is vast and efficient exploration is desirable when using reinforcement learning agents. In this study, we propose an algorithm to aid efficient exploration. The algorithm is inspired by a concept known in the literature as curiosity. We show on three benchmarks that a curious agent finds better performing molecules. This indicates an exciting new research direction for reinforcement learning agents that can explore the chemical space out of their own motivation. This has the potential to eventually lead to unexpected new molecules that no human has thought about so far.
Deep Molecular Dreaming: Inverse machine learning for de-novo molecular design and interpretability with surjective representations
Dec 17, 2020
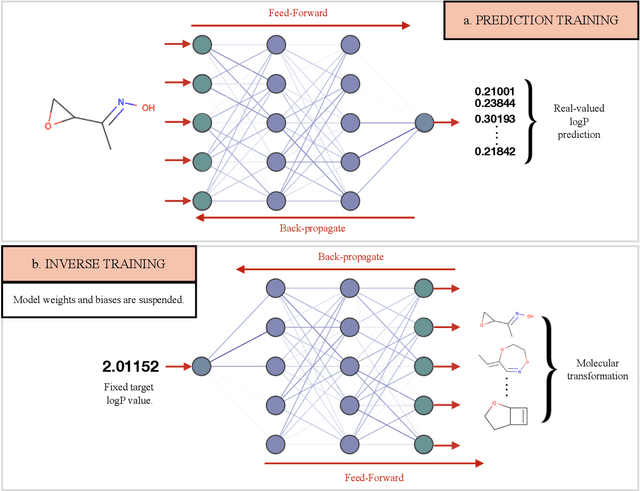
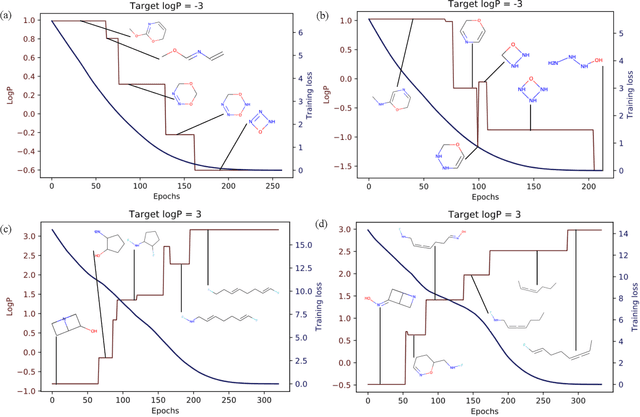
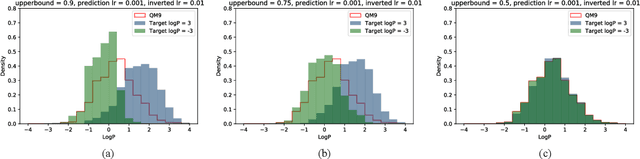
Abstract:Computer-based de-novo design of functional molecules is one of the most prominent challenges in cheminformatics today. As a result, generative and evolutionary inverse designs from the field of artificial intelligence have emerged at a rapid pace, with aims to optimize molecules for a particular chemical property. These models 'indirectly' explore the chemical space; by learning latent spaces, policies, distributions or by applying mutations on populations of molecules. However, the recent development of the SELFIES string representation of molecules, a surjective alternative to SMILES, have made possible other potential techniques. Based on SELFIES, we therefore propose PASITHEA, a direct gradient-based molecule optimization that applies inceptionism techniques from computer vision. PASITHEA exploits the use of gradients by directly reversing the learning process of a neural network, which is trained to predict real-valued chemical properties. Effectively, this forms an inverse regression model, which is capable of generating molecular variants optimized for a certain property. Although our results are preliminary, we observe a shift in distribution of a chosen property during inverse-training, a clear indication of PASITHEA's viability. A striking property of inceptionism is that we can directly probe the model's understanding of the chemical space it was trained on. We expect that extending PASITHEA to larger datasets, molecules and more complex properties will lead to advances in the design of new functional molecules as well as the interpretation and explanation of machine learning models.
Scientific intuition inspired by machine learning generated hypotheses
Oct 27, 2020
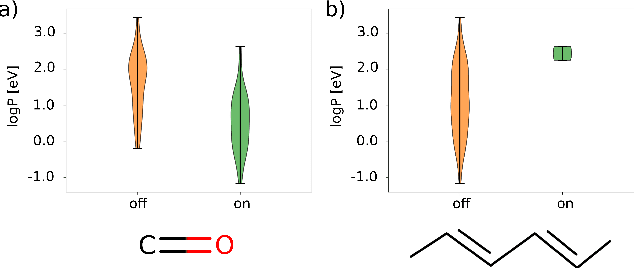
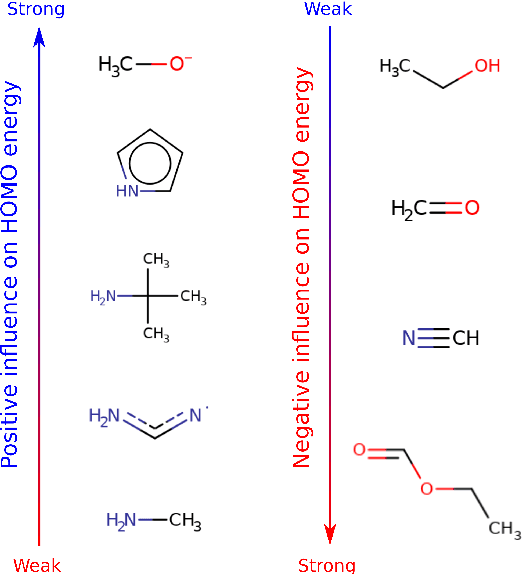
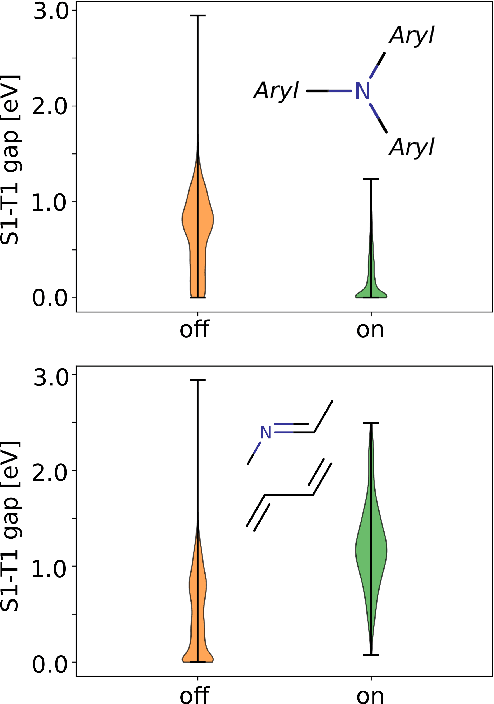
Abstract:Machine learning with application to questions in the physical sciences has become a widely used tool, successfully applied to classification, regression and optimization tasks in many areas. Research focus mostly lies in improving the accuracy of the machine learning models in numerical predictions, while scientific understanding is still almost exclusively generated by human researchers analysing numerical results and drawing conclusions. In this work, we shift the focus on the insights and the knowledge obtained by the machine learning models themselves. In particular, we study how it can be extracted and used to inspire human scientists to increase their intuitions and understanding of natural systems. We apply gradient boosting in decision trees to extract human interpretable insights from big data sets from chemistry and physics. In chemistry, we not only rediscover widely know rules of thumb but also find new interesting motifs that tell us how to control solubility and energy levels of organic molecules. At the same time, in quantum physics, we gain new understanding on experiments for quantum entanglement. The ability to go beyond numerics and to enter the realm of scientific insight and hypothesis generation opens the door to use machine learning to accelerate the discovery of conceptual understanding in some of the most challenging domains of science.
Computer-inspired Quantum Experiments
Feb 23, 2020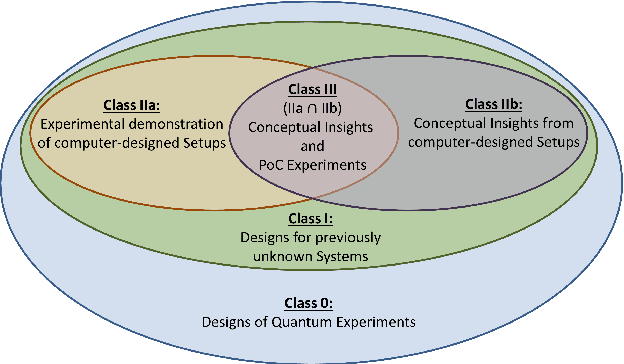
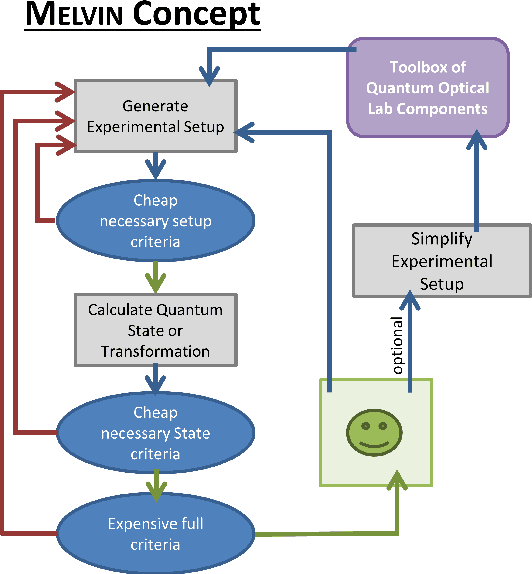
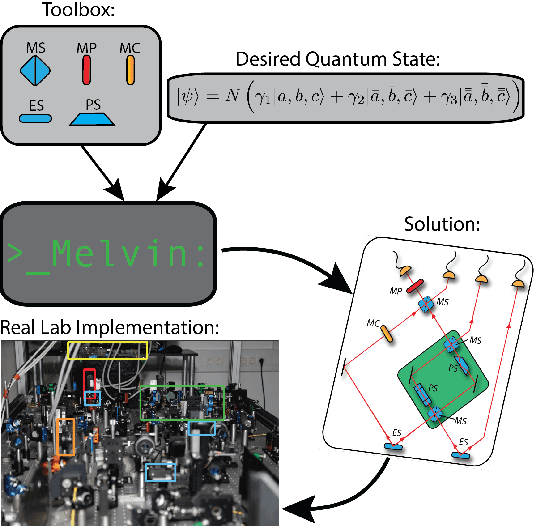
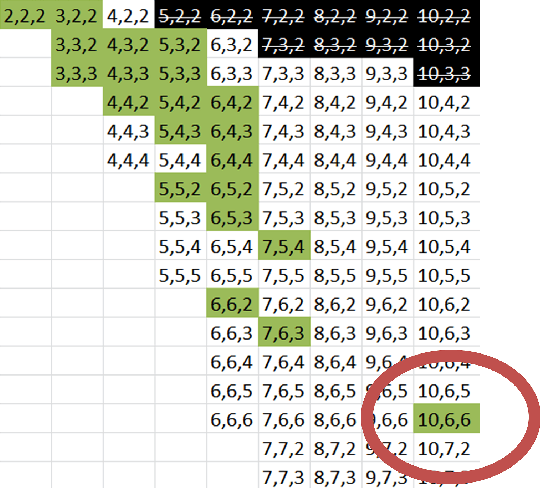
Abstract:The design of new devices and experiments in science and engineering has historically relied on the intuitions of human experts. This credo, however, has changed. In many disciplines, computer-inspired design processes, also known as inverse-design, have augmented the capability of scientists. Here we visit different fields of physics in which computer-inspired designs are applied. We will meet vastly diverse computational approaches based on topological optimization, evolutionary strategies, deep learning, reinforcement learning or automated reasoning. Then we draw our attention specifically on quantum physics. In the quest for designing new quantum experiments, we face two challenges: First, quantum phenomena are unintuitive. Second, the number of possible configurations of quantum experiments explodes combinatorially. To overcome these challenges, physicists began to use algorithms for computer-designed quantum experiments. We focus on the most mature and \textit{practical} approaches that scientists used to find new complex quantum experiments, which experimentalists subsequently have realized in the laboratories. The underlying idea is a highly-efficient topological search, which allows for scientific interpretability. In that way, some of the computer-designs have led to the discovery of new scientific concepts and ideas -- demonstrating how computer algorithm can genuinely contribute to science by providing unexpected inspirations. We discuss several extensions and alternatives based on optimization and machine learning techniques, with the potential of accelerating the discovery of practical computer-inspired experiments or concepts in the future. Finally, we discuss what we can learn from the different approaches in the fields of physics, and raise several fascinating possibilities for future research.
Quantum Optical Experiments Modeled by Long Short-Term Memory
Oct 30, 2019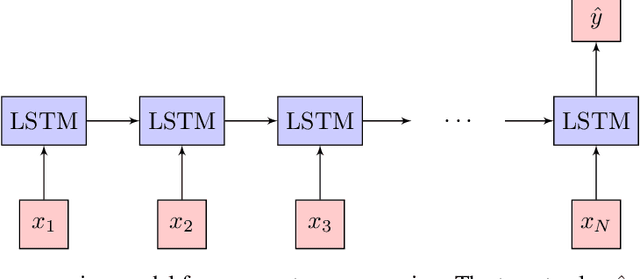
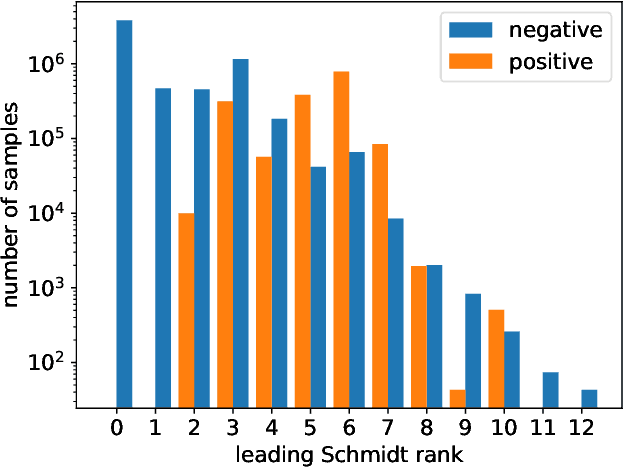
Abstract:We demonstrate how machine learning is able to model experiments in quantum physics. Quantum entanglement is a cornerstone for upcoming quantum technologies such as quantum computation and quantum cryptography. Of particular interest are complex quantum states with more than two particles and a large number of entangled quantum levels. Given such a multiparticle high-dimensional quantum state, it is usually impossible to reconstruct an experimental setup that produces it. To search for interesting experiments, one thus has to randomly create millions of setups on a computer and calculate the respective output states. In this work, we show that machine learning models can provide significant improvement over random search. We demonstrate that a long short-term memory (LSTM) neural network can successfully learn to model quantum experiments by correctly predicting output state characteristics for given setups without the necessity of computing the states themselves. This approach not only allows for faster search but is also an essential step towards automated design of multiparticle high-dimensional quantum experiments using generative machine learning models.
Augmenting Genetic Algorithms with Deep Neural Networks for Exploring the Chemical Space
Sep 30, 2019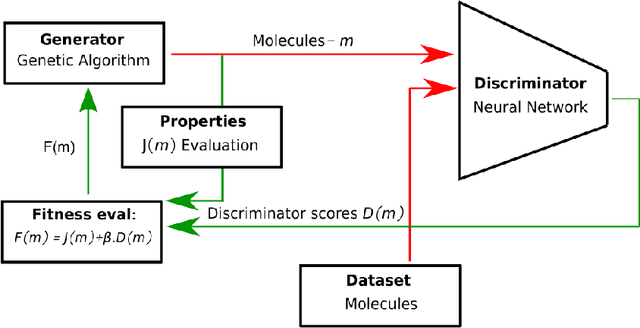
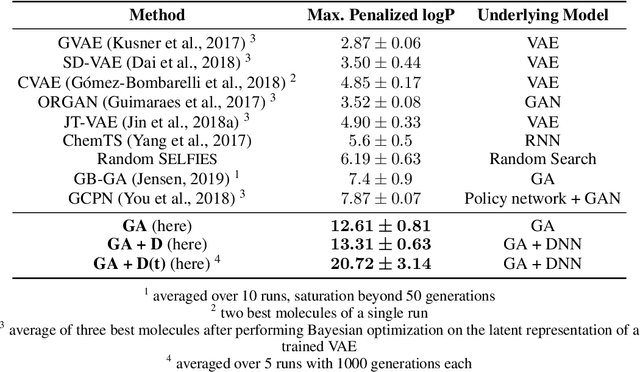
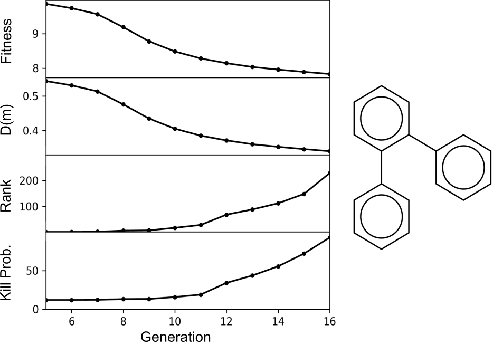
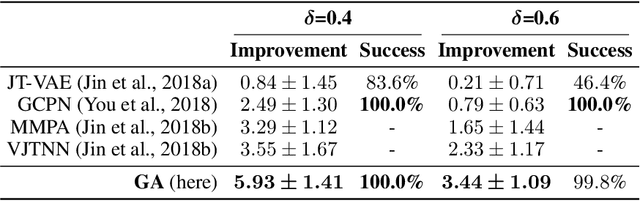
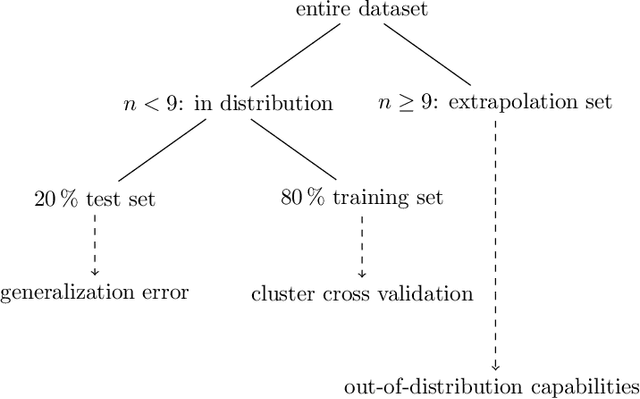
Abstract:Challenges in natural sciences can often be phrased as optimization problems. Machine learning techniques have recently been applied to solve such problems. One example in chemistry is the design of tailor-made organic materials and molecules, which requires efficient methods to explore the chemical space. We present a genetic algorithm (GA) that is enhanced with a neural network (DNN) based discriminator model to improve the diversity of generated molecules and at the same time steer the GA. We show that our algorithm outperforms other generative models in optimization tasks. We furthermore present a way to increase interpretability of genetic algorithms, which helped us to derive design principles.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge