Marc M. Van Hulle
CORTEG: Foundation Models Enable Cross-Modality Representation Transfer from Scalp to Intracranial Brain Recordings
May 11, 2026Abstract:Intracranial electrocorticography (ECoG) offers high-signal-to-noise access to cortical activity for brain-computer interfaces, yet limited per-patient data has led most prior work to rely on small, subject-specific decoders that neglect information shared across patients. We investigate whether large pretrained scalp-EEG foundation models (EEG FMs) can be adapted to ECoG, enabling cross-patient learning and competitive decoding performance while calibrating to a held-out patient in 10-30 minutes on a single GPU. We introduce CORTEG, a cross-modality transfer framework that combines a pretrained EEG FM backbone, an electrode-aware KNNSoftFourier spatial adapter, a dual-stream tokenizer for low-frequency and high-gamma activity, and a leave-one-subject-out fine-tuning strategy. We evaluate CORTEG on two challenging regression tasks: public finger trajectory regression (n=9) and private audio envelope regression (n=16). CORTEG matches or exceeds the strongest task-specific baselines on both tasks: it reaches the highest mean correlation among compared methods on the public finger benchmark (gain not statistically significant on n=9 subjects), with larger and statistically significant gains on the audio task and in low-data per-patient calibration. Feature analyses align with neurophysiology, and latent manifolds capture low-dimensional finger-movement structure. CORTEG provides systematic evidence that scalp-EEG pretraining can be repurposed for ECoG decoding, enabling data-efficient intracranial BCIs that can adapt to new patients.
An open-source implementation of a closed-loop electrocorticographic Brain-Computer Interface using Micromed, FieldTrip, and PsychoPy
Feb 10, 2026Abstract:We present an open-source implementation of a closed-loop Brain-Computer Interface (BCI) system based on electrocorticographic (ECoG) recordings. Our setup integrates FieldTrip for interfacing with a Micromed acquisition system and PsychoPy for implementing experiments. We open-source three custom Python libraries (psychopylib, pymarkerlib, and pyfieldtriplib) each covering different aspects of a closed-loop BCI interface: designing interactive experiments, sending event information, and real-time signal processing. Our modules facilitate the design and operation of a transparent BCI system, promoting customization and flexibility in BCI research, and lowering the barrier for researchers to translate advances in ECoG decoding into BCI applications.
BTTDA: Block-Term Tensor Discriminant Analysis for Brain-Computer Interfacing
Nov 06, 2025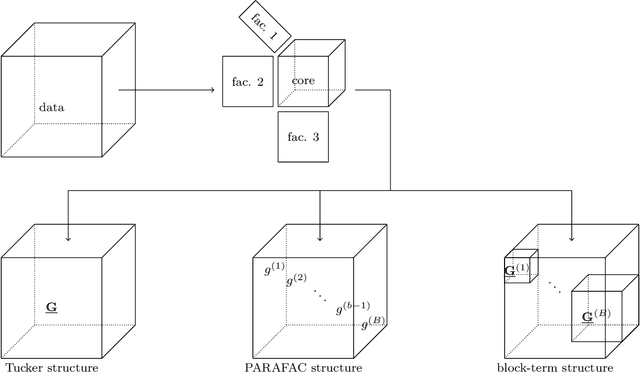
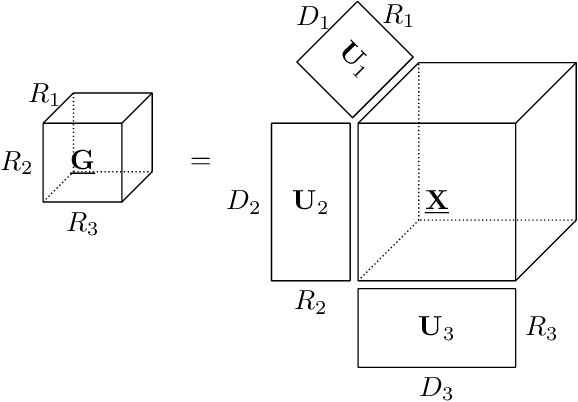
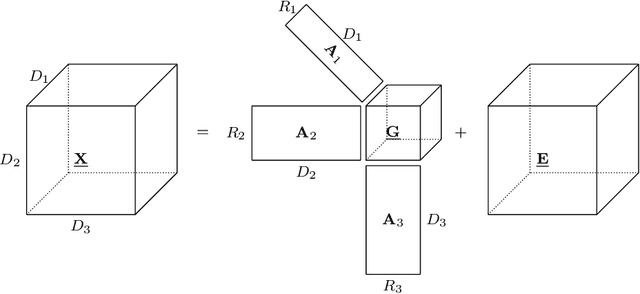
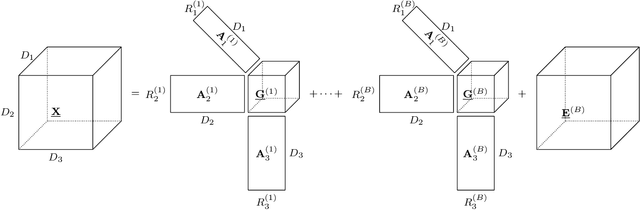
Abstract:Brain-computer interfaces (BCIs) allow direct communication between the brain and external devices, frequently using electroencephalography (EEG) to record neural activity. Dimensionality reduction and structured regularization are essential for effectively classifying task-related brain signals, including event-related potentials (ERPs) and motor imagery (MI) rhythms. Current tensor-based approaches, such as Tucker and PARAFAC decompositions, often lack the flexibility needed to fully capture the complexity of EEG data. This study introduces Block-Term Tensor Discriminant Analysis (BTTDA): a novel tensor-based and supervised feature extraction method designed to enhance classification accuracy by providing flexible multilinear dimensionality reduction. Extending Higher Order Discriminant Analysis (HODA), BTTDA uses a novel and interpretable forward model for HODA combined with a deflation scheme to iteratively extract discriminant block terms, improving feature representation for classification. BTTDA and a sum-of-rank-1-terms variant PARAFACDA were evaluated on publicly available ERP (second-order tensors) and MI (third-order tensors) EEG datasets from the MOABB benchmarking framework. Benchmarking revealed that BTTDA and PARAFACDA significantly outperform the traditional HODA method in ERP decoding, resulting in state-of-the art performance (ROC-AUC = 91.25%). For MI, decoding results of HODA, BTTDA and PARAFACDA were subpar, but BTTDA still significantly outperformed HODA (64.52% > 61.00%). The block-term structure of BTTDA enables interpretable and more efficient dimensionality reduction without compromising discriminative power. This offers a promising and adaptable approach for feature extraction in BCI and broader neuroimaging applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge