Mahir Jain
PRISM: Exploring Heterogeneous Pretrained EEG Foundation Model Transfer to Clinical Differential Diagnosis
Feb 28, 2026Abstract:EEG foundation models are typically pretrained on narrow-source clinical archives and evaluated on benchmarks from the same ecosystem, leaving unclear whether representations encode neural physiology or recording-distribution artifacts. We introduce PRISM (Population Representative Invariant Signal Model), a masked autoencoder ablated along two axes -- pretraining population and downstream adaptation -- with architecture and preprocessing fixed. We compare a narrow-source EU/US corpus (TUH + PhysioNet) against a geographically diverse pool augmented with multi-center South Asian clinical recordings across multiple EEG systems. Three findings emerge. First, narrow-source pretraining yields stronger linear probes on distribution-matched benchmarks, while diverse pretraining produces more adaptable representations under fine-tuning -- a trade-off invisible under single-protocol evaluation. Trained on three source corpora, PRISM matches or outperforms REVE (92 datasets, 60,000+ hours) on the majority of tasks, demonstrating that targeted diversity can substitute for indiscriminate scale and that dataset count is a confounding variable in model comparison. Second, on a clinically challenging and previously untested task -- distinguishing epilepsy from diagnostic mimickers via interictal EEG -- the diverse checkpoint outperforms the narrow-source checkpoint by +12.3 pp balanced accuracy, the largest gap across all evaluations. Third, systematic inconsistencies between EEG-Bench and EEG-FM-Bench reverse model rankings on identical datasets by up to 24 pp; we identify six concrete sources including split construction, checkpoint selection, segment length, and normalization, showing these factors compound non-additively.
Revisiting Bayesian Autoencoders with MCMC
Apr 13, 2021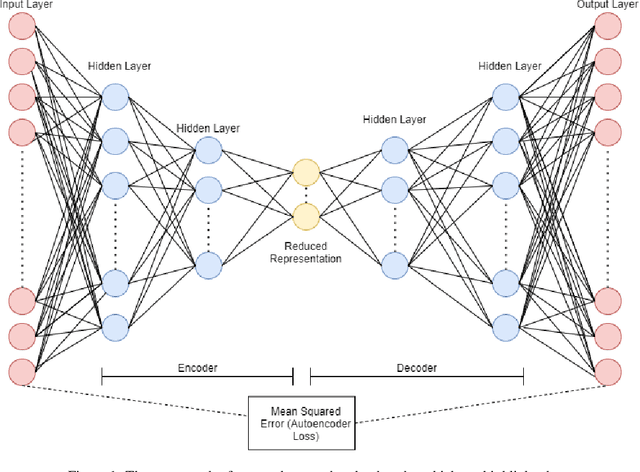

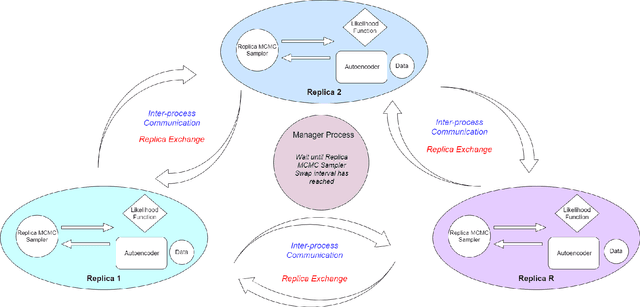

Abstract:Autoencoders gained popularity in the deep learning revolution given their ability to compress data and provide dimensionality reduction. Although prominent deep learning methods have been used to enhance autoencoders, the need to provide robust uncertainty quantification remains a challenge. This has been addressed with variational autoencoders so far. Bayesian inference via MCMC methods have faced limitations but recent advances with parallel computing and advanced proposal schemes that incorporate gradients have opened routes less travelled. In this paper, we present Bayesian autoencoders powered MCMC sampling implemented using parallel computing and Langevin gradient proposal scheme. Our proposed Bayesian autoencoder provides similar performance accuracy when compared to related methods from the literature, with the additional feature of robust uncertainty quantification in compressed datasets. This motivates further application of the Bayesian autoencoder framework for other deep learning models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge