Jeet Bandhu Lahiri
PRISM: Exploring Heterogeneous Pretrained EEG Foundation Model Transfer to Clinical Differential Diagnosis
Feb 28, 2026Abstract:EEG foundation models are typically pretrained on narrow-source clinical archives and evaluated on benchmarks from the same ecosystem, leaving unclear whether representations encode neural physiology or recording-distribution artifacts. We introduce PRISM (Population Representative Invariant Signal Model), a masked autoencoder ablated along two axes -- pretraining population and downstream adaptation -- with architecture and preprocessing fixed. We compare a narrow-source EU/US corpus (TUH + PhysioNet) against a geographically diverse pool augmented with multi-center South Asian clinical recordings across multiple EEG systems. Three findings emerge. First, narrow-source pretraining yields stronger linear probes on distribution-matched benchmarks, while diverse pretraining produces more adaptable representations under fine-tuning -- a trade-off invisible under single-protocol evaluation. Trained on three source corpora, PRISM matches or outperforms REVE (92 datasets, 60,000+ hours) on the majority of tasks, demonstrating that targeted diversity can substitute for indiscriminate scale and that dataset count is a confounding variable in model comparison. Second, on a clinically challenging and previously untested task -- distinguishing epilepsy from diagnostic mimickers via interictal EEG -- the diverse checkpoint outperforms the narrow-source checkpoint by +12.3 pp balanced accuracy, the largest gap across all evaluations. Third, systematic inconsistencies between EEG-Bench and EEG-FM-Bench reverse model rankings on identical datasets by up to 24 pp; we identify six concrete sources including split construction, checkpoint selection, segment length, and normalization, showing these factors compound non-additively.
SVM and ANN based Classification of EMG signals by using PCA and LDA
Oct 22, 2021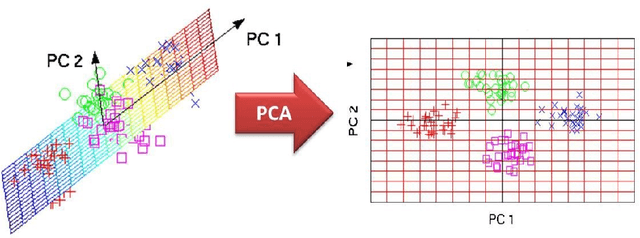
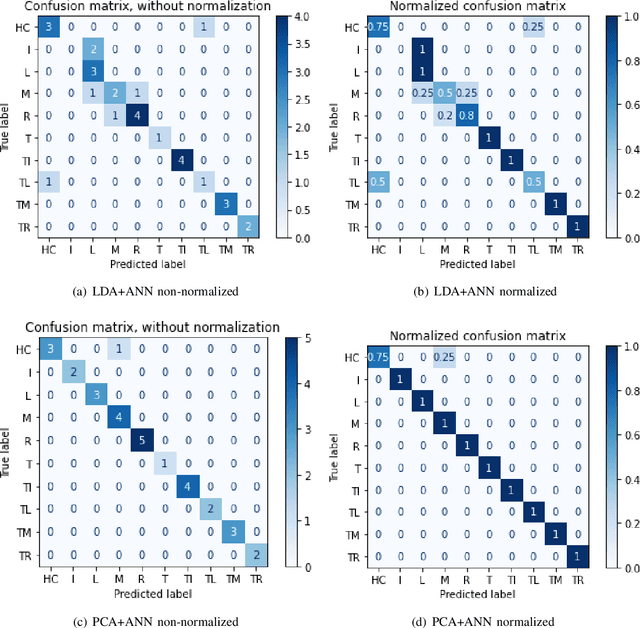
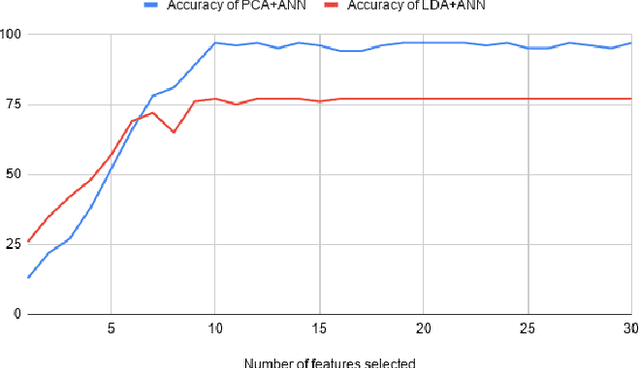
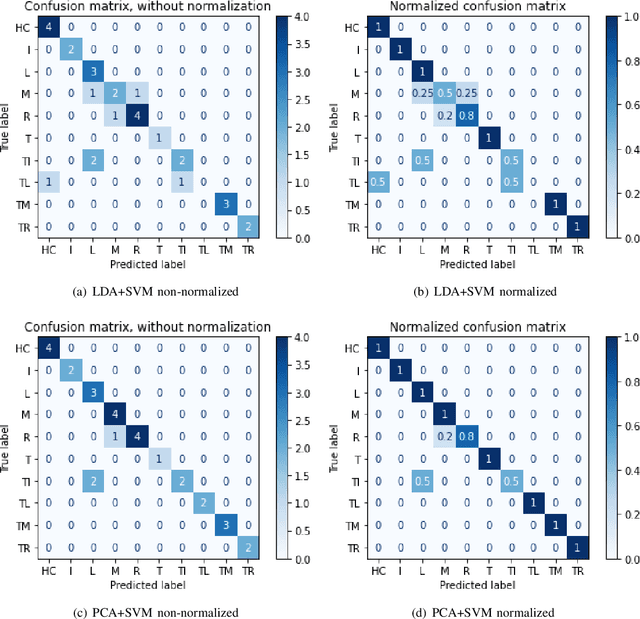
Abstract:In recent decades, biomedical signals have been used for communication in Human-Computer Interfaces (HCI) for medical applications; an instance of these signals are the myoelectric signals (MES), which are generated in the muscles of the human body as unidimensional patterns. Because of this, the methods and algorithms developed for pattern recognition in signals can be applied for their analyses once these signals have been sampled and turned into electromyographic (EMG) signals. Additionally, in recent years, many researchers have dedicated their efforts to studying prosthetic control utilizing EMG signal classification, that is, by logging a set of MES in a proper range of frequencies to classify the corresponding EMG signals. The feature classification can be carried out on the time domain or by using other domains such as the frequency domain (also known as the spectral domain), time scale, and time-frequency, amongst others. One of the main methods used for pattern recognition in myoelectric signals is the Support Vector Machines (SVM) technique whose primary function is to identify an n-dimensional hyperplane to separate a set of input feature points into different classes. This technique has the potential to recognize complex patterns and on several occasions, it has proven its worth when compared to other classifiers such as Artificial Neural Network (ANN), Linear Discriminant Analysis (LDA), and Principal Component Analysis(PCA). The key concepts underlying the SVM are (a) the hyperplane separator; (b) the kernel function; (c) the optimal separation hyperplane; and (d) a soft margin (hyperplane tolerance).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge