Maarten de Rooij
and on behalf of the UNICORN consortium
Designing UNICORN: a Unified Benchmark for Imaging in Computational Pathology, Radiology, and Natural Language
Mar 03, 2026Abstract:Medical foundation models show promise to learn broadly generalizable features from large, diverse datasets. This could be the base for reliable cross-modality generalization and rapid adaptation to new, task-specific goals, with only a few task-specific examples. Yet, evidence for this is limited by the lack of public, standardized, and reproducible evaluation frameworks, as existing public benchmarks are often fragmented across task-, organ-, or modality-specific settings, limiting assessment of cross-task generalization. We introduce UNICORN, a public benchmark designed to systematically evaluate medical foundation models under a unified protocol. To isolate representation quality, we built the benchmark on a novel two-step framework that decouples model inference from task-specific evaluation based on standardized few-shot adaptation. As a central design choice, we constructed indirectly accessible sequestered test sets derived from clinically relevant cohorts, along with standardized evaluation code and a submission interface on an open benchmarking platform. Performance is aggregated into a single UNICORN Score, a new metric that we introduce to support direct comparison of foundation models across diverse medical domains, modalities, and task types. The UNICORN test dataset includes data from more than 2,400 patients, including over 3,700 vision cases and over 2,400 clinical reports collected from 17 institutions across eight countries. The benchmark spans eight anatomical regions and four imaging modalities. Both task-specific and aggregated leaderboards enable accessible, standardized, and reproducible evaluation. By standardizing multi-task, multi-modality assessment, UNICORN establishes a foundation for reproducible benchmarking of medical foundation models. Data, baseline methods, and the evaluation platform are publicly available via unicorn.grand-challenge.org.
Deformable MRI Sequence Registration for AI-based Prostate Cancer Diagnosis
Apr 15, 2024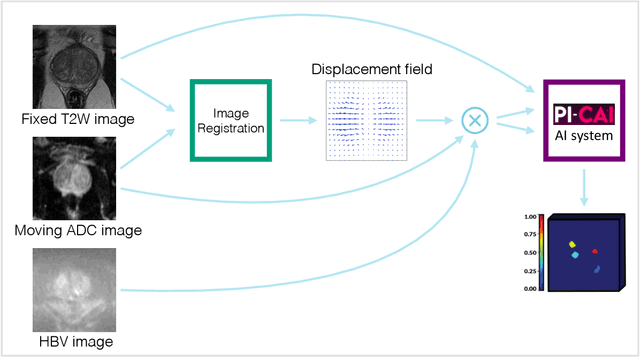
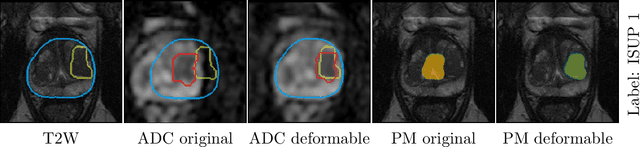
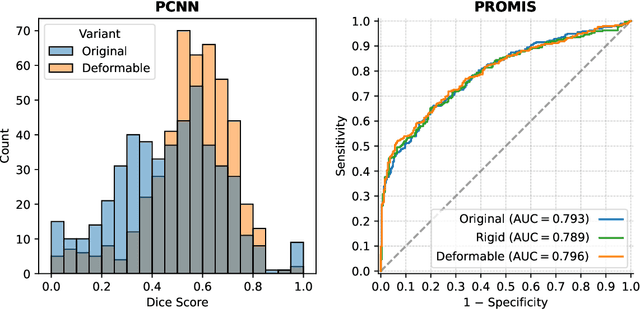
Abstract:The PI-CAI (Prostate Imaging: Cancer AI) challenge led to expert-level diagnostic algorithms for clinically significant prostate cancer detection. The algorithms receive biparametric MRI scans as input, which consist of T2-weighted and diffusion-weighted scans. These scans can be misaligned due to multiple factors in the scanning process. Image registration can alleviate this issue by predicting the deformation between the sequences. We investigate the effect of image registration on the diagnostic performance of AI-based prostate cancer diagnosis. First, the image registration algorithm, developed in MeVisLab, is analyzed using a dataset with paired lesion annotations. Second, the effect on diagnosis is evaluated by comparing case-level cancer diagnosis performance between using the original dataset, rigidly aligned diffusion-weighted scans, or deformably aligned diffusion-weighted scans. Rigid registration showed no improvement. Deformable registration demonstrated a substantial improvement in lesion overlap (+10% median Dice score) and a positive yet non-significant improvement in diagnostic performance (+0.3% AUROC, p=0.18). Our investigation shows that a substantial improvement in lesion alignment does not directly lead to a significant improvement in diagnostic performance. Qualitative analysis indicated that jointly developing image registration methods and diagnostic AI algorithms could enhance diagnostic accuracy and patient outcomes.
Report-Guided Automatic Lesion Annotation for Deep Learning-Based Prostate Cancer Detection in bpMRI
Dec 09, 2021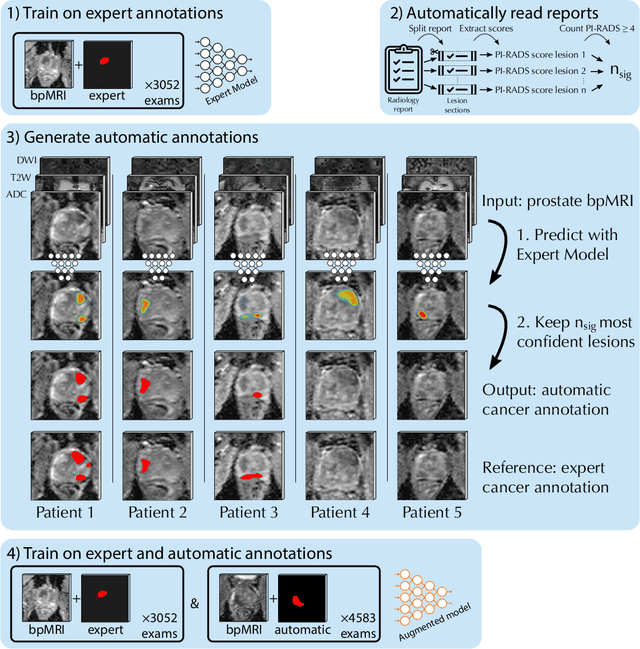

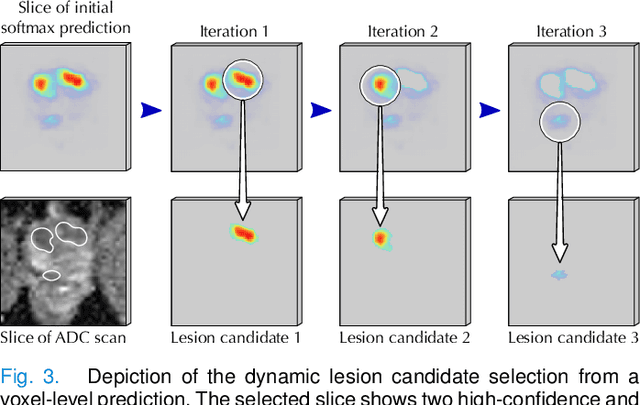
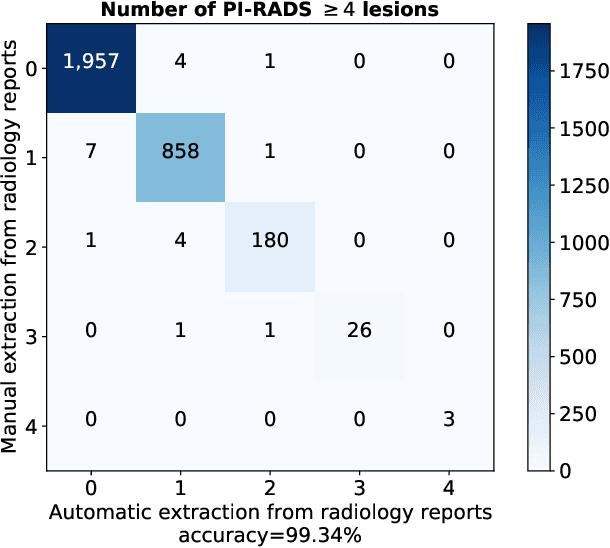
Abstract:Deep learning-based diagnostic performance increases with more annotated data, but manual annotation is a bottleneck in most fields. Experts evaluate diagnostic images during clinical routine, and write their findings in reports. Automatic annotation based on clinical reports could overcome the manual labelling bottleneck. We hypothesise that dense annotations for detection tasks can be generated using model predictions, guided by sparse information from these reports. To demonstrate efficacy, we generated clinically significant prostate cancer (csPCa) annotations, guided by the number of clinically significant findings in the radiology reports. We included 7,756 prostate MRI examinations, of which 3,050 were manually annotated and 4,706 were automatically annotated. We evaluated the automatic annotation quality on the manually annotated subset: our score extraction correctly identified the number of csPCa lesions for $99.3\%$ of the reports and our csPCa segmentation model correctly localised $83.8 \pm 1.1\%$ of the lesions. We evaluated prostate cancer detection performance on 300 exams from an external centre with histopathology-confirmed ground truth. Augmenting the training set with automatically labelled exams improved patient-based diagnostic area under the receiver operating characteristic curve from $88.1\pm 1.1\%$ to $89.8\pm 1.0\%$ ($P = 1.2 \cdot 10^{-4}$) and improved lesion-based sensitivity at one false positive per case from $79.2 \pm 2.8\%$ to $85.4 \pm 1.9\%$ ($P<10^{-4}$), with $mean \pm std.$ over 15 independent runs. This improved performance demonstrates the feasibility of our report-guided automatic annotations. Source code is made publicly available at https://github.com/DIAGNijmegen/Report-Guided-Annotation. Best csPCa detection algorithm is made available at https://grand-challenge.org/algorithms/bpmri-cspca-detection-report-guided-annotations/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge