Lukas Lange
BoschAI @ Causal News Corpus 2023: Robust Cause-Effect Span Extraction using Multi-Layer Sequence Tagging and Data Augmentation
Dec 11, 2023Abstract:Understanding causality is a core aspect of intelligence. The Event Causality Identification with Causal News Corpus Shared Task addresses two aspects of this challenge: Subtask 1 aims at detecting causal relationships in texts, and Subtask 2 requires identifying signal words and the spans that refer to the cause or effect, respectively. Our system, which is based on pre-trained transformers, stacked sequence tagging, and synthetic data augmentation, ranks third in Subtask 1 and wins Subtask 2 with an F1 score of 72.8, corresponding to a margin of 13 pp. to the second-best system.
DelucionQA: Detecting Hallucinations in Domain-specific Question Answering
Dec 08, 2023



Abstract:Hallucination is a well-known phenomenon in text generated by large language models (LLMs). The existence of hallucinatory responses is found in almost all application scenarios e.g., summarization, question-answering (QA) etc. For applications requiring high reliability (e.g., customer-facing assistants), the potential existence of hallucination in LLM-generated text is a critical problem. The amount of hallucination can be reduced by leveraging information retrieval to provide relevant background information to the LLM. However, LLMs can still generate hallucinatory content for various reasons (e.g., prioritizing its parametric knowledge over the context, failure to capture the relevant information from the context, etc.). Detecting hallucinations through automated methods is thus paramount. To facilitate research in this direction, we introduce a sophisticated dataset, DelucionQA, that captures hallucinations made by retrieval-augmented LLMs for a domain-specific QA task. Furthermore, we propose a set of hallucination detection methods to serve as baselines for future works from the research community. Analysis and case study are also provided to share valuable insights on hallucination phenomena in the target scenario.
GradSim: Gradient-Based Language Grouping for Effective Multilingual Training
Oct 23, 2023Abstract:Most languages of the world pose low-resource challenges to natural language processing models. With multilingual training, knowledge can be shared among languages. However, not all languages positively influence each other and it is an open research question how to select the most suitable set of languages for multilingual training and avoid negative interference among languages whose characteristics or data distributions are not compatible. In this paper, we propose GradSim, a language grouping method based on gradient similarity. Our experiments on three diverse multilingual benchmark datasets show that it leads to the largest performance gains compared to other similarity measures and it is better correlated with cross-lingual model performance. As a result, we set the new state of the art on AfriSenti, a benchmark dataset for sentiment analysis on low-resource African languages. In our extensive analysis, we further reveal that besides linguistic features, the topics of the datasets play an important role for language grouping and that lower layers of transformer models encode language-specific features while higher layers capture task-specific information.
TADA: Efficient Task-Agnostic Domain Adaptation for Transformers
May 22, 2023



Abstract:Intermediate training of pre-trained transformer-based language models on domain-specific data leads to substantial gains for downstream tasks. To increase efficiency and prevent catastrophic forgetting alleviated from full domain-adaptive pre-training, approaches such as adapters have been developed. However, these require additional parameters for each layer, and are criticized for their limited expressiveness. In this work, we introduce TADA, a novel task-agnostic domain adaptation method which is modular, parameter-efficient, and thus, data-efficient. Within TADA, we retrain the embeddings to learn domain-aware input representations and tokenizers for the transformer encoder, while freezing all other parameters of the model. Then, task-specific fine-tuning is performed. We further conduct experiments with meta-embeddings and newly introduced meta-tokenizers, resulting in one model per task in multi-domain use cases. Our broad evaluation in 4 downstream tasks for 14 domains across single- and multi-domain setups and high- and low-resource scenarios reveals that TADA is an effective and efficient alternative to full domain-adaptive pre-training and adapters for domain adaptation, while not introducing additional parameters or complex training steps.
NLNDE at SemEval-2023 Task 12: Adaptive Pretraining and Source Language Selection for Low-Resource Multilingual Sentiment Analysis
Apr 28, 2023



Abstract:This paper describes our system developed for the SemEval-2023 Task 12 "Sentiment Analysis for Low-resource African Languages using Twitter Dataset". Sentiment analysis is one of the most widely studied applications in natural language processing. However, most prior work still focuses on a small number of high-resource languages. Building reliable sentiment analysis systems for low-resource languages remains challenging, due to the limited training data in this task. In this work, we propose to leverage language-adaptive and task-adaptive pretraining on African texts and study transfer learning with source language selection on top of an African language-centric pretrained language model. Our key findings are: (1) Adapting the pretrained model to the target language and task using a small yet relevant corpus improves performance remarkably by more than 10 F1 score points. (2) Selecting source languages with positive transfer gains during training can avoid harmful interference from dissimilar languages, leading to better results in multilingual and cross-lingual settings. In the shared task, our system wins 8 out of 15 tracks and, in particular, performs best in the multilingual evaluation.
SwitchPrompt: Learning Domain-Specific Gated Soft Prompts for Classification in Low-Resource Domains
Feb 14, 2023Abstract:Prompting pre-trained language models leads to promising results across natural language processing tasks but is less effective when applied in low-resource domains, due to the domain gap between the pre-training data and the downstream task. In this work, we bridge this gap with a novel and lightweight prompting methodology called SwitchPrompt for the adaptation of language models trained on datasets from the general domain to diverse low-resource domains. Using domain-specific keywords with a trainable gated prompt, SwitchPrompt offers domain-oriented prompting, that is, effective guidance on the target domains for general-domain language models. Our few-shot experiments on three text classification benchmarks demonstrate the efficacy of the general-domain pre-trained language models when used with SwitchPrompt. They often even outperform their domain-specific counterparts trained with baseline state-of-the-art prompting methods by up to 10.7% performance increase in accuracy. This result indicates that SwitchPrompt effectively reduces the need for domain-specific language model pre-training.
Multilingual Normalization of Temporal Expressions with Masked Language Models
May 20, 2022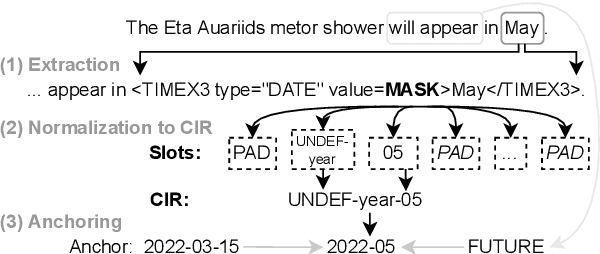
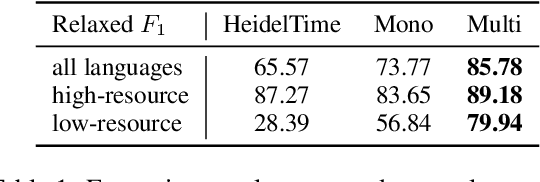
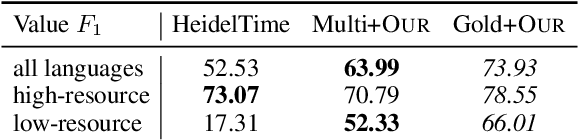
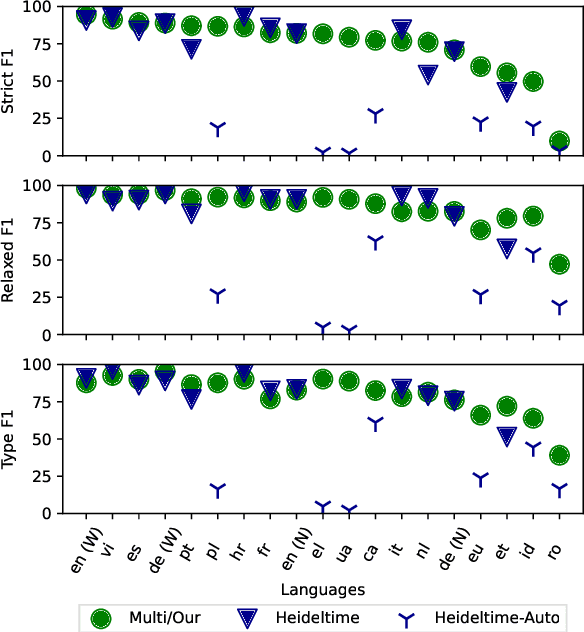
Abstract:The detection and normalization of temporal expressions is an important task and a preprocessing step for many applications. However, prior work on normalization is rule-based, which severely limits the applicability in real-world multilingual settings, due to the costly creation of new rules. We propose a novel neural method for normalizing temporal expressions based on masked language modeling. Our multilingual method outperforms prior rule-based systems in many languages, and in particular, for low-resource languages with performance improvements of up to 35 F1 on average compared to the state of the art.
CLIN-X: pre-trained language models and a study on cross-task transfer for concept extraction in the clinical domain
Dec 17, 2021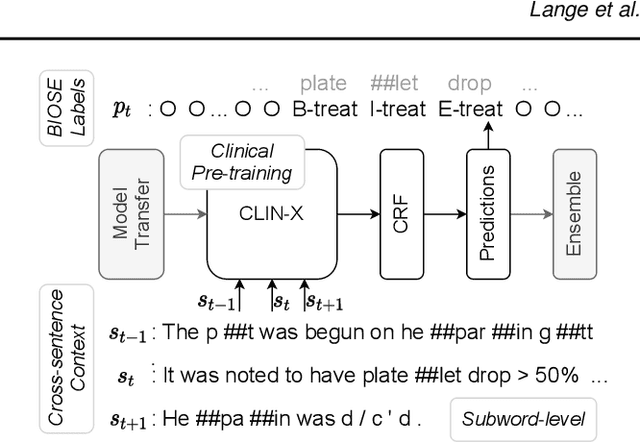
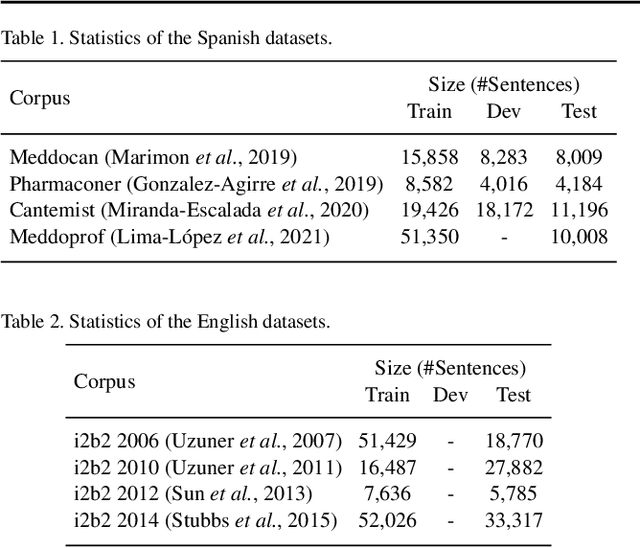
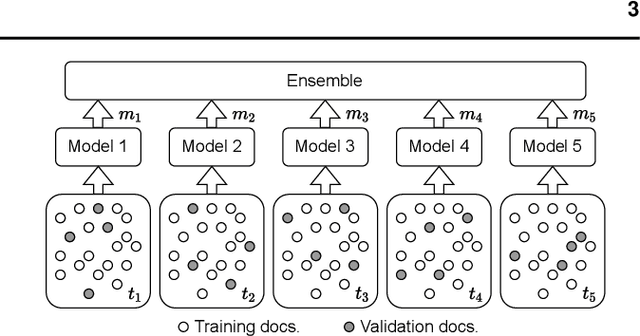
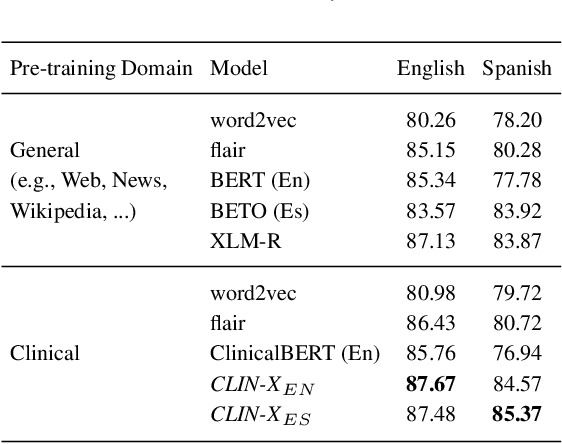
Abstract:The field of natural language processing (NLP) has recently seen a large change towards using pre-trained language models for solving almost any task. Despite showing great improvements in benchmark datasets for various tasks, these models often perform sub-optimal in non-standard domains like the clinical domain where a large gap between pre-training documents and target documents is observed. In this paper, we aim at closing this gap with domain-specific training of the language model and we investigate its effect on a diverse set of downstream tasks and settings. We introduce the pre-trained CLIN-X (Clinical XLM-R) language models and show how CLIN-X outperforms other pre-trained transformer models by a large margin for ten clinical concept extraction tasks from two languages. In addition, we demonstrate how the transformer model can be further improved with our proposed task- and language-agnostic model architecture based on ensembles over random splits and cross-sentence context. Our studies in low-resource and transfer settings reveal stable model performance despite a lack of annotated data with improvements of up to 47 F1 points when only 250 labeled sentences are available. Our results highlight the importance of specialized language models as CLIN-X for concept extraction in non-standard domains, but also show that our task-agnostic model architecture is robust across the tested tasks and languages so that domain- or task-specific adaptations are not required.
Boosting Transformers for Job Expression Extraction and Classification in a Low-Resource Setting
Sep 17, 2021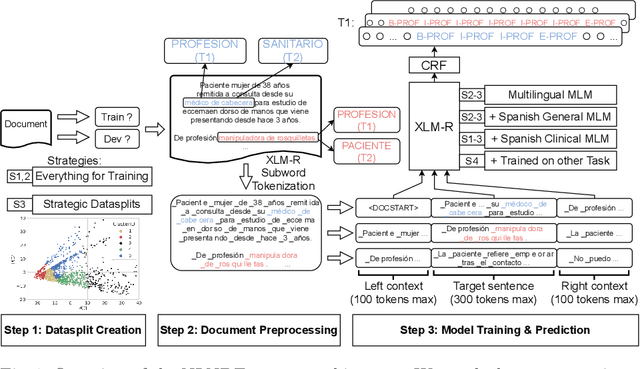
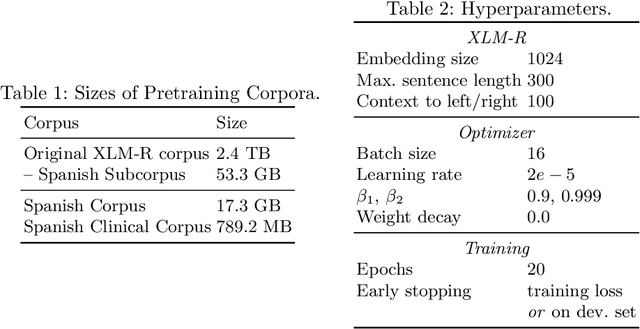
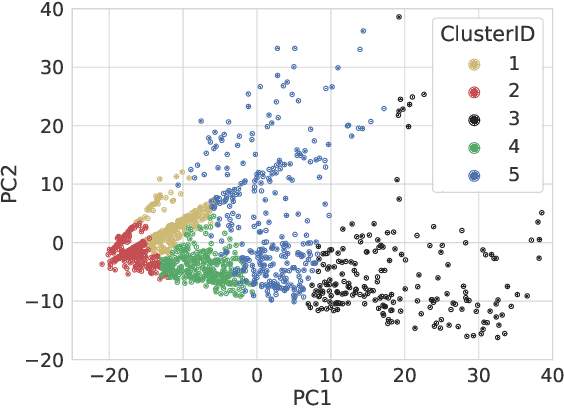
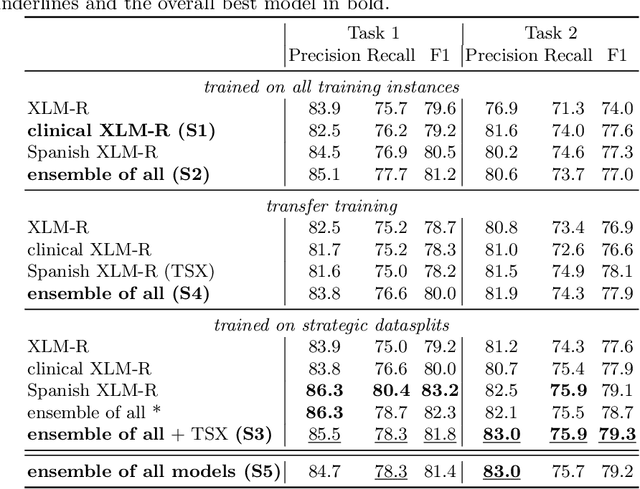
Abstract:In this paper, we explore possible improvements of transformer models in a low-resource setting. In particular, we present our approaches to tackle the first two of three subtasks of the MEDDOPROF competition, i.e., the extraction and classification of job expressions in Spanish clinical texts. As neither language nor domain experts, we experiment with the multilingual XLM-R transformer model and tackle these low-resource information extraction tasks as sequence-labeling problems. We explore domain- and language-adaptive pretraining, transfer learning and strategic datasplits to boost the transformer model. Our results show strong improvements using these methods by up to 5.3 F1 points compared to a fine-tuned XLM-R model. Our best models achieve 83.2 and 79.3 F1 for the first two tasks, respectively.
To Share or not to Share: Predicting Sets of Sources for Model Transfer Learning
Apr 16, 2021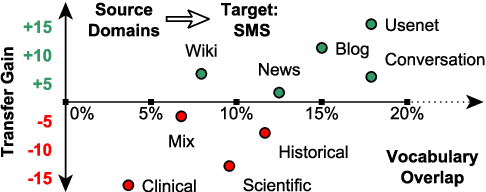
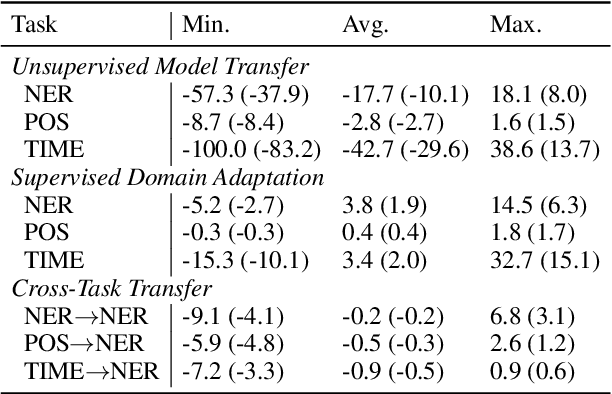
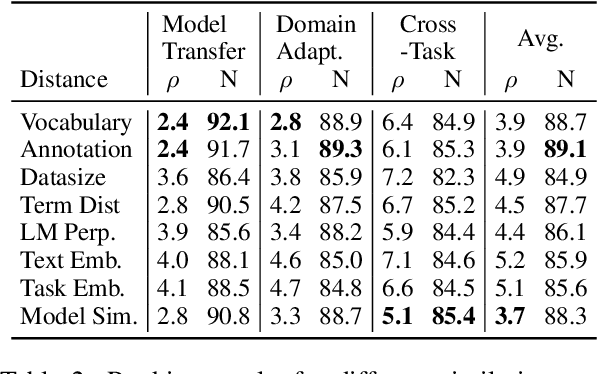
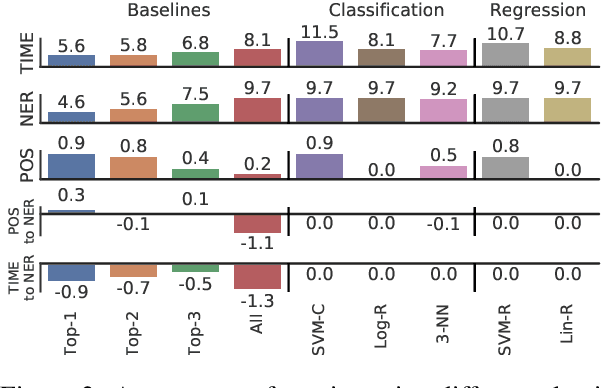
Abstract:In low-resource settings, model transfer can help to overcome a lack of labeled data for many tasks and domains. However, predicting useful transfer sources is a challenging problem, as even the most similar sources might lead to unexpected negative transfer results. Thus, ranking methods based on task and text similarity may not be sufficient to identify promising sources. To tackle this problem, we propose a method to automatically determine which and how many sources should be exploited. For this, we study the effects of model transfer on sequence labeling across various domains and tasks and show that our methods based on model similarity and support vector machines are able to predict promising sources, resulting in performance increases of up to 24 F1 points.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge