Karsten Borgwardt
The magnitude vector of images
Oct 28, 2021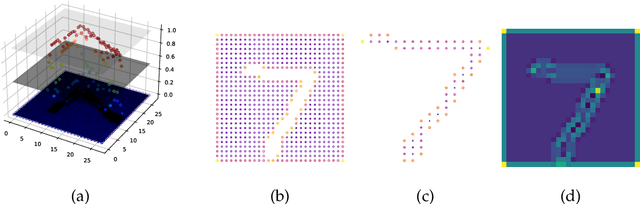

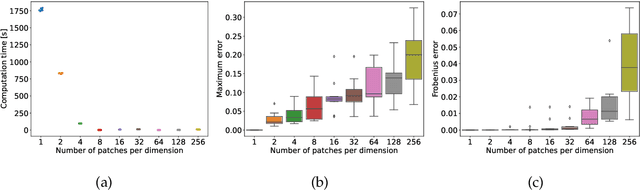
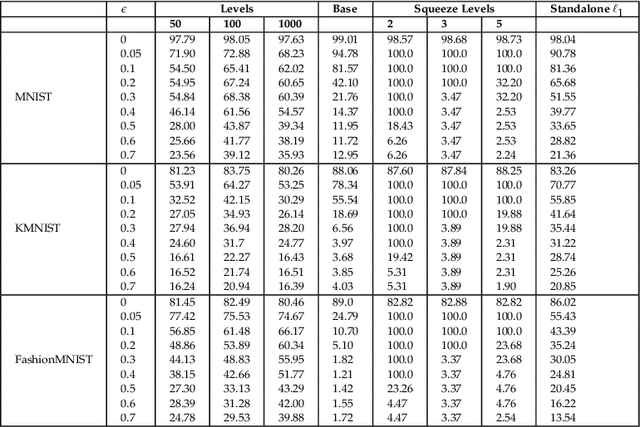
Abstract:The magnitude of a finite metric space is a recently-introduced invariant quantity. Despite beneficial theoretical and practical properties, such as a general utility for outlier detection, and a close connection to Laplace radial basis kernels, magnitude has received little attention by the machine learning community so far. In this work, we investigate the properties of magnitude on individual images, with each image forming its own metric space. We show that the known properties of outlier detection translate to edge detection in images and we give supporting theoretical justifications. In addition, we provide a proof of concept of its utility by using a novel magnitude layer to defend against adversarial attacks. Since naive magnitude calculations may be computationally prohibitive, we introduce an algorithm that leverages the regular structure of images to dramatically reduce the computational cost.
Predicting sepsis in multi-site, multi-national intensive care cohorts using deep learning
Jul 12, 2021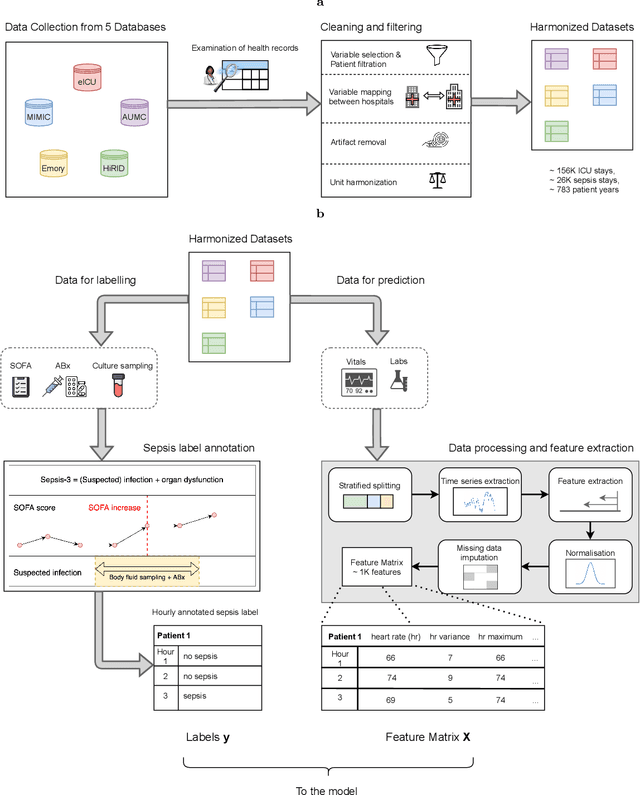
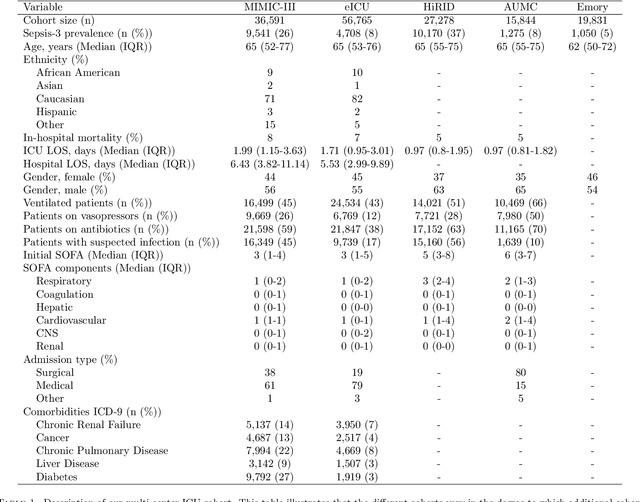
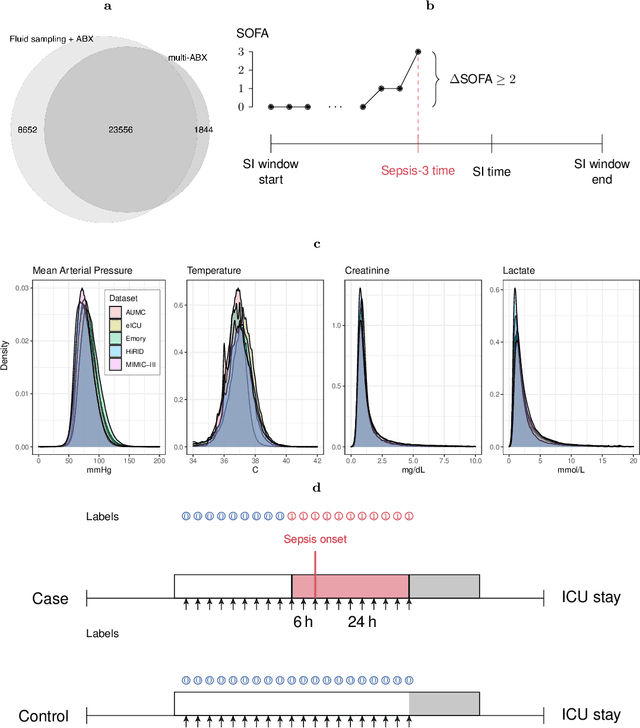
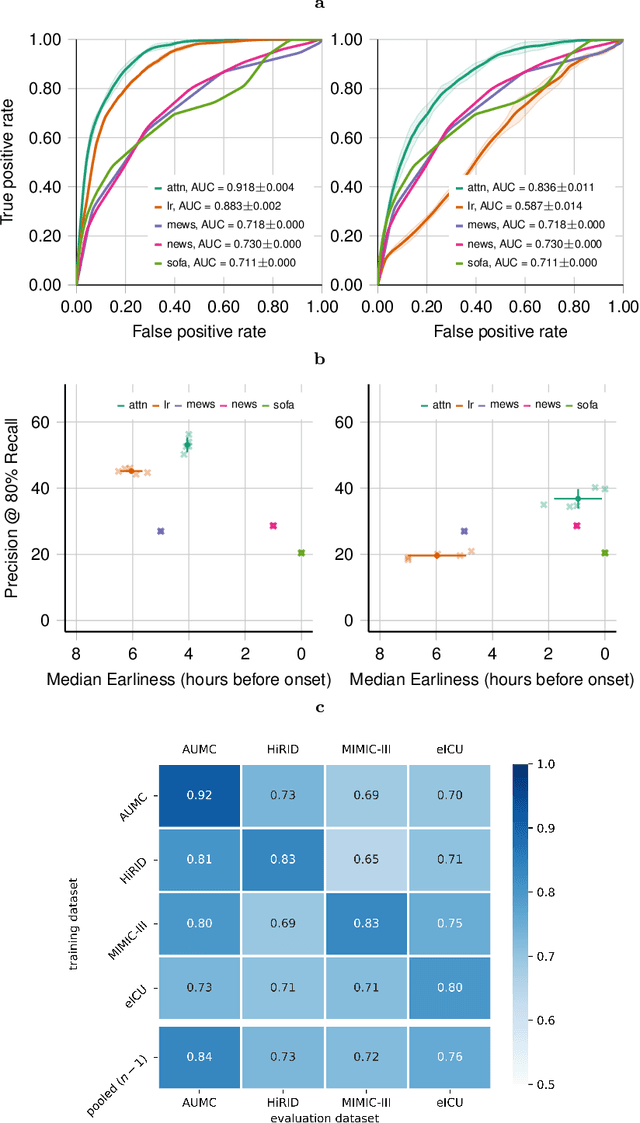
Abstract:Despite decades of clinical research, sepsis remains a global public health crisis with high mortality, and morbidity. Currently, when sepsis is detected and the underlying pathogen is identified, organ damage may have already progressed to irreversible stages. Effective sepsis management is therefore highly time-sensitive. By systematically analysing trends in the plethora of clinical data available in the intensive care unit (ICU), an early prediction of sepsis could lead to earlier pathogen identification, resistance testing, and effective antibiotic and supportive treatment, and thereby become a life-saving measure. Here, we developed and validated a machine learning (ML) system for the prediction of sepsis in the ICU. Our analysis represents the largest multi-national, multi-centre in-ICU study for sepsis prediction using ML to date. Our dataset contains $156,309$ unique ICU admissions, which represent a refined and harmonised subset of five large ICU databases originating from three countries. Using the international consensus definition Sepsis-3, we derived hourly-resolved sepsis label annotations, amounting to $26,734$ ($17.1\%$) septic stays. We compared our approach, a deep self-attention model, to several clinical baselines as well as ML baselines and performed an extensive internal and external validation within and across databases. On average, our model was able to predict sepsis with an AUROC of $0.847 \pm 0.050$ (internal out-of sample validation) and $0.761 \pm 0.052$ (external validation). For a harmonised prevalence of $17\%$, at $80\%$ recall our model detects septic patients with $39\%$ precision 3.7 hours in advance.
Evaluation Metrics for Graph Generative Models: Problems, Pitfalls, and Practical Solutions
Jun 02, 2021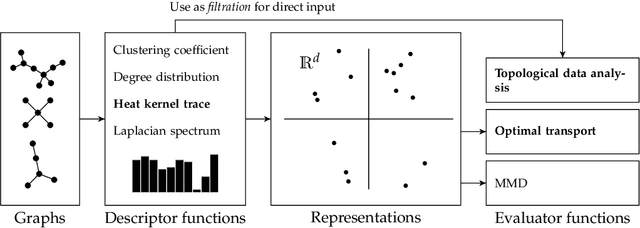
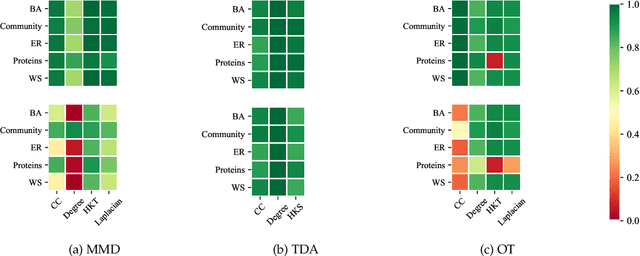
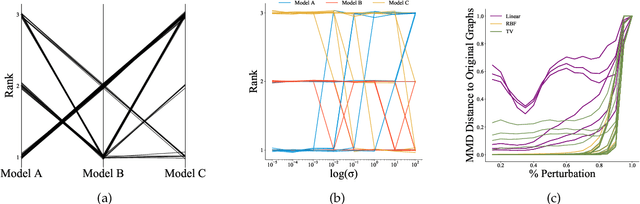
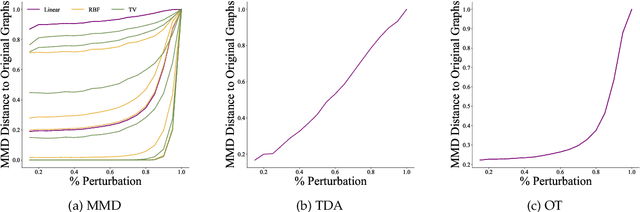

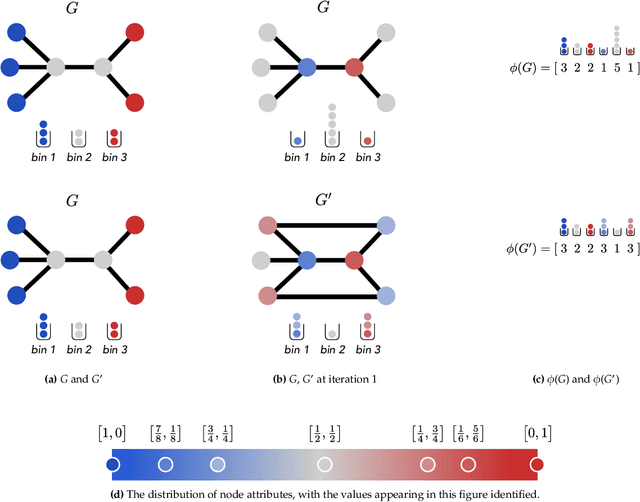
Abstract:Graph generative models are a highly active branch of machine learning. Given the steady development of new models of ever-increasing complexity, it is necessary to provide a principled way to evaluate and compare them. In this paper, we enumerate the desirable criteria for comparison metrics, discuss the development of such metrics, and provide a comparison of their respective expressive power. We perform a systematic evaluation of the main metrics in use today, highlighting some of the challenges and pitfalls researchers inadvertently can run into. We then describe a collection of suitable metrics, give recommendations as to their practical suitability, and analyse their behaviour on synthetically generated perturbed graphs as well as on recently proposed graph generative models.
Topological Graph Neural Networks
Feb 15, 2021
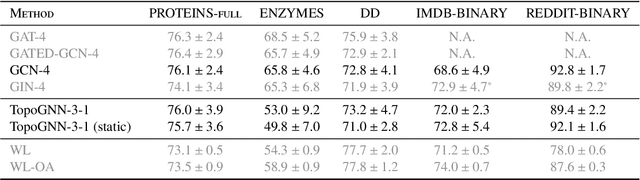
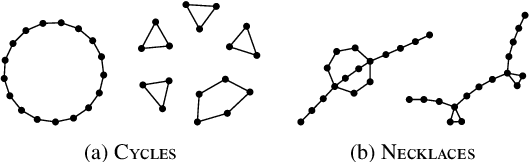
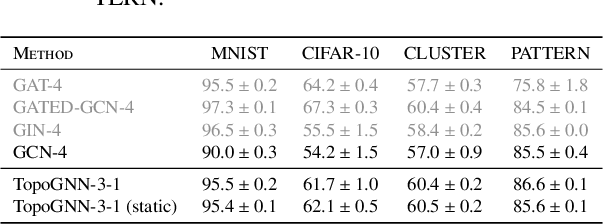
Abstract:Graph neural networks (GNNs) are a powerful architecture for tackling graph learning tasks, yet have been shown to be oblivious to eminent substructures, such as cycles. We present TOGL, a novel layer that incorporates global topological information of a graph using persistent homology. TOGL can be easily integrated into any type of GNN and is strictly more expressive in terms of the Weisfeiler--Lehman test of isomorphism. Augmenting GNNs with our layer leads to beneficial predictive performance, both on synthetic data sets, which can be trivially classified by humans but not by ordinary GNNs, and on real-world data.
Graph Kernels: State-of-the-Art and Future Challenges
Nov 10, 2020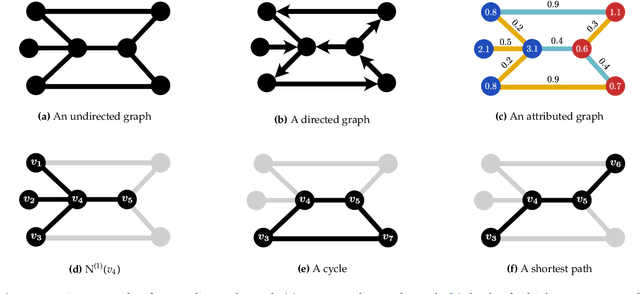
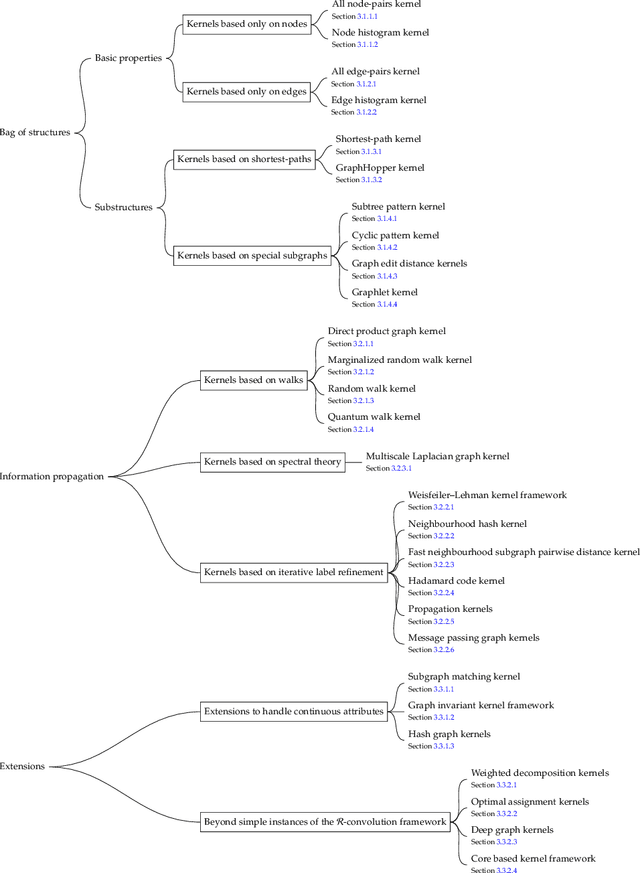
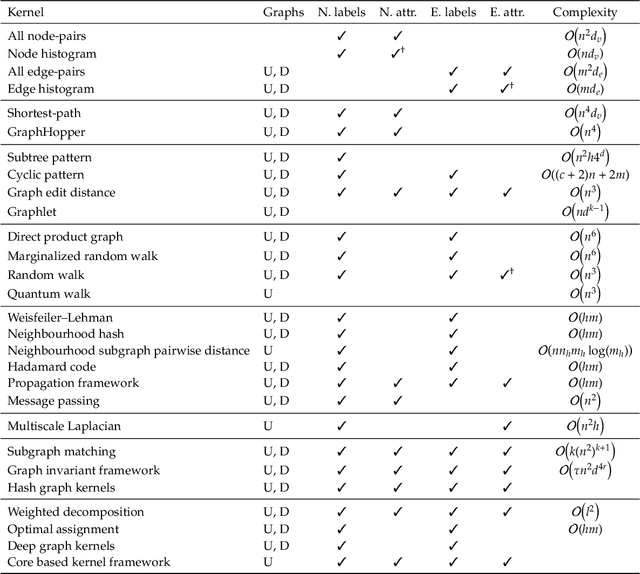
Abstract:Graph-structured data are an integral part of many application domains, including chemoinformatics, computational biology, neuroimaging, and social network analysis. Over the last two decades, numerous graph kernels, i.e. kernel functions between graphs, have been proposed to solve the problem of assessing the similarity between graphs, thereby making it possible to perform predictions in both classification and regression settings. This manuscript provides a review of existing graph kernels, their applications, software plus data resources, and an empirical comparison of state-of-the-art graph kernels.
Accelerating COVID-19 Differential Diagnosis with Explainable Ultrasound Image Analysis
Sep 13, 2020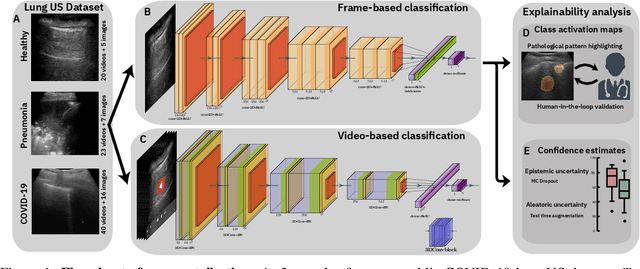
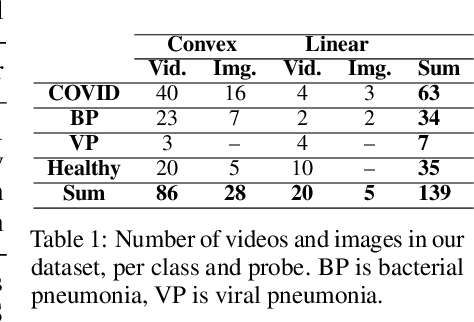

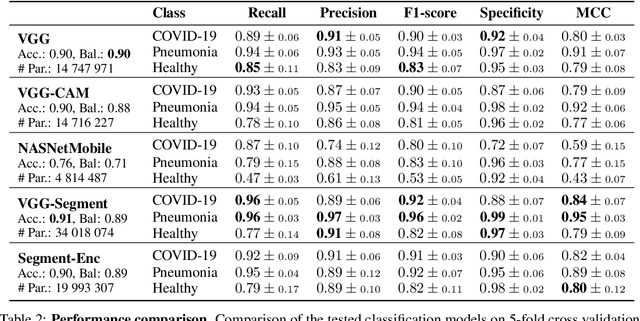
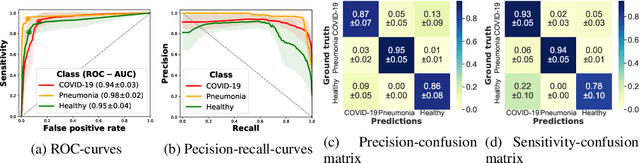
Abstract:Controlling the COVID-19 pandemic largely hinges upon the existence of fast, safe, and highly-available diagnostic tools. Ultrasound, in contrast to CT or X-Ray, has many practical advantages and can serve as a globally-applicable first-line examination technique. We provide the largest publicly available lung ultrasound (US) dataset for COVID-19 consisting of 106 videos from three classes (COVID-19, bacterial pneumonia, and healthy controls); curated and approved by medical experts. On this dataset, we perform an in-depth study of the value of deep learning methods for differential diagnosis of COVID-19. We propose a frame-based convolutional neural network that correctly classifies COVID-19 US videos with a sensitivity of 0.98+-0.04 and a specificity of 0.91+-08 (frame-based sensitivity 0.93+-0.05, specificity 0.87+-0.07). We further employ class activation maps for the spatio-temporal localization of pulmonary biomarkers, which we subsequently validate for human-in-the-loop scenarios in a blindfolded study with medical experts. Aiming for scalability and robustness, we perform ablation studies comparing mobile-friendly, frame- and video-based architectures and show reliability of the best model by aleatoric and epistemic uncertainty estimates. We hope to pave the road for a community effort toward an accessible, efficient and interpretable screening method and we have started to work on a clinical validation of the proposed method. Data and code are publicly available.
Uncovering the Topology of Time-Varying fMRI Data using Cubical Persistence
Jun 14, 2020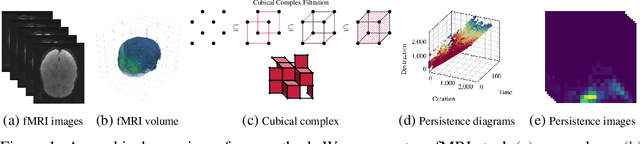
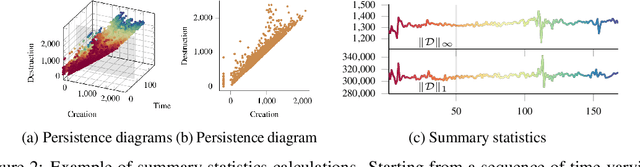
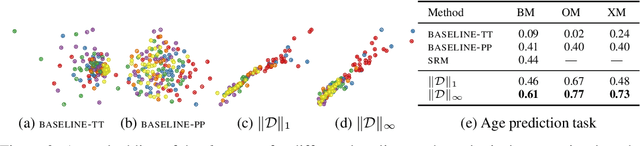
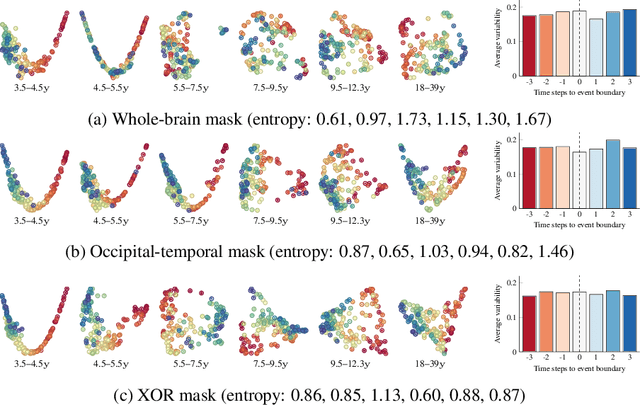
Abstract:Functional magnetic resonance imaging (fMRI) is a crucial technology for gaining insights into cognitive processes in humans. Data amassed from fMRI measurements result in volumetric data sets that vary over time. However, analysing such data presents a challenge due to the large degree of noise and person-to-person variation in how information is represented in the brain. To address this challenge, we present a novel topological approach that encodes each time point in an fMRI data set as a persistence diagram of topological features, i.e. high-dimensional voids present in the data. This representation naturally does not rely on voxel-by-voxel correspondence and is robust towards noise. We show that these time-varying persistence diagrams can be clustered to find meaningful groupings between participants, and that they are also useful in studying within-subject brain state trajectories of subjects performing a particular task. Here, we apply both clustering and trajectory analysis techniques to a group of participants watching the movie 'Partly Cloudy'. We observe significant differences in both brain state trajectories and overall topological activity between adults and children watching the same movie.
Path Imputation Strategies for Signature Models of Irregular Time Series
Jun 06, 2020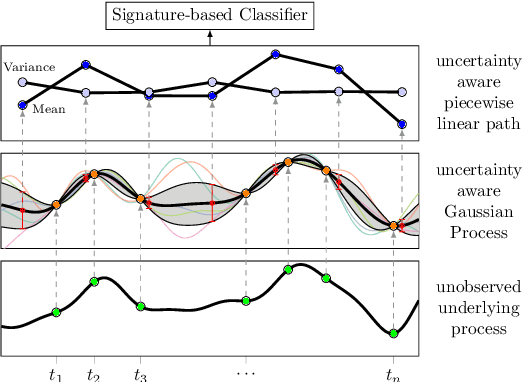
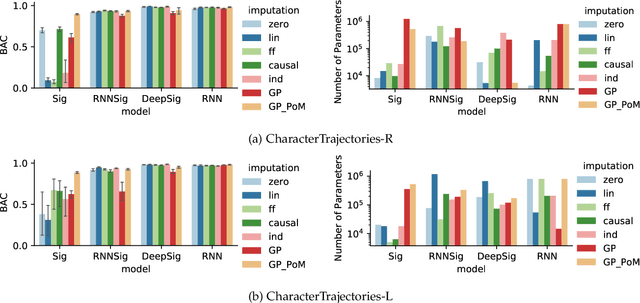
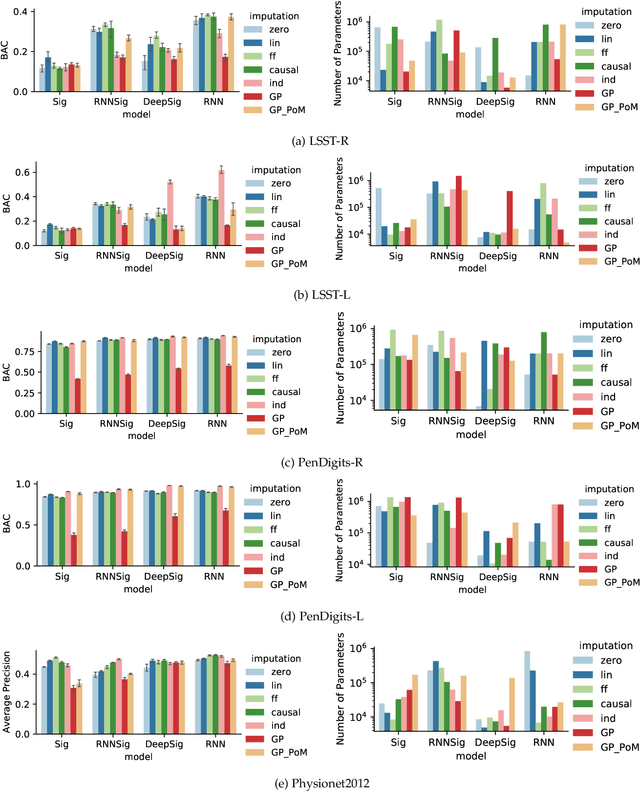
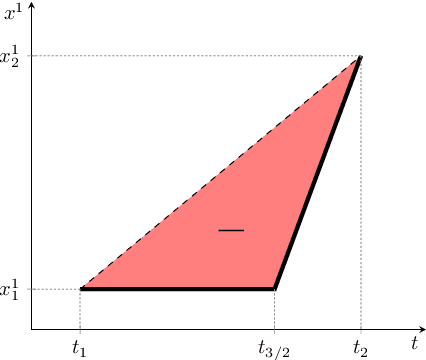
Abstract:The signature transform is a 'universal nonlinearity' on the space of continuous vector-valued paths, and has received attention for use in machine learning on time series. However, real-world temporal data is typically observed at discrete points in time, and must first be transformed into a continuous path before signature techniques can be applied. We make this step explicit by characterising it as an imputation problem, and empirically assess the impact of various imputation strategies when applying signature-based neural nets to irregular time series data. For one of these strategies, Gaussian process (GP) adapters, we propose an extension~(GP-PoM) that makes uncertainty information directly available to the subsequent classifier while at the same time preventing costly Monte-Carlo (MC) sampling. In our experiments, we find that the choice of imputation drastically affects shallow signature models, whereas deeper architectures are more robust. Next, we observe that uncertainty-aware predictions (based on GP-PoM or indicator imputations) are beneficial for predictive performance, even compared to the uncertainty-aware training of conventional GP adapters. In conclusion, we have demonstrated that the path construction is indeed crucial for signature models and that our proposed strategy leads to competitive performance in general, while improving robustness of signature models in particular.
Set Functions for Time Series
Sep 26, 2019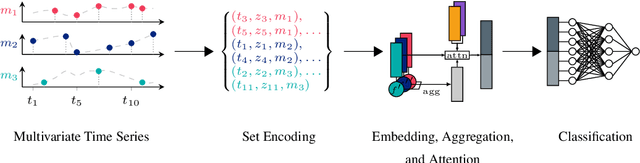
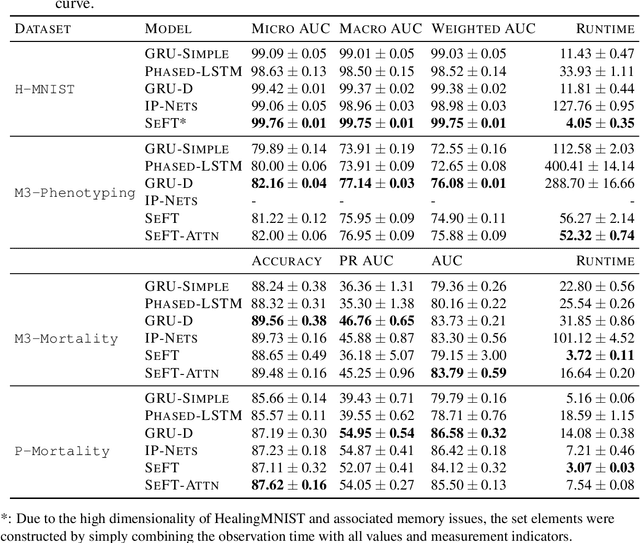
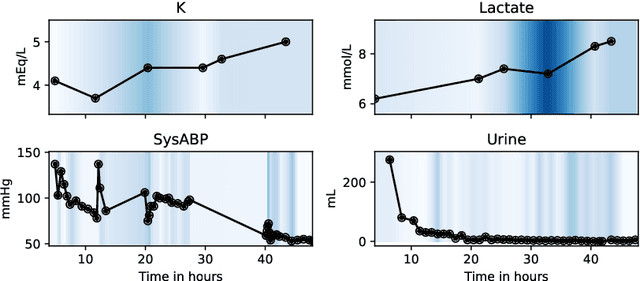
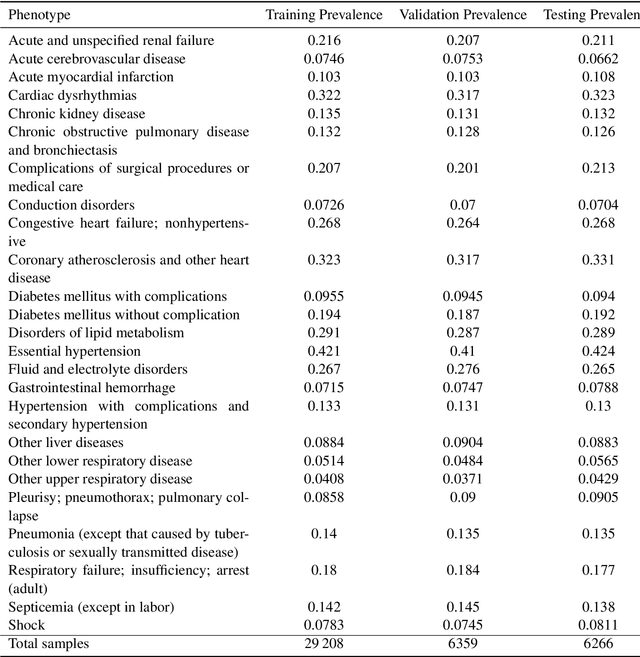
Abstract:Despite the eminent successes of deep neural networks, many architectures are often hard to transfer to irregularly-sampled and asynchronous time series that occur in many real-world datasets, such as healthcare applications. This paper proposes a novel framework for classifying irregularly sampled time series with unaligned measurements, focusing on high scalability and data efficiency. Our method SEFT (Set Functions for Time Series) is based on recent advances in differentiable set function learning, extremely parallelizable, and scales well to very large datasets and online monitoring scenarios. We extensively compare our method to competitors on multiple healthcare time series datasets and show that it performs competitively whilst significantly reducing runtime.
Wasserstein Weisfeiler-Lehman Graph Kernels
Jun 04, 2019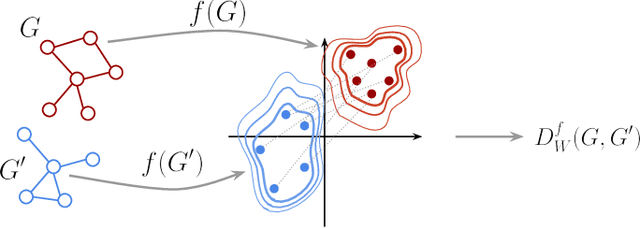
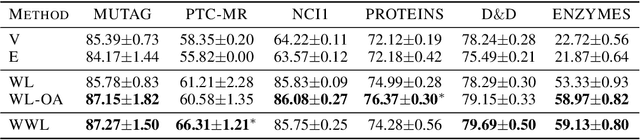
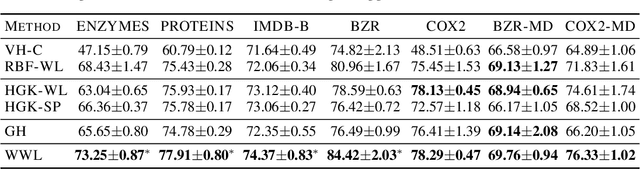
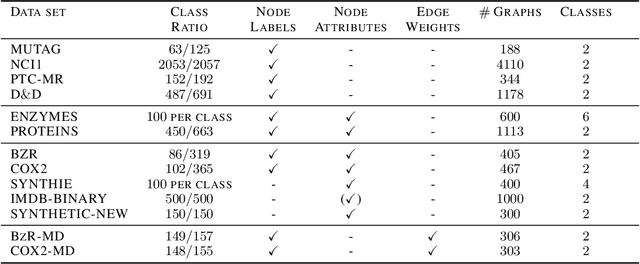
Abstract:Graph kernels are an instance of the class of $\mathcal{R}$-Convolution kernels, which measure the similarity of objects by comparing their substructures. Despite their empirical success, most graph kernels use a naive aggregation of the final set of substructures, usually a sum or average, thereby potentially discarding valuable information about the distribution of individual components. Furthermore, only a limited instance of these approaches can be extended to continuously attributed graphs. We propose a novel method that relies on the Wasserstein distance between the node feature vector distributions of two graphs, which allows to find subtler differences in data sets by considering graphs as high-dimensional objects, rather than simple means. We further propose a Weisfeiler-Lehman inspired embedding scheme for graphs with continuous node attributes and weighted edges, enhance it with the computed Wasserstein distance, and thus improve the state-of-the-art prediction performance on several graph classification tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge