John A. Lee
Perplexity-free Parametric t-SNE
Oct 03, 2020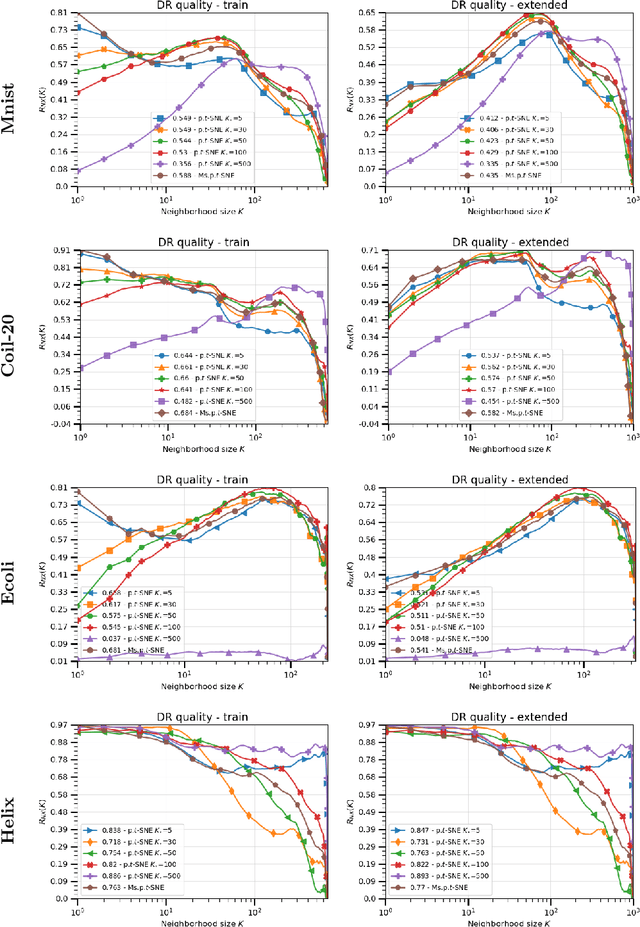
Abstract:The t-distributed Stochastic Neighbor Embedding (t-SNE) algorithm is a ubiquitously employed dimensionality reduction (DR) method. Its non-parametric nature and impressive efficacy motivated its parametric extension. It is however bounded to a user-defined perplexity parameter, restricting its DR quality compared to recently developed multi-scale perplexity-free approaches. This paper hence proposes a multi-scale parametric t-SNE scheme, relieved from the perplexity tuning and with a deep neural network implementing the mapping. It produces reliable embeddings with out-of-sample extensions, competitive with the best perplexity adjustments in terms of neighborhood preservation on multiple data sets.
Deep Learning to Detect Bacterial Colonies for the Production of Vaccines
Sep 02, 2020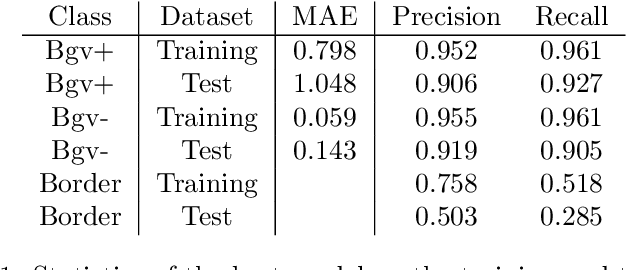
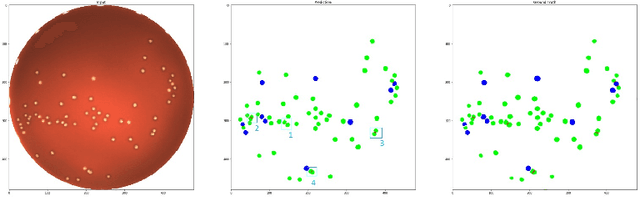
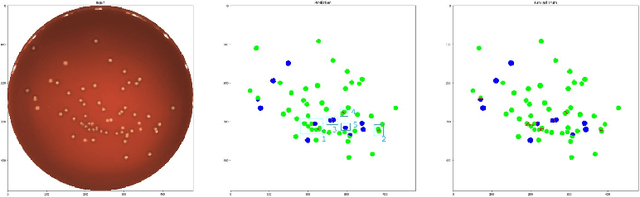
Abstract:During the development of vaccines, bacterial colony forming units (CFUs) are counted in order to quantify the yield in the fermentation process. This manual task is time-consuming and error-prone. In this work we test multiple segmentation algorithms based on the U-Net CNN architecture and show that these offer robust, automated CFU counting. We show that the multiclass generalisation with a bespoke loss function allows distinguishing virulent and avirulent colonies with acceptable accuracy. While many possibilities are left to explore, our results show the potential of deep learning for separating and classifying bacterial colonies.
Knowing what you know in brain segmentation using deep neural networks
Dec 18, 2018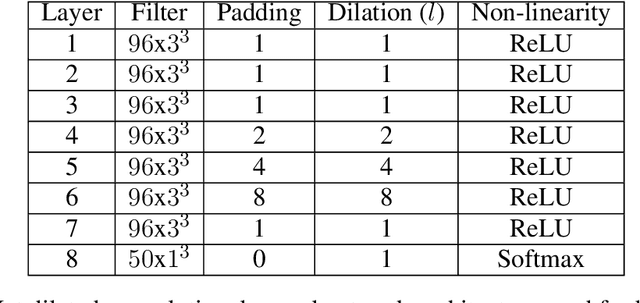
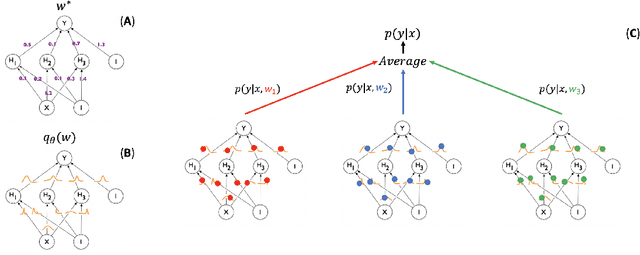
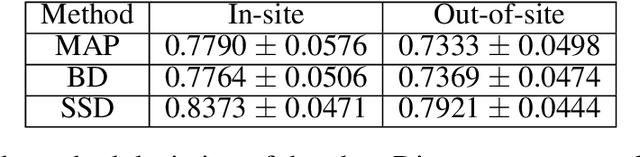
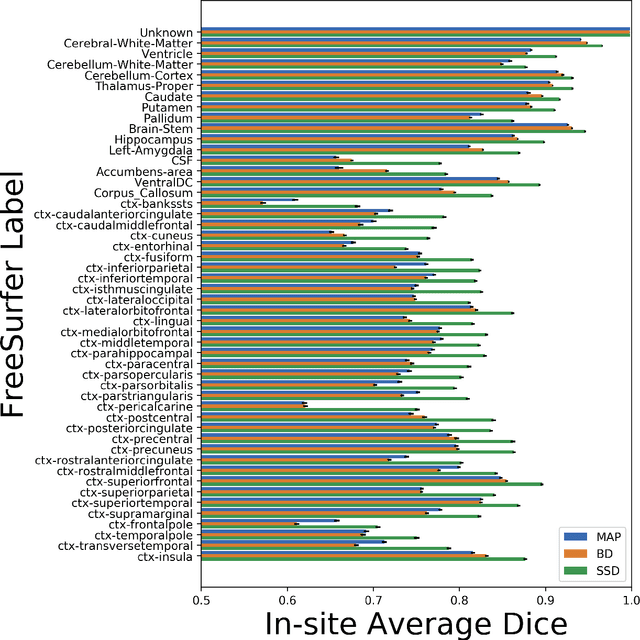
Abstract:In this paper, we describe a deep neural network trained to predict FreeSurfer segmentations of structural MRI volumes, in seconds rather than hours. The network was trained and evaluated on an extremely large dataset (n = 11,148), obtained by combining data from more than a hundred sites. We also show that the prediction uncertainty of the network at each voxel is a good indicator of whether the network has made an error. The resulting uncertainty volume can be used in conjunction with the predicted segmentation to improve downstream uses, such as calculation of measures derived from segmentation regions of interest or the building of prediction models. Finally, we demonstrate that the average prediction uncertainty across voxels in the brain is an excellent indicator of manual quality control ratings, outperforming the best available automated solutions.
Capturing Variabilities from Computed Tomography Images with Generative Adversarial Networks
May 29, 2018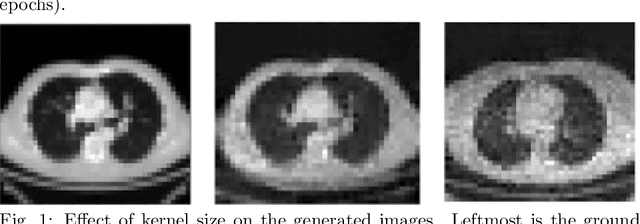
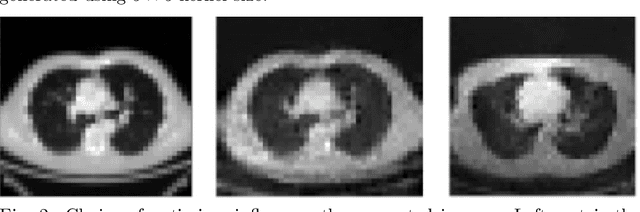
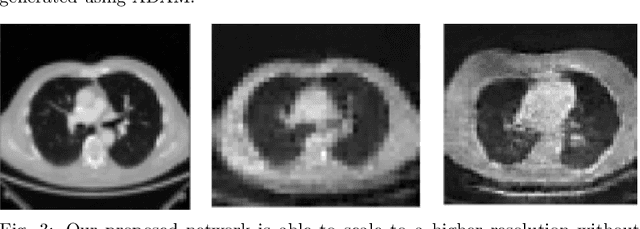
Abstract:With the advent of Deep Learning (DL) techniques, especially Generative Adversarial Networks (GANs), data augmentation and generation are quickly evolving domains that have raised much interest recently. However, the DL techniques are data demanding and since, medical data is not easily accessible, they suffer from data insufficiency. To deal with this limitation, different data augmentation techniques are used. Here, we propose a novel unsupervised data-driven approach for data augmentation that can generate 2D Computed Tomography (CT) images using a simple GAN. The generated CT images have good global and local features of a real CT image and can be used to augment the training datasets for effective learning. In this proof-of-concept study, we show that our proposed solution using GANs is able to capture some of the global and local CT variabilities. Our network is able to generate visually realistic CT images and we aim to further enhance its output by scaling it to a higher resolution and potentially from 2D to 3D.
Blind Deconvolution of PET Images using Anatomical Priors
Aug 05, 2016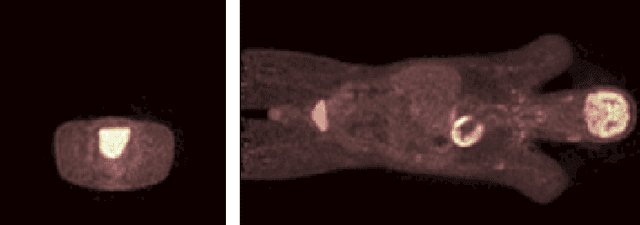
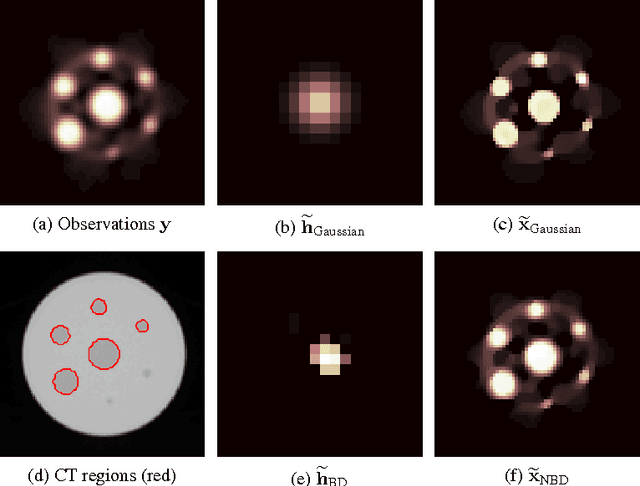
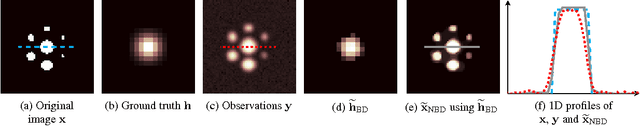
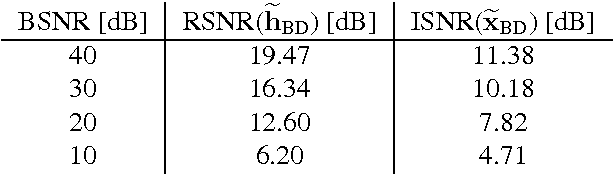


Abstract:Images from positron emission tomography (PET) provide metabolic information about the human body. They present, however, a spatial resolution that is limited by physical and instrumental factors often modeled by a blurring function. Since this function is typically unknown, blind deconvolution (BD) techniques are needed in order to produce a useful restored PET image. In this work, we propose a general BD technique that restores a low resolution blurry image using information from data acquired with a high resolution modality (e.g., CT-based delineation of regions with uniform activity in PET images). The proposed BD method is validated on synthetic and actual phantoms.
Post-Reconstruction Deconvolution of PET Images by Total Generalized Variation Regularization
Jun 16, 2015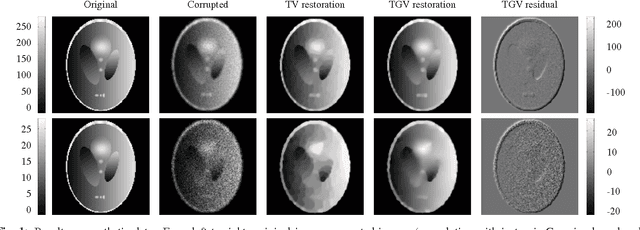
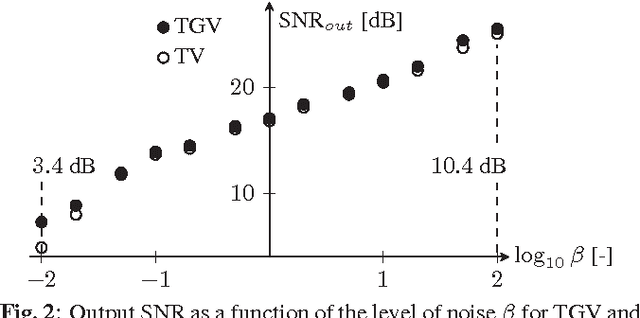
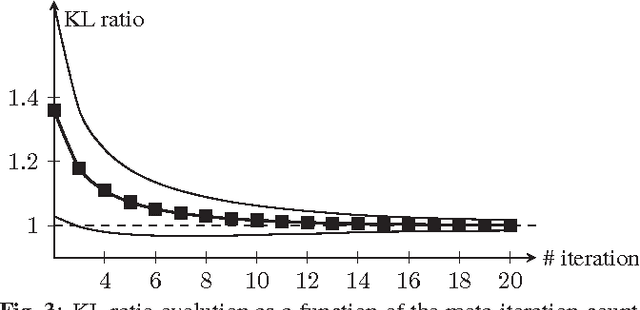
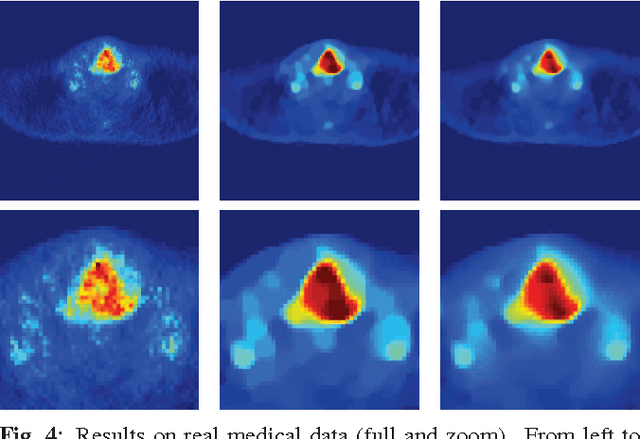
Abstract:Improving the quality of positron emission tomography (PET) images, affected by low resolution and high level of noise, is a challenging task in nuclear medicine and radiotherapy. This work proposes a restoration method, achieved after tomographic reconstruction of the images and targeting clinical situations where raw data are often not accessible. Based on inverse problem methods, our contribution introduces the recently developed total generalized variation (TGV) norm to regularize PET image deconvolution. Moreover, we stabilize this procedure with additional image constraints such as positivity and photometry invariance. A criterion for updating and adjusting automatically the regularization parameter in case of Poisson noise is also presented. Experiments are conducted on both synthetic data and real patient images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge