Jesper B. Christensen
Ctrl-A: Control-Driven Online Data Augmentation
Mar 23, 2026Abstract:We introduce ControlAugment (Ctrl-A), an automated data augmentation algorithm for image-vision tasks, which incorporates principles from control theory for online adjustment of augmentation strength distributions during model training. Ctrl-A eliminates the need for initialization of individual augmentation strengths. Instead, augmentation strength distributions are dynamically, and individually, adapted during training based on a control-loop architecture and what we define as relative operation response curves. Using an operation-dependent update procedure provides Ctrl-A with the potential to suppress augmentation styles that negatively impact model performance, alleviating the need for manually engineering augmentation policies for new image-vision tasks. Experiments on the CIFAR-10, CIFAR-100, and SVHN-core benchmark datasets using the common WideResNet-28-10 architecture demonstrate that Ctrl-A is highly competitive with existing state-of-the-art data augmentation strategies.
Accurate and fast identification of minimally prepared bacteria phenotypes using Raman spectroscopy assisted by machine learning
Jun 27, 2022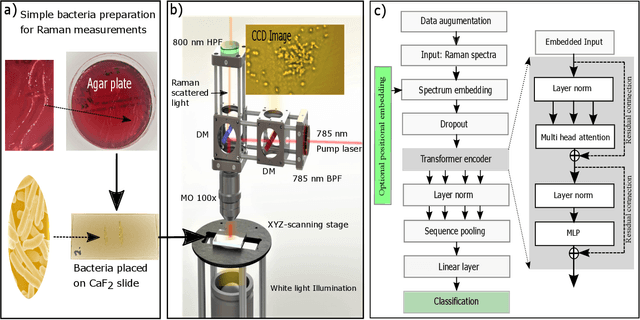
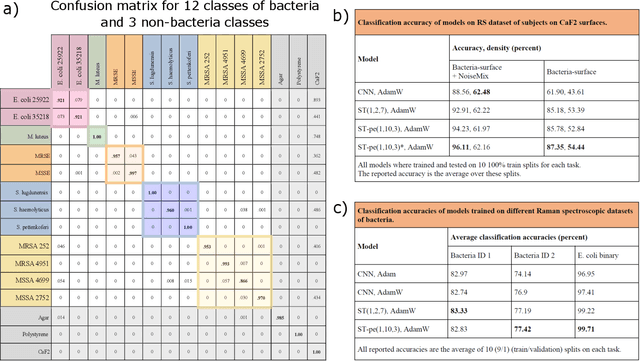
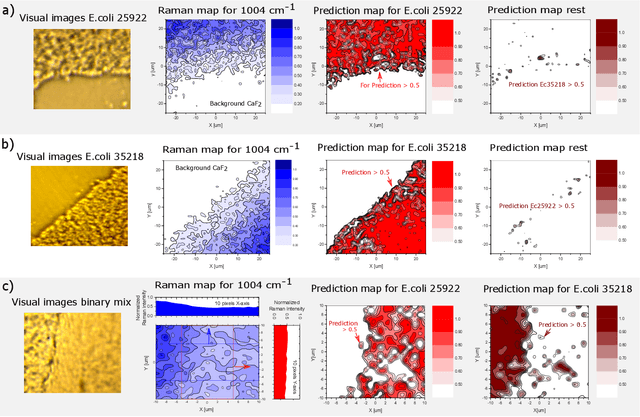
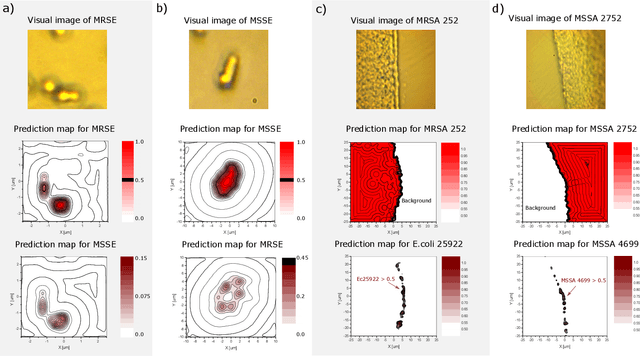
Abstract:The worldwide increase of antimicrobial resistance (AMR) is a serious threat to human health. To avert the spread of AMR, fast reliable diagnostics tools that facilitate optimal antibiotic stewardship are an unmet need. In this regard, Raman spectroscopy promises rapid label- and culture-free identification and antimicrobial susceptibility testing (AST) in a single step. However, even though many Raman-based bacteria-identification and AST studies have demonstrated impressive results, some shortcomings must be addressed. To bridge the gap between proof-of-concept studies and clinical application, we have developed machine learning techniques in combination with a novel data-augmentation algorithm, for fast identification of minimally prepared bacteria phenotypes and the distinctions of methicillin-resistant (MR) from methicillin-susceptible (MS) bacteria. For this we have implemented a spectral transformer model for hyper-spectral Raman images of bacteria. We show that our model outperforms the standard convolutional neural network models on a multitude of classification problems, both in terms of accuracy and in terms of training time. We attain more than 96$\%$ classification accuracy on a dataset consisting of 15 different classes and 95.6$\%$ classification accuracy for six MR-MS bacteria species. More importantly, our results are obtained using only fast and easy-to-produce training and test data
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge