Jasper Friedrichs
Did you take the pill? - Detecting Personal Intake of Medicine from Twitter
Aug 03, 2018
Abstract:Mining social media messages such as tweets, articles, and Facebook posts for health and drug related information has received significant interest in pharmacovigilance research. Social media sites (e.g., Twitter), have been used for monitoring drug abuse, adverse reactions of drug usage and analyzing expression of sentiments related to drugs. Most of these studies are based on aggregated results from a large population rather than specific sets of individuals. In order to conduct studies at an individual level or specific cohorts, identifying posts mentioning intake of medicine by the user is necessary. Towards this objective we develop a classifier for identifying mentions of personal intake of medicine in tweets. We train a stacked ensemble of shallow convolutional neural network (CNN) models on an annotated dataset. We use random search for tuning the hyper-parameters of the CNN models and present an ensemble of best models for the prediction task. Our system produces state-of-the-art result, with a micro-averaged F-score of 0.693. We believe that the developed classifier has direct uses in the areas of psychology, health informatics, pharmacovigilance and affective computing for tracking moods, emotions and sentiments of patients expressing intake of medicine in social media.
#phramacovigilance - Exploring Deep Learning Techniques for Identifying Mentions of Medication Intake from Twitter
May 16, 2018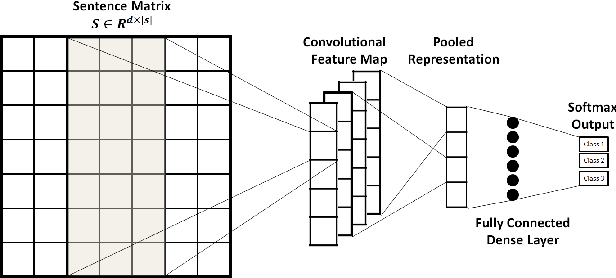
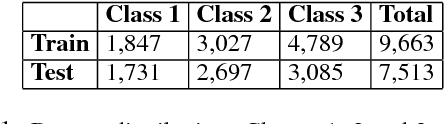
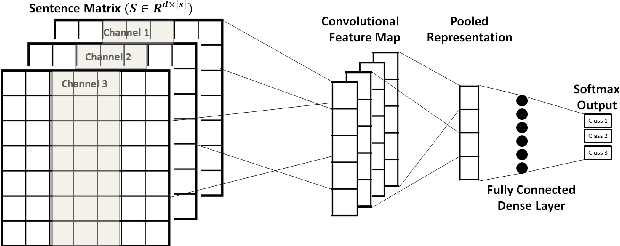
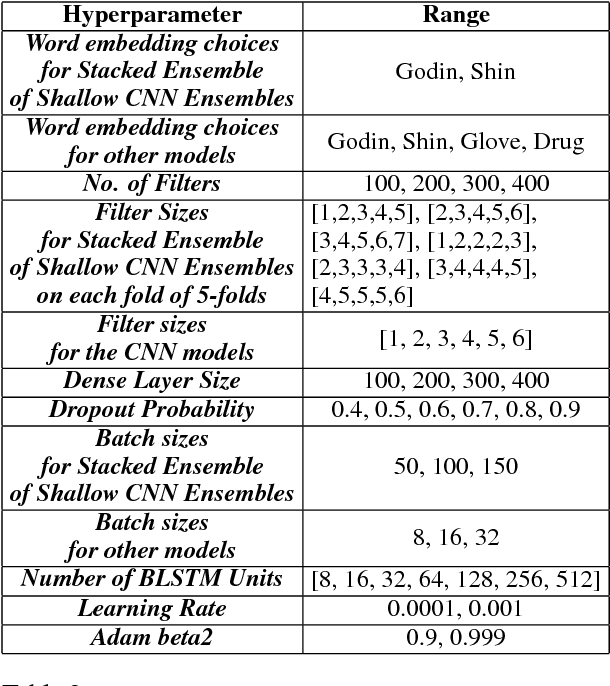
Abstract:Mining social media messages for health and drug related information has received significant interest in pharmacovigilance research. Social media sites (e.g., Twitter), have been used for monitoring drug abuse, adverse reactions of drug usage and analyzing expression of sentiments related to drugs. Most of these studies are based on aggregated results from a large population rather than specific sets of individuals. In order to conduct studies at an individual level or specific cohorts, identifying posts mentioning intake of medicine by the user is necessary. Towards this objective, we train different deep neural network classification models on a publicly available annotated dataset and study their performances on identifying mentions of personal intake of medicine in tweets. We also design and train a new architecture of a stacked ensemble of shallow convolutional neural network (CNN) ensembles. We use random search for tuning the hyperparameters of the models and share the details of the values taken by the hyperparameters for the best learnt model in different deep neural network architectures. Our system produces state-of-the-art results, with a micro- averaged F-score of 0.693.
InfyNLP at SMM4H Task 2: Stacked Ensemble of Shallow Convolutional Neural Networks for Identifying Personal Medication Intake from Twitter
Mar 21, 2018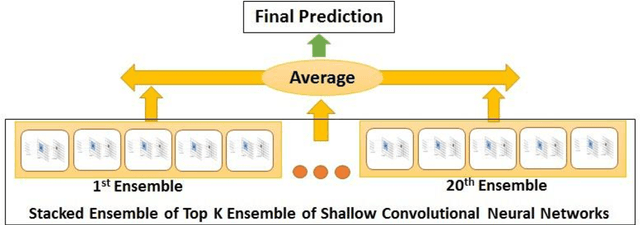
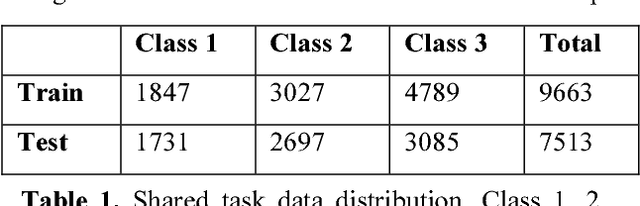
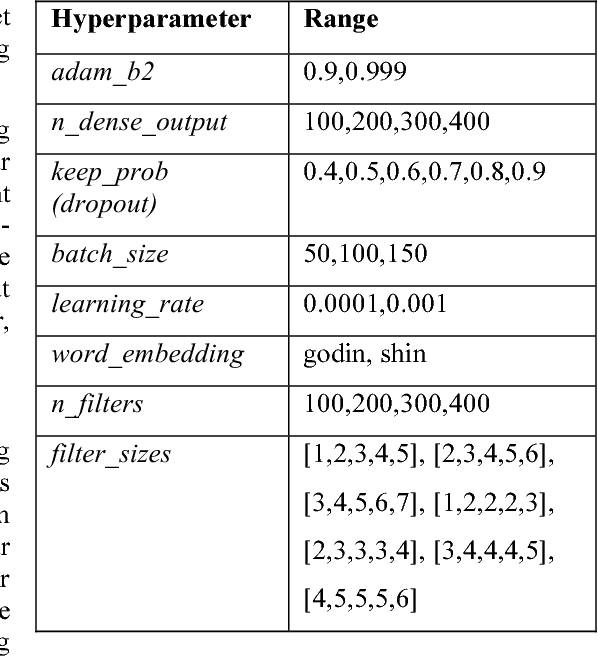
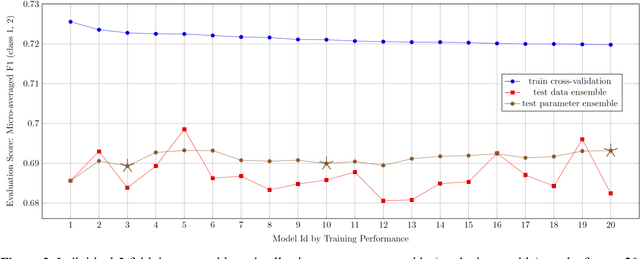
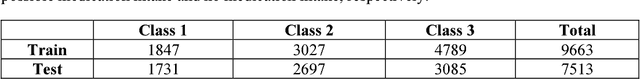
Abstract:This paper describes Infosys's participation in the "2nd Social Media Mining for Health Applications Shared Task at AMIA, 2017, Task 2". Mining social media messages for health and drug related information has received significant interest in pharmacovigilance research. This task targets at developing automated classification models for identifying tweets containing descriptions of personal intake of medicines. Towards this objective we train a stacked ensemble of shallow convolutional neural network (CNN) models on an annotated dataset provided by the organizers. We use random search for tuning the hyper-parameters of the CNN and submit an ensemble of best models for the prediction task. Our system secured first place among 9 teams, with a micro-averaged F-score of 0.693.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge