Jason Dunkelberger
AstaBench: Rigorous Benchmarking of AI Agents with a Scientific Research Suite
Oct 24, 2025Abstract:AI agents hold the potential to revolutionize scientific productivity by automating literature reviews, replicating experiments, analyzing data, and even proposing new directions of inquiry; indeed, there are now many such agents, ranging from general-purpose "deep research" systems to specialized science-specific agents, such as AI Scientist and AIGS. Rigorous evaluation of these agents is critical for progress. Yet existing benchmarks fall short on several fronts: they (1) fail to provide holistic, product-informed measures of real-world use cases such as science research; (2) lack reproducible agent tools necessary for a controlled comparison of core agentic capabilities; (3) do not account for confounding variables such as model cost and tool access; (4) do not provide standardized interfaces for quick agent prototyping and evaluation; and (5) lack comprehensive baseline agents necessary to identify true advances. In response, we define principles and tooling for more rigorously benchmarking agents. Using these, we present AstaBench, a suite that provides the first holistic measure of agentic ability to perform scientific research, comprising 2400+ problems spanning the entire scientific discovery process and multiple scientific domains, and including many problems inspired by actual user requests to deployed Asta agents. Our suite comes with the first scientific research environment with production-grade search tools that enable controlled, reproducible evaluation, better accounting for confounders. Alongside, we provide a comprehensive suite of nine science-optimized classes of Asta agents and numerous baselines. Our extensive evaluation of 57 agents across 22 agent classes reveals several interesting findings, most importantly that despite meaningful progress on certain individual aspects, AI remains far from solving the challenge of science research assistance.
olmOCR: Unlocking Trillions of Tokens in PDFs with Vision Language Models
Feb 25, 2025Abstract:PDF documents have the potential to provide trillions of novel, high-quality tokens for training language models. However, these documents come in a diversity of types with differing formats and visual layouts that pose a challenge when attempting to extract and faithfully represent the underlying content for language model use. We present olmOCR, an open-source Python toolkit for processing PDFs into clean, linearized plain text in natural reading order while preserving structured content like sections, tables, lists, equations, and more. Our toolkit runs a fine-tuned 7B vision language model (VLM) trained on a sample of 260,000 pages from over 100,000 crawled PDFs with diverse properties, including graphics, handwritten text and poor quality scans. olmOCR is optimized for large-scale batch processing, able to scale flexibly to different hardware setups and convert a million PDF pages for only $190 USD. We release all components of olmOCR including VLM weights, data and training code, as well as inference code built on serving frameworks including vLLM and SGLang.
The Semantic Scholar Open Data Platform
Jan 24, 2023



Abstract:The volume of scientific output is creating an urgent need for automated tools to help scientists keep up with developments in their field. Semantic Scholar (S2) is an open data platform and website aimed at accelerating science by helping scholars discover and understand scientific literature. We combine public and proprietary data sources using state-of-the-art techniques for scholarly PDF content extraction and automatic knowledge graph construction to build the Semantic Scholar Academic Graph, the largest open scientific literature graph to-date, with 200M+ papers, 80M+ authors, 550M+ paper-authorship edges, and 2.4B+ citation edges. The graph includes advanced semantic features such as structurally parsed text, natural language summaries, and vector embeddings. In this paper, we describe the components of the S2 data processing pipeline and the associated APIs offered by the platform. We will update this living document to reflect changes as we add new data offerings and improve existing services.
Construction of the Literature Graph in Semantic Scholar
May 06, 2018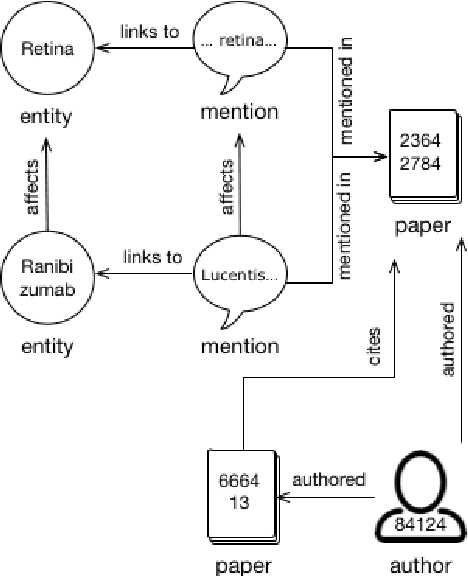
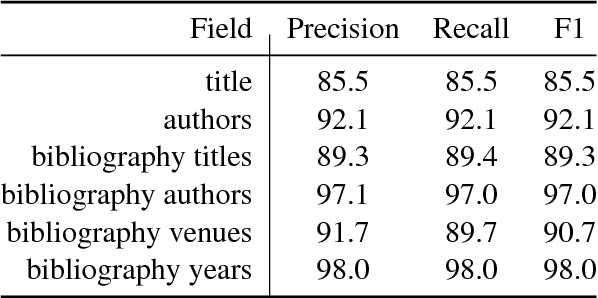
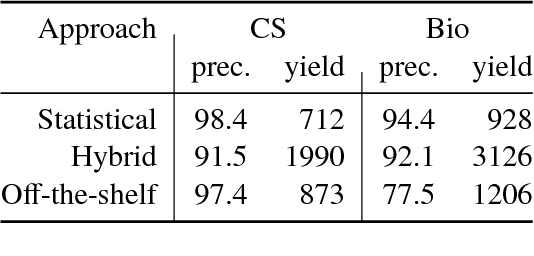

Abstract:We describe a deployed scalable system for organizing published scientific literature into a heterogeneous graph to facilitate algorithmic manipulation and discovery. The resulting literature graph consists of more than 280M nodes, representing papers, authors, entities and various interactions between them (e.g., authorships, citations, entity mentions). We reduce literature graph construction into familiar NLP tasks (e.g., entity extraction and linking), point out research challenges due to differences from standard formulations of these tasks, and report empirical results for each task. The methods described in this paper are used to enable semantic features in www.semanticscholar.org
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge