Jérémie Decouchant
Dynamic Topology Optimization for Non-IID Data in Decentralized Learning
Feb 03, 2026Abstract:Decentralized learning (DL) enables a set of nodes to train a model collaboratively without central coordination, offering benefits for privacy and scalability. However, DL struggles to train a high accuracy model when the data distribution is non-independent and identically distributed (non-IID) and when the communication topology is static. To address these issues, we propose Morph, a topology optimization algorithm for DL. In Morph, nodes adaptively choose peers for model exchange based on maximum model dissimilarity. Morph maintains a fixed in-degree while dynamically reshaping the communication graph through gossip-based peer discovery and diversity-driven neighbor selection, thereby improving robustness to data heterogeneity. Experiments on CIFAR-10 and FEMNIST with up to 100 nodes show that Morph consistently outperforms static and epidemic baselines, while closely tracking the fully connected upper bound. On CIFAR-10, Morph achieves a relative improvement of 1.12x in test accuracy compared to the state-of-the-art baselines. On FEMNIST, Morph achieves an accuracy that is 1.08x higher than Epidemic Learning. Similar trends hold for 50 node deployments, where Morph narrows the gap to the fully connected upper bound within 0.5 percentage points on CIFAR-10. These results demonstrate that Morph achieves higher final accuracy, faster convergence, and more stable learning as quantified by lower inter-node variance, while requiring fewer communication rounds than baselines and no global knowledge.
Training Diffusion Models with Federated Learning
Jun 18, 2024



Abstract:The training of diffusion-based models for image generation is predominantly controlled by a select few Big Tech companies, raising concerns about privacy, copyright, and data authority due to their lack of transparency regarding training data. To ad-dress this issue, we propose a federated diffusion model scheme that enables the independent and collaborative training of diffusion models without exposing local data. Our approach adapts the Federated Averaging (FedAvg) algorithm to train a Denoising Diffusion Model (DDPM). Through a novel utilization of the underlying UNet backbone, we achieve a significant reduction of up to 74% in the number of parameters exchanged during training,compared to the naive FedAvg approach, whilst simultaneously maintaining image quality comparable to the centralized setting, as evaluated by the FID score.
Parameterizing Federated Continual Learning for Reproducible Research
Jun 04, 2024



Abstract:Federated Learning (FL) systems evolve in heterogeneous and ever-evolving environments that challenge their performance. Under real deployments, the learning tasks of clients can also evolve with time, which calls for the integration of methodologies such as Continual Learning. To enable research reproducibility, we propose a set of experimental best practices that precisely capture and emulate complex learning scenarios. Our framework, Freddie, is the first entirely configurable framework for Federated Continual Learning (FCL), and it can be seamlessly deployed on a large number of machines thanks to the use of Kubernetes and containerization. We demonstrate the effectiveness of Freddie on two use cases, (i) large-scale FL on CIFAR100 and (ii) heterogeneous task sequence on FCL, which highlight unaddressed performance challenges in FCL scenarios.
Asynchronous Multi-Server Federated Learning for Geo-Distributed Clients
Jun 03, 2024



Abstract:Federated learning (FL) systems enable multiple clients to train a machine learning model iteratively through synchronously exchanging the intermediate model weights with a single server. The scalability of such FL systems can be limited by two factors: server idle time due to synchronous communication and the risk of a single server becoming the bottleneck. In this paper, we propose a new FL architecture, to our knowledge, the first multi-server FL system that is entirely asynchronous, and therefore addresses these two limitations simultaneously. Our solution keeps both servers and clients continuously active. As in previous multi-server methods, clients interact solely with their nearest server, ensuring efficient update integration into the model. Differently, however, servers also periodically update each other asynchronously, and never postpone interactions with clients. We compare our solution to three representative baselines - FedAvg, FedAsync and HierFAVG - on the MNIST and CIFAR-10 image classification datasets and on the WikiText-2 language modeling dataset. Our solution converges to similar or higher accuracy levels than previous baselines and requires 61% less time to do so in geo-distributed settings.
Asynchronous Byzantine Federated Learning
Jun 03, 2024Abstract:Federated learning (FL) enables a set of geographically distributed clients to collectively train a model through a server. Classically, the training process is synchronous, but can be made asynchronous to maintain its speed in presence of slow clients and in heterogeneous networks. The vast majority of Byzantine fault-tolerant FL systems however rely on a synchronous training process. Our solution is one of the first Byzantine-resilient and asynchronous FL algorithms that does not require an auxiliary server dataset and is not delayed by stragglers, which are shortcomings of previous works. Intuitively, the server in our solution waits to receive a minimum number of updates from clients on its latest model to safely update it, and is later able to safely leverage the updates that late clients might send. We compare the performance of our solution with state-of-the-art algorithms on both image and text datasets under gradient inversion, perturbation, and backdoor attacks. Our results indicate that our solution trains a model faster than previous synchronous FL solution, and maintains a higher accuracy, up to 1.54x and up to 1.75x for perturbation and gradient inversion attacks respectively, in the presence of Byzantine clients than previous asynchronous FL solutions.
Share Your Secrets for Privacy! Confidential Forecasting with Vertical Federated Learning
May 31, 2024



Abstract:Vertical federated learning (VFL) is a promising area for time series forecasting in industrial applications, such as predictive maintenance and machine control. Critical challenges to address in manufacturing include data privacy and over-fitting on small and noisy datasets during both training and inference. Additionally, to increase industry adaptability, such forecasting models must scale well with the number of parties while ensuring strong convergence and low-tuning complexity. We address those challenges and propose 'Secret-shared Time Series Forecasting with VFL' (STV), a novel framework that exhibits the following key features: i) a privacy-preserving algorithm for forecasting with SARIMAX and autoregressive trees on vertically partitioned data; ii) serverless forecasting using secret sharing and multi-party computation; iii) novel N-party algorithms for matrix multiplication and inverse operations for direct parameter optimization, giving strong convergence with minimal hyperparameter tuning complexity. We conduct evaluations on six representative datasets from public and industry-specific contexts. Our results demonstrate that STV's forecasting accuracy is comparable to those of centralized approaches. They also show that our direct optimization can outperform centralized methods, which include state-of-the-art diffusion models and long-short-term memory, by 23.81% on forecasting accuracy. We also conduct a scalability analysis by examining the communication costs of direct and iterative optimization to navigate the choice between the two. Code and appendix are available: https://github.com/adis98/STV
CCBNet: Confidential Collaborative Bayesian Networks Inference
May 23, 2024



Abstract:Effective large-scale process optimization in manufacturing industries requires close cooperation between different human expert parties who encode their knowledge of related domains as Bayesian network models. For instance, Bayesian networks for domains such as lithography equipment, processes, and auxiliary tools must be conjointly used to effectively identify process optimizations in the semiconductor industry. However, business confidentiality across domains hinders such collaboration, and encourages alternatives to centralized inference. We propose CCBNet, the first Confidentiality-preserving Collaborative Bayesian Network inference framework. CCBNet leverages secret sharing to securely perform analysis on the combined knowledge of party models by joining two novel subprotocols: (i) CABN, which augments probability distributions for features across parties by modeling them into secret shares of their normalized combination; and (ii) SAVE, which aggregates party inference result shares through distributed variable elimination. We extensively evaluate CCBNet via 9 public Bayesian networks. Our results show that CCBNet achieves predictive quality that is similar to the ones of centralized methods while preserving model confidentiality. We further demonstrate that CCBNet scales to challenging manufacturing use cases that involve 16-128 parties in large networks of 223-1003 features, and decreases, on average, computational overhead by 23%, while communicating 71k values per request. Finally, we showcase possible attacks and mitigations for partially reconstructing party networks in the two subprotocols.
Aergia: Leveraging Heterogeneity in Federated Learning Systems
Oct 12, 2022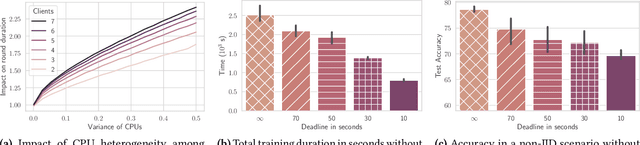
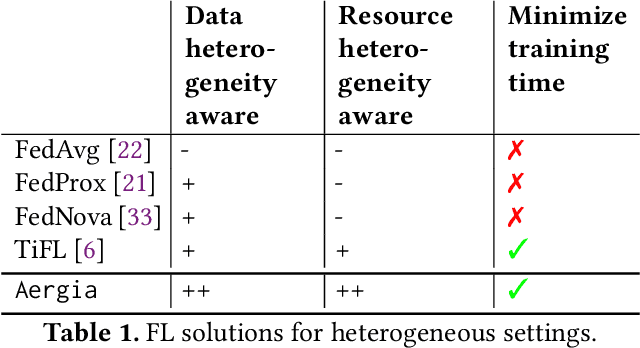
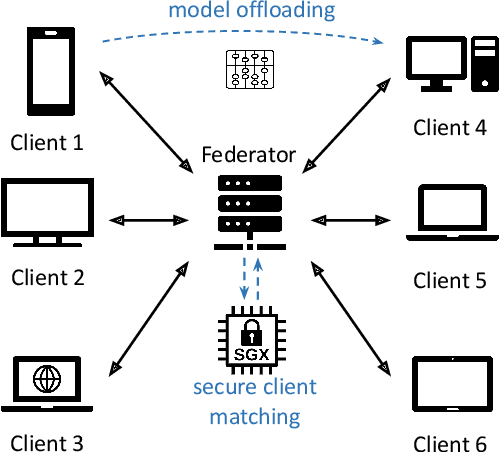
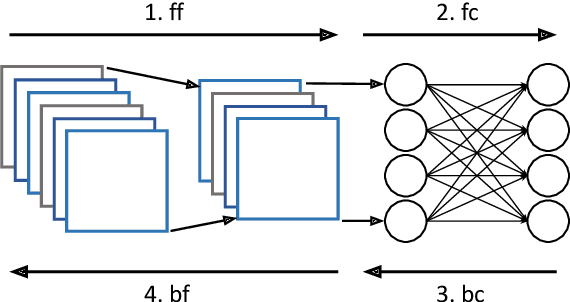
Abstract:Federated Learning (FL) is a popular approach for distributed deep learning that prevents the pooling of large amounts of data in a central server. FL relies on clients to update a global model using their local datasets. Classical FL algorithms use a central federator that, for each training round, waits for all clients to send their model updates before aggregating them. In practical deployments, clients might have different computing powers and network capabilities, which might lead slow clients to become performance bottlenecks. Previous works have suggested to use a deadline for each learning round so that the federator ignores the late updates of slow clients, or so that clients send partially trained models before the deadline. To speed up the training process, we instead propose Aergia, a novel approach where slow clients (i) freeze the part of their model that is the most computationally intensive to train; (ii) train the unfrozen part of their model; and (iii) offload the training of the frozen part of their model to a faster client that trains it using its own dataset. The offloading decisions are orchestrated by the federator based on the training speed that clients report and on the similarities between their datasets, which are privately evaluated thanks to a trusted execution environment. We show through extensive experiments that Aergia maintains high accuracy and significantly reduces the training time under heterogeneous settings by up to 27% and 53% compared to FedAvg and TiFL, respectively.
I-GWAS: Privacy-Preserving Interdependent Genome-Wide Association Studies
Aug 17, 2022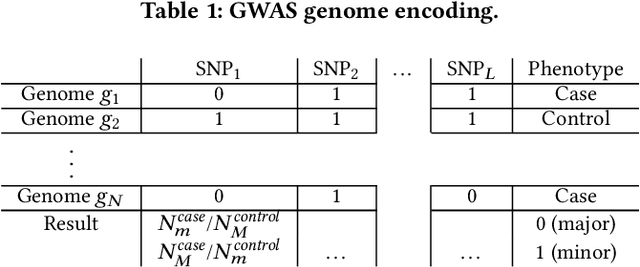
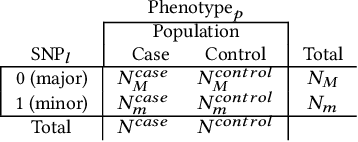
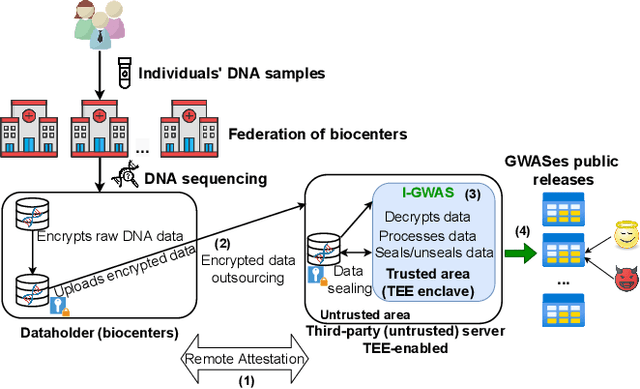

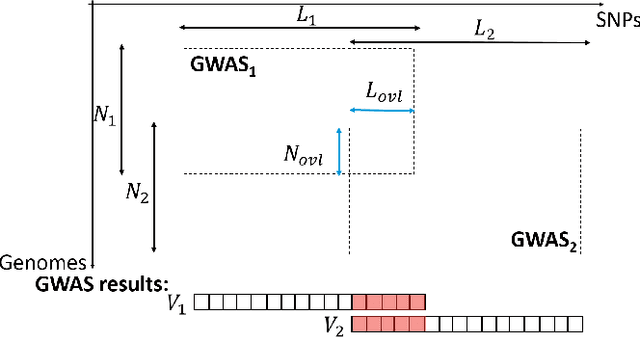
Abstract:Genome-wide Association Studies (GWASes) identify genomic variations that are statistically associated with a trait, such as a disease, in a group of individuals. Unfortunately, careless sharing of GWAS statistics might give rise to privacy attacks. Several works attempted to reconcile secure processing with privacy-preserving releases of GWASes. However, we highlight that these approaches remain vulnerable if GWASes utilize overlapping sets of individuals and genomic variations. In such conditions, we show that even when relying on state-of-the-art techniques for protecting releases, an adversary could reconstruct the genomic variations of up to 28.6% of participants, and that the released statistics of up to 92.3% of the genomic variations would enable membership inference attacks. We introduce I-GWAS, a novel framework that securely computes and releases the results of multiple possibly interdependent GWASes. I-GWAScontinuously releases privacy-preserving and noise-free GWAS results as new genomes become available.
AGIC: Approximate Gradient Inversion Attack on Federated Learning
Apr 28, 2022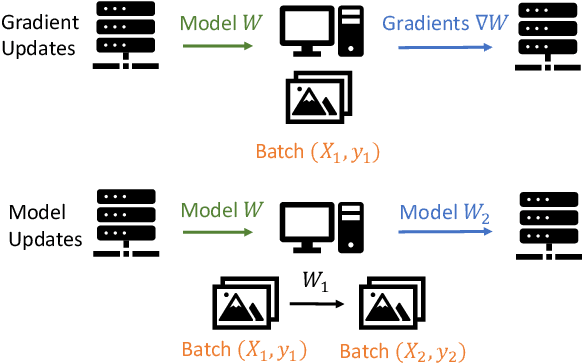

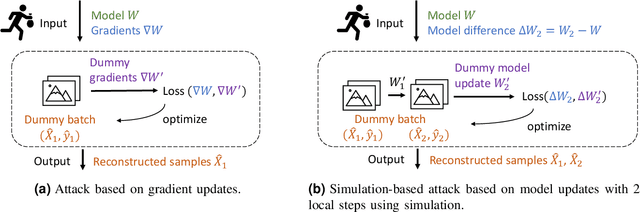
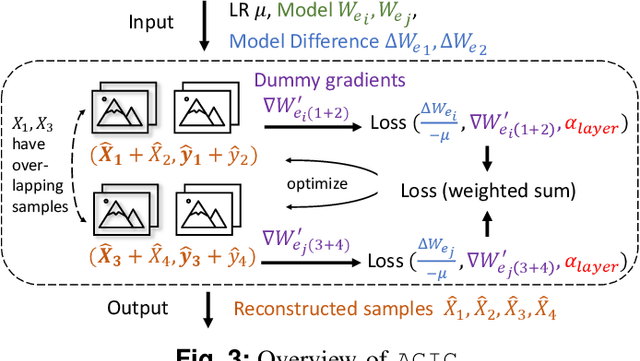
Abstract:Federated learning is a private-by-design distributed learning paradigm where clients train local models on their own data before a central server aggregates their local updates to compute a global model. Depending on the aggregation method used, the local updates are either the gradients or the weights of local learning models. Recent reconstruction attacks apply a gradient inversion optimization on the gradient update of a single minibatch to reconstruct the private data used by clients during training. As the state-of-the-art reconstruction attacks solely focus on single update, realistic adversarial scenarios are overlooked, such as observation across multiple updates and updates trained from multiple mini-batches. A few studies consider a more challenging adversarial scenario where only model updates based on multiple mini-batches are observable, and resort to computationally expensive simulation to untangle the underlying samples for each local step. In this paper, we propose AGIC, a novel Approximate Gradient Inversion Attack that efficiently and effectively reconstructs images from both model or gradient updates, and across multiple epochs. In a nutshell, AGIC (i) approximates gradient updates of used training samples from model updates to avoid costly simulation procedures, (ii) leverages gradient/model updates collected from multiple epochs, and (iii) assigns increasing weights to layers with respect to the neural network structure for reconstruction quality. We extensively evaluate AGIC on three datasets, CIFAR-10, CIFAR-100 and ImageNet. Our results show that AGIC increases the peak signal-to-noise ratio (PSNR) by up to 50% compared to two representative state-of-the-art gradient inversion attacks. Furthermore, AGIC is faster than the state-of-the-art simulation based attack, e.g., it is 5x faster when attacking FedAvg with 8 local steps in between model updates.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge