Irfan Al-Hussaini
CCS Explorer: Relevance Prediction, Extractive Summarization, and Named Entity Recognition from Clinical Cohort Studies
Nov 16, 2022



Abstract:Clinical Cohort Studies (CCS), such as randomized clinical trials, are a great source of documented clinical research. Ideally, a clinical expert inspects these articles for exploratory analysis ranging from drug discovery for evaluating the efficacy of existing drugs in tackling emerging diseases to the first test of newly developed drugs. However, more than 100 articles are published daily on a single prevalent disease like COVID-19 in PubMed. As a result, it can take days for a physician to find articles and extract relevant information. Can we develop a system to sift through the long list of these articles faster and document the crucial takeaways from each of these articles? In this work, we propose CCS Explorer, an end-to-end system for relevance prediction of sentences, extractive summarization, and patient, outcome, and intervention entity detection from CCS. CCS Explorer is packaged in a web-based graphical user interface where the user can provide any disease name. CCS Explorer then extracts and aggregates all relevant information from articles on PubMed based on the results of an automatically generated query produced on the back-end. For each task, CCS Explorer fine-tunes pre-trained language representation models based on transformers with additional layers. The models are evaluated using two publicly available datasets. CCS Explorer obtains a recall of 80.2%, AUC-ROC of 0.843, and an accuracy of 88.3% on sentence relevance prediction using BioBERT and achieves an average Micro F1-Score of 77.8% on Patient, Intervention, Outcome detection (PIO) using PubMedBERT. Thus, CCS Explorer can reliably extract relevant information to summarize articles, saving time by $\sim \text{660}\times$.
Performance and utility trade-off in interpretable sleep staging
Nov 15, 2022



Abstract:Recent advances in deep learning have led to the development of models approaching the human level of accuracy. However, healthcare remains an area lacking in widespread adoption. The safety-critical nature of healthcare results in a natural reticence to put these black-box deep learning models into practice. This paper explores interpretable methods for a clinical decision support system called sleep staging, an essential step in diagnosing sleep disorders. Clinical sleep staging is an arduous process requiring manual annotation for each 30s of sleep using physiological signals such as electroencephalogram (EEG). Recent work has shown that sleep staging using simple models and an exhaustive set of features can perform nearly as well as deep learning approaches but only for some specific datasets. Moreover, the utility of those features from a clinical standpoint is ambiguous. On the other hand, the proposed framework, NormIntSleep demonstrates exceptional performance across different datasets by representing deep learning embeddings using normalized features. NormIntSleep performs 4.5% better than the exhaustive feature-based approach and 1.5% better than other representation learning approaches. An empirical comparison between the utility of the interpretations of these models highlights the improved alignment with clinical expectations when performance is traded-off slightly. NormIntSleep paired with a clinically meaningful set of features can best balance this trade-off by providing reliable, clinically relevant interpretation with robust performance.
SERF: Interpretable Sleep Staging using Embeddings, Rules, and Features
Sep 25, 2022
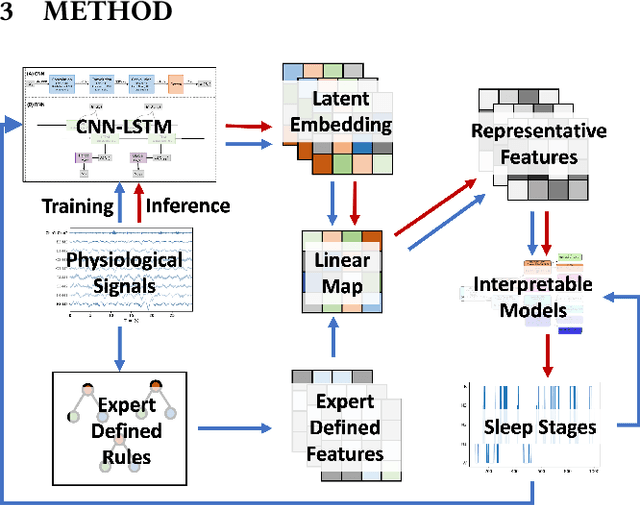
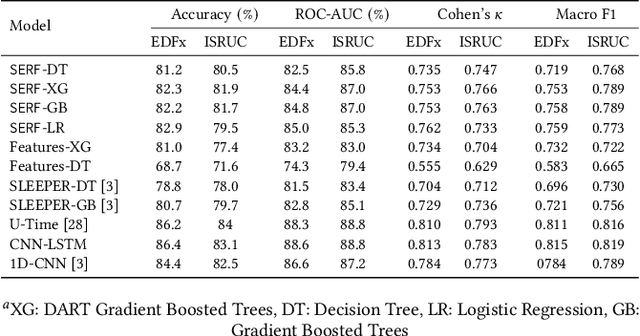
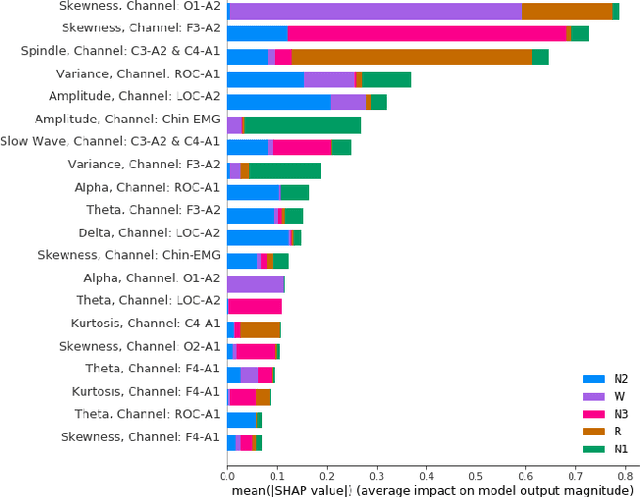
Abstract:The accuracy of recent deep learning based clinical decision support systems is promising. However, lack of model interpretability remains an obstacle to widespread adoption of artificial intelligence in healthcare. Using sleep as a case study, we propose a generalizable method to combine clinical interpretability with high accuracy derived from black-box deep learning. Clinician-determined sleep stages from polysomnogram (PSG) remain the gold standard for evaluating sleep quality. However, PSG manual annotation by experts is expensive and time-prohibitive. We propose SERF, interpretable Sleep staging using Embeddings, Rules, and Features to read PSG. SERF provides interpretation of classified sleep stages through meaningful features derived from the AASM Manual for the Scoring of Sleep and Associated Events. In SERF, the embeddings obtained from a hybrid of convolutional and recurrent neural networks are transposed to the interpretable feature space. These representative interpretable features are used to train simple models like a shallow decision tree for classification. Model results are validated on two publicly available datasets. SERF surpasses the current state-of-the-art for interpretable sleep staging by 2%. Using Gradient Boosted Trees as the classifier, SERF obtains 0.766 $\kappa$ and 0.870 AUC-ROC, within 2% of the current state-of-the-art black-box models.
SLEEPER: interpretable Sleep staging via Prototypes from Expert Rules
Oct 14, 2019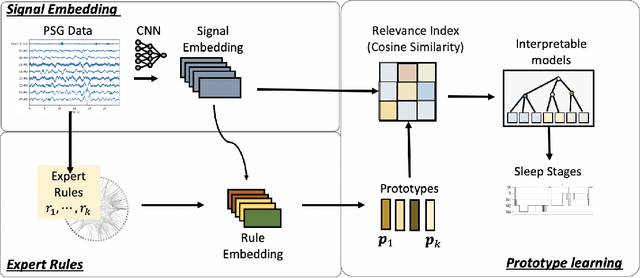
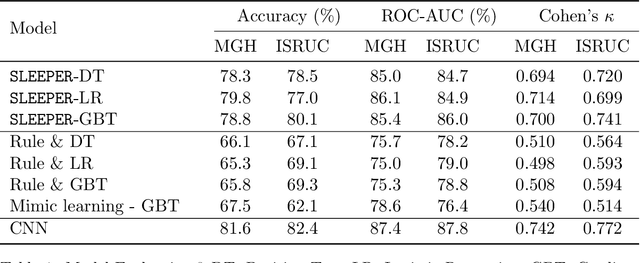
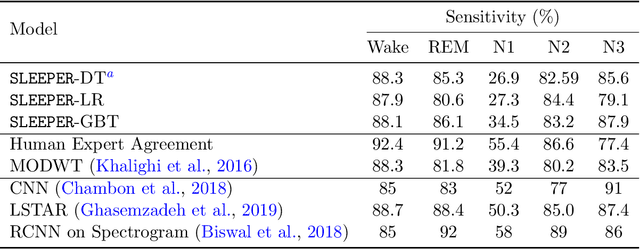
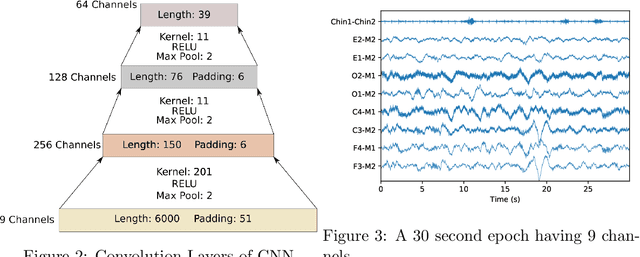
Abstract:Sleep staging is a crucial task for diagnosing sleep disorders. It is tedious and complex as it can take a trained expert several hours to annotate just one patient's polysomnogram (PSG) from a single night. Although deep learning models have demonstrated state-of-the-art performance in automating sleep staging, interpretability which defines other desiderata, has largely remained unexplored. In this study, we propose Sleep staging via Prototypes from Expert Rules (SLEEPER), which combines deep learning models with expert defined rules using a prototype learning framework to generate simple interpretable models. In particular, SLEEPER utilizes sleep scoring rules and expert defined features to derive prototypes which are embeddings of PSG data fragments via convolutional neural networks. The final models are simple interpretable models like a shallow decision tree defined over those phenotypes. We evaluated SLEEPER using two PSG datasets collected from sleep studies and demonstrated that SLEEPER could provide accurate sleep stage classification comparable to human experts and deep neural networks with about 85% ROC-AUC and .7 kappa.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge