Holger R. Roth
NVIDIA, USA
Precise Estimation of Renal Vascular Dominant Regions Using Spatially Aware Fully Convolutional Networks, Tensor-Cut and Voronoi Diagrams
Aug 05, 2019
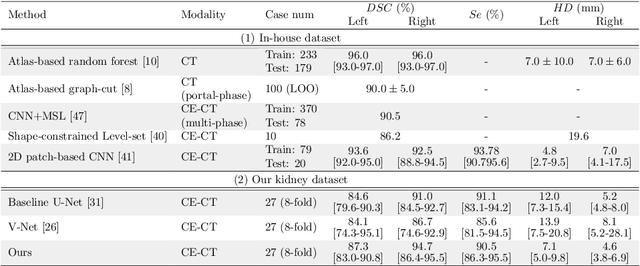
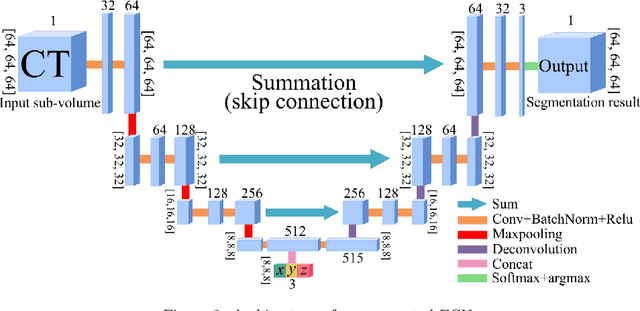
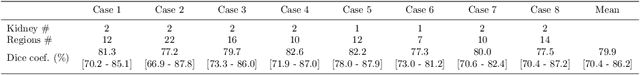
Abstract:This paper presents a new approach for precisely estimating the renal vascular dominant region using a Voronoi diagram. To provide computer-assisted diagnostics for the pre-surgical simulation of partial nephrectomy surgery, we must obtain information on the renal arteries and the renal vascular dominant regions. We propose a fully automatic segmentation method that combines a neural network and tensor-based graph-cut methods to precisely extract the kidney and renal arteries. First, we use a convolutional neural network to localize the kidney regions and extract tiny renal arteries with a tensor-based graph-cut method. Then we generate a Voronoi diagram to estimate the renal vascular dominant regions based on the segmented kidney and renal arteries. The accuracy of kidney segmentation in 27 cases with 8-fold cross validation reached a Dice score of 95%. The accuracy of renal artery segmentation in 8 cases obtained a centerline overlap ratio of 80%. Each partition region corresponds to a renal vascular dominant region. The final dominant-region estimation accuracy achieved a Dice coefficient of 80%. A clinical application showed the potential of our proposed estimation approach in a real clinical surgical environment. Further validation using large-scale database is our future work.
3D FCN Feature Driven Regression Forest-Based Pancreas Localization and Segmentation
Jun 08, 2018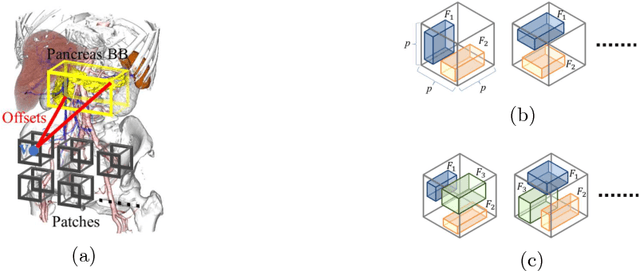
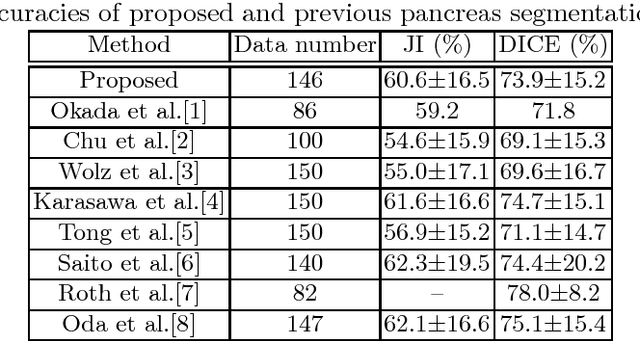
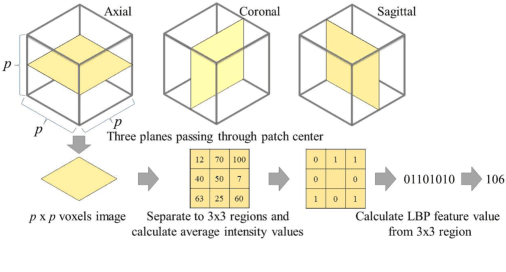
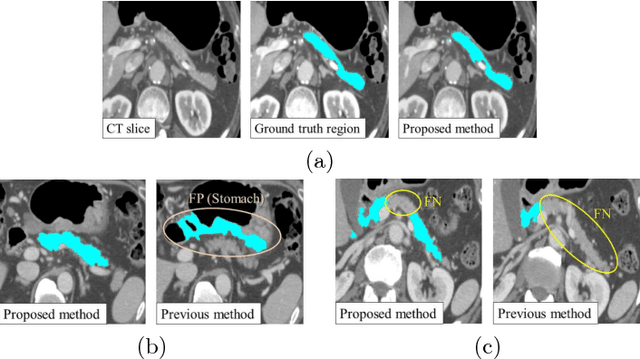
Abstract:This paper presents a fully automated atlas-based pancreas segmentation method from CT volumes utilizing 3D fully convolutional network (FCN) feature-based pancreas localization. Segmentation of the pancreas is difficult because it has larger inter-patient spatial variations than other organs. Previous pancreas segmentation methods failed to deal with such variations. We propose a fully automated pancreas segmentation method that contains novel localization and segmentation. Since the pancreas neighbors many other organs, its position and size are strongly related to the positions of the surrounding organs. We estimate the position and the size of the pancreas (localized) from global features by regression forests. As global features, we use intensity differences and 3D FCN deep learned features, which include automatically extracted essential features for segmentation. We chose 3D FCN features from a trained 3D U-Net, which is trained to perform multi-organ segmentation. The global features include both the pancreas and surrounding organ information. After localization, a patient-specific probabilistic atlas-based pancreas segmentation is performed. In evaluation results with 146 CT volumes, we achieved 60.6% of the Jaccard index and 73.9% of the Dice overlap.
* Presented in MICCAI 2017 workshop, DLMIA 2017 (Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support)
A multi-scale pyramid of 3D fully convolutional networks for abdominal multi-organ segmentation
Jun 06, 2018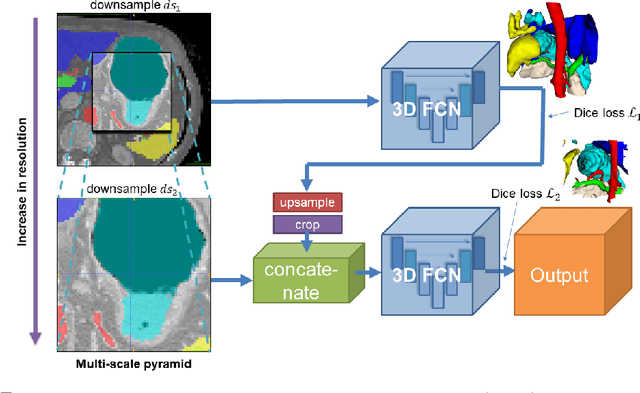
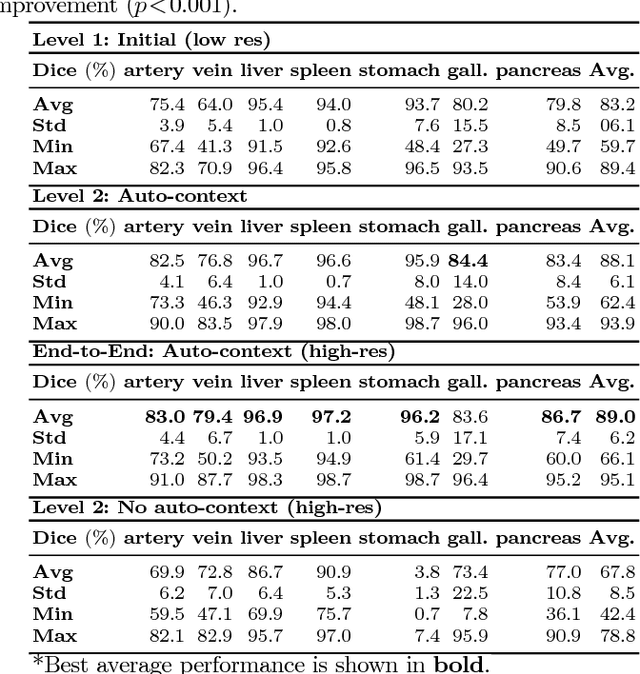
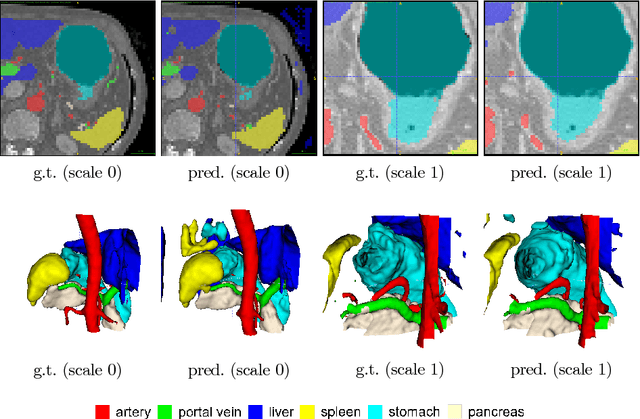
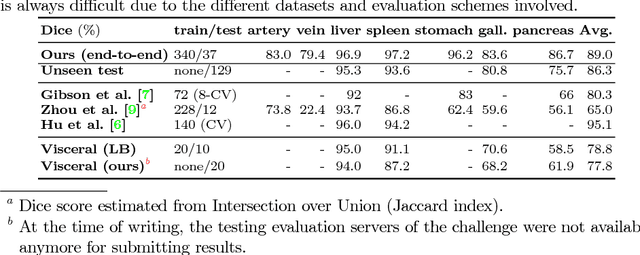
Abstract:Recent advances in deep learning, like 3D fully convolutional networks (FCNs), have improved the state-of-the-art in dense semantic segmentation of medical images. However, most network architectures require severely downsampling or cropping the images to meet the memory limitations of today's GPU cards while still considering enough context in the images for accurate segmentation. In this work, we propose a novel approach that utilizes auto-context to perform semantic segmentation at higher resolutions in a multi-scale pyramid of stacked 3D FCNs. We train and validate our models on a dataset of manually annotated abdominal organs and vessels from 377 clinical CT images used in gastric surgery, and achieve promising results with close to 90% Dice score on average. For additional evaluation, we perform separate testing on datasets from different sources and achieve competitive results, illustrating the robustness of the model and approach.
Unsupervised Segmentation of 3D Medical Images Based on Clustering and Deep Representation Learning
Apr 11, 2018Abstract:This paper presents a novel unsupervised segmentation method for 3D medical images. Convolutional neural networks (CNNs) have brought significant advances in image segmentation. However, most of the recent methods rely on supervised learning, which requires large amounts of manually annotated data. Thus, it is challenging for these methods to cope with the growing amount of medical images. This paper proposes a unified approach to unsupervised deep representation learning and clustering for segmentation. Our proposed method consists of two phases. In the first phase, we learn deep feature representations of training patches from a target image using joint unsupervised learning (JULE) that alternately clusters representations generated by a CNN and updates the CNN parameters using cluster labels as supervisory signals. We extend JULE to 3D medical images by utilizing 3D convolutions throughout the CNN architecture. In the second phase, we apply k-means to the deep representations from the trained CNN and then project cluster labels to the target image in order to obtain the fully segmented image. We evaluated our methods on three images of lung cancer specimens scanned with micro-computed tomography (micro-CT). The automatic segmentation of pathological regions in micro-CT could further contribute to the pathological examination process. Hence, we aim to automatically divide each image into the regions of invasive carcinoma, noninvasive carcinoma, and normal tissue. Our experiments show the potential abilities of unsupervised deep representation learning for medical image segmentation.
* This paper was presented at SPIE Medical Imaging 2018, Houston, TX, USA
Unsupervised Pathology Image Segmentation Using Representation Learning with Spherical K-means
Apr 11, 2018Abstract:This paper presents a novel method for unsupervised segmentation of pathology images. Staging of lung cancer is a major factor of prognosis. Measuring the maximum dimensions of the invasive component in a pathology images is an essential task. Therefore, image segmentation methods for visualizing the extent of invasive and noninvasive components on pathology images could support pathological examination. However, it is challenging for most of the recent segmentation methods that rely on supervised learning to cope with unlabeled pathology images. In this paper, we propose a unified approach to unsupervised representation learning and clustering for pathology image segmentation. Our method consists of two phases. In the first phase, we learn feature representations of training patches from a target image using the spherical k-means. The purpose of this phase is to obtain cluster centroids which could be used as filters for feature extraction. In the second phase, we apply conventional k-means to the representations extracted by the centroids and then project cluster labels to the target images. We evaluated our methods on pathology images of lung cancer specimen. Our experiments showed that the proposed method outperforms traditional k-means segmentation and the multithreshold Otsu method both quantitatively and qualitatively with an improved normalized mutual information (NMI) score of 0.626 compared to 0.168 and 0.167, respectively. Furthermore, we found that the centroids can be applied to the segmentation of other slices from the same sample.
* This paper was presented at SPIE Medical Imaging 2018, Houston, TX, USA
Deep learning and its application to medical image segmentation
Mar 23, 2018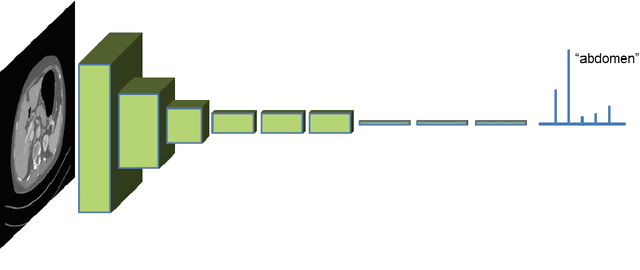
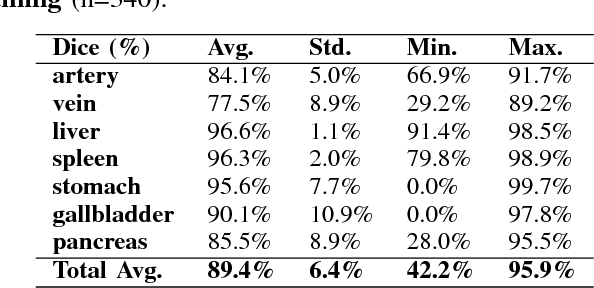
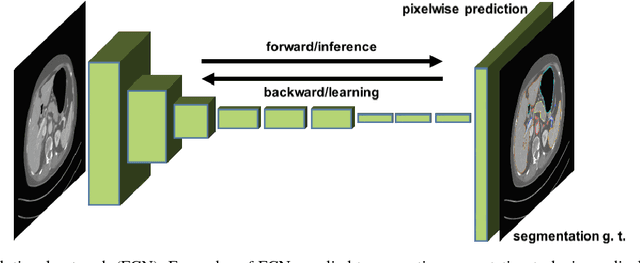
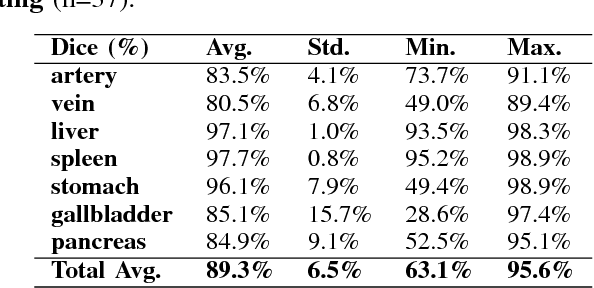
Abstract:One of the most common tasks in medical imaging is semantic segmentation. Achieving this segmentation automatically has been an active area of research, but the task has been proven very challenging due to the large variation of anatomy across different patients. However, recent advances in deep learning have made it possible to significantly improve the performance of image recognition and semantic segmentation methods in the field of computer vision. Due to the data driven approaches of hierarchical feature learning in deep learning frameworks, these advances can be translated to medical images without much difficulty. Several variations of deep convolutional neural networks have been successfully applied to medical images. Especially fully convolutional architectures have been proven efficient for segmentation of 3D medical images. In this article, we describe how to build a 3D fully convolutional network (FCN) that can process 3D images in order to produce automatic semantic segmentations. The model is trained and evaluated on a clinical computed tomography (CT) dataset and shows state-of-the-art performance in multi-organ segmentation.
* Accepted for publication in the journal of the Japanese Society of Medical Imaging Technology (JAMIT)
An application of cascaded 3D fully convolutional networks for medical image segmentation
Mar 20, 2018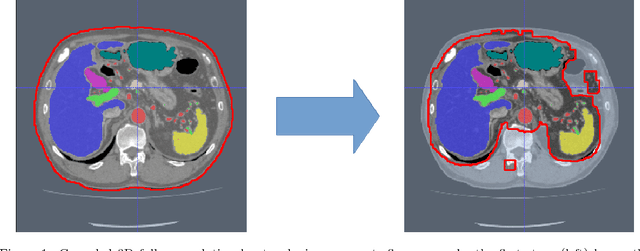
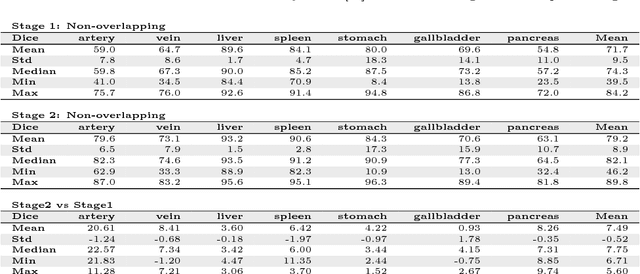
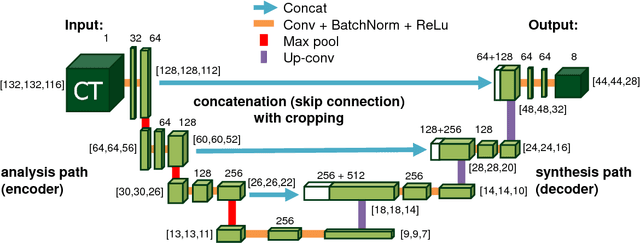
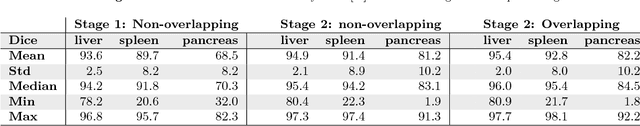
Abstract:Recent advances in 3D fully convolutional networks (FCN) have made it feasible to produce dense voxel-wise predictions of volumetric images. In this work, we show that a multi-class 3D FCN trained on manually labeled CT scans of several anatomical structures (ranging from the large organs to thin vessels) can achieve competitive segmentation results, while avoiding the need for handcrafting features or training class-specific models. To this end, we propose a two-stage, coarse-to-fine approach that will first use a 3D FCN to roughly define a candidate region, which will then be used as input to a second 3D FCN. This reduces the number of voxels the second FCN has to classify to ~10% and allows it to focus on more detailed segmentation of the organs and vessels. We utilize training and validation sets consisting of 331 clinical CT images and test our models on a completely unseen data collection acquired at a different hospital that includes 150 CT scans, targeting three anatomical organs (liver, spleen, and pancreas). In challenging organs such as the pancreas, our cascaded approach improves the mean Dice score from 68.5 to 82.2%, achieving the highest reported average score on this dataset. We compare with a 2D FCN method on a separate dataset of 240 CT scans with 18 classes and achieve a significantly higher performance in small organs and vessels. Furthermore, we explore fine-tuning our models to different datasets. Our experiments illustrate the promise and robustness of current 3D FCN based semantic segmentation of medical images, achieving state-of-the-art results. Our code and trained models are available for download: https://github.com/holgerroth/3Dunet_abdomen_cascade.
* Preprint accepted for publication in Computerized Medical Imaging and Graphics. Substantial extension of arXiv:1704.06382; Corrected references to figure numbers in this version
On the influence of Dice loss function in multi-class organ segmentation of abdominal CT using 3D fully convolutional networks
Jan 18, 2018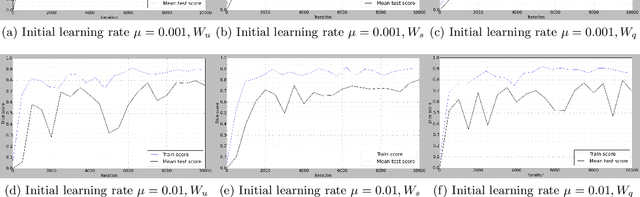
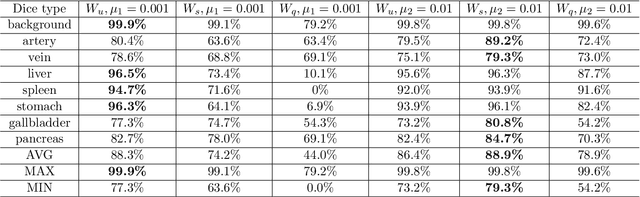
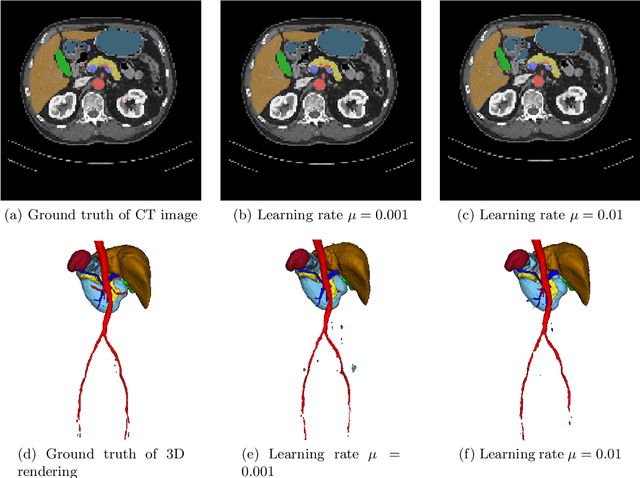
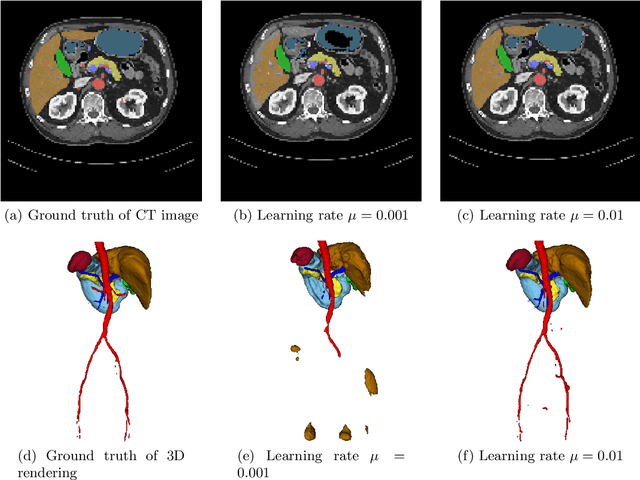
Abstract:Deep learning-based methods achieved impressive results for the segmentation of medical images. With the development of 3D fully convolutional networks (FCNs), it has become feasible to produce improved results for multi-organ segmentation of 3D computed tomography (CT) images. The results of multi-organ segmentation using deep learning-based methods not only depend on the choice of networks architecture, but also strongly rely on the choice of loss function. In this paper, we present a discussion on the influence of Dice-based loss functions for multi-class organ segmentation using a dataset of abdominal CT volumes. We investigated three different types of weighting the Dice loss functions based on class label frequencies (uniform, simple and square) and evaluate their influence on segmentation accuracies. Furthermore, we compared the influence of different initial learning rates. We achieved average Dice scores of 81.3%, 59.5% and 31.7% for uniform, simple and square types of weighting when the learning rate is 0.001, and 78.2%, 81.0% and 58.5% for each weighting when the learning rate is 0.01. Our experiments indicated a strong relationship between class balancing weights and initial learning rate in training.
Hierarchical 3D fully convolutional networks for multi-organ segmentation
Apr 21, 2017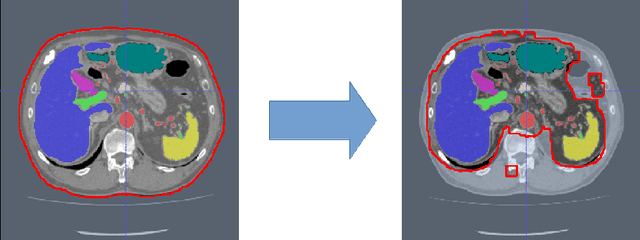
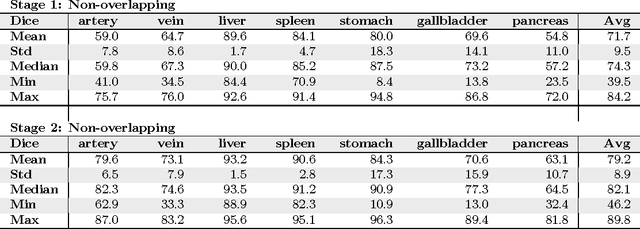
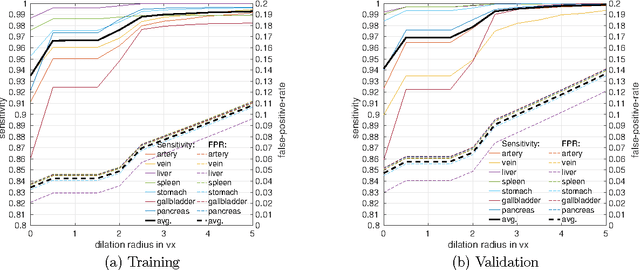
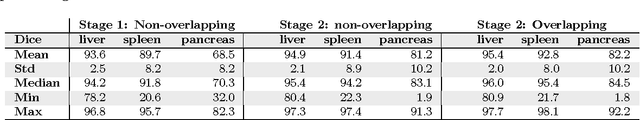
Abstract:Recent advances in 3D fully convolutional networks (FCN) have made it feasible to produce dense voxel-wise predictions of full volumetric images. In this work, we show that a multi-class 3D FCN trained on manually labeled CT scans of seven abdominal structures (artery, vein, liver, spleen, stomach, gallbladder, and pancreas) can achieve competitive segmentation results, while avoiding the need for handcrafting features or training organ-specific models. To this end, we propose a two-stage, coarse-to-fine approach that trains an FCN model to roughly delineate the organs of interest in the first stage (seeing $\sim$40% of the voxels within a simple, automatically generated binary mask of the patient's body). We then use these predictions of the first-stage FCN to define a candidate region that will be used to train a second FCN. This step reduces the number of voxels the FCN has to classify to $\sim$10% while maintaining a recall high of $>$99%. This second-stage FCN can now focus on more detailed segmentation of the organs. We respectively utilize training and validation sets consisting of 281 and 50 clinical CT images. Our hierarchical approach provides an improved Dice score of 7.5 percentage points per organ on average in our validation set. We furthermore test our models on a completely unseen data collection acquired at a different hospital that includes 150 CT scans with three anatomical labels (liver, spleen, and pancreas). In such challenging organs as the pancreas, our hierarchical approach improves the mean Dice score from 68.5 to 82.2%, achieving the highest reported average score on this dataset.
Comparison of the Deep-Learning-Based Automated Segmentation Methods for the Head Sectioned Images of the Virtual Korean Human Project
Mar 15, 2017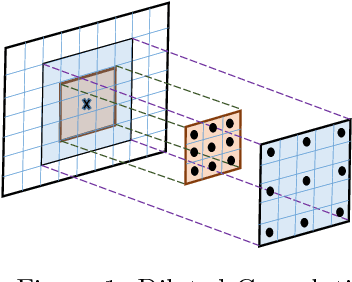
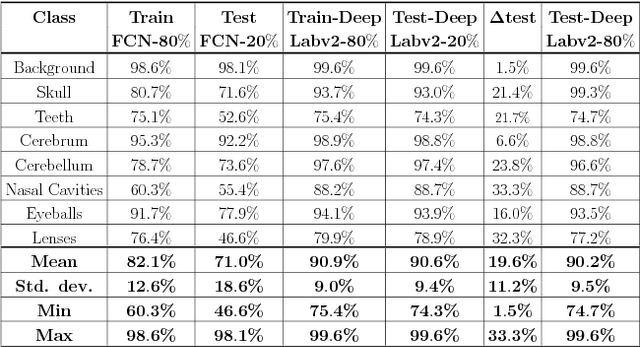
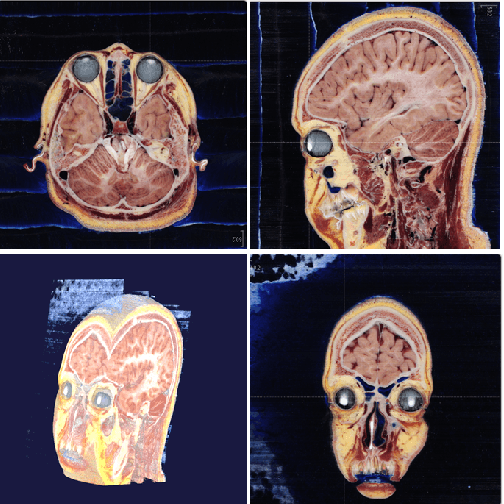
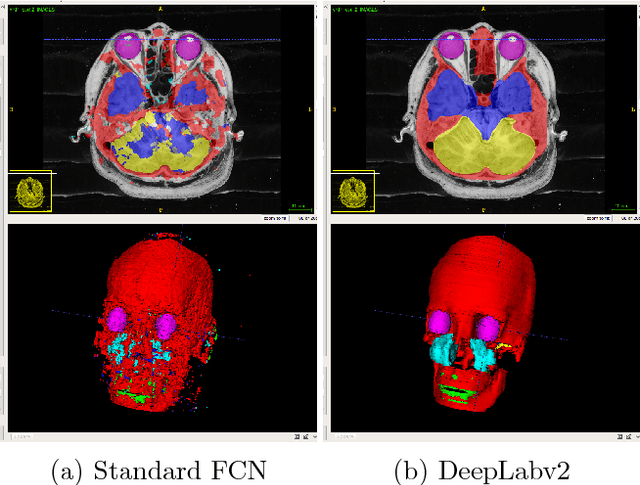
Abstract:This paper presents an end-to-end pixelwise fully automated segmentation of the head sectioned images of the Visible Korean Human (VKH) project based on Deep Convolutional Neural Networks (DCNNs). By converting classification networks into Fully Convolutional Networks (FCNs), a coarse prediction map, with smaller size than the original input image, can be created for segmentation purposes. To refine this map and to obtain a dense pixel-wise output, standard FCNs use deconvolution layers to upsample the coarse map. However, upsampling based on deconvolution increases the number of network parameters and causes loss of detail because of interpolation. On the other hand, dilated convolution is a new technique introduced recently that attempts to capture multi-scale contextual information without increasing the network parameters while keeping the resolution of the prediction maps high. We used both a standard FCN and a dilated convolution based FCN for semantic segmentation of the head sectioned images of the VKH dataset. Quantitative results showed approximately 20% improvement in the segmentation accuracy when using FCNs with dilated convolutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge